Abstract
Niabella soli Weon et al. 2008 is a member of the Chitinophagaceae, a family within the class Sphingobacteriia that is poorly characterized at the genome level, thus far. N. soli strain JS13-8T is of interest for its ability to produce a variety of glycosyl hydrolases. The genome of N. soli strain JS13-8T is only the second genome sequence of a type strain from the family Chitinophagaceae to be published, and the first one from the genus Niabella. Here we describe the features of this organism, together with the complete genome sequence and annotation. The 4,697,343 bp long chromosome with its 3,931 protein-coding and 49 RNA genes is a part of the Genomic Encyclopedia of Bacteria and Archaea project.
Keywords: aerobic, non-motile, Gram-negative, mesophilic, chemoorganotrophic, glycosyl hydrolases, soil, Chitinophagaceae, GEBA
Introduction
Strain JS13-8T (= DSM 19437 = KACC 12604) is the type strain of the species Niabella soli [1], one out of five species in the genus Niabella [2]. The strain was originally isolated from soil sampled from Jeju Island, Republic of Korea [1]. The genus name was derived from the arbitrary word NIAB, National Institute of Agricultural Biotechnology, where taxonomic studies of this organism were conducted [3]; the species epithet was derived from the Latin word soli, of soil [1]. Strain JS13-8T was found to assimilate several mono- and disaccharides and to produce numerous glycosyl hydrolases [1]. There are no PubMed records that document the use of the strain for any biotechnological studies; only comparative analyses performed for the description of later members of the genus Niabella are recorded. Here we present a summary classification and a set of features for N. soli JS13-8T, together with the description of the genomic sequencing and annotation.
Classification and features
A representative genomic 16S rRNA sequence of N. soli JS13-8T was compared using NCBI BLAST [4,5] under default settings (e.g., considering only the high-scoring segment pairs (HSPs) from the best 250 hits) with the most recent release of the Greengenes database [6]. The relative frequencies of taxa and keywords (reduced to their stem [7]) were determined, weighted by BLAST scores. The most frequently occurring genera were Niabella (34.8%), Terrimonas (21.0%), Flavobacterium (14.9%), 'Niablella' (8.5%; an apparent misspelling of Niabella) and Niastella (8.2%) (13 hits in total). Regarding the single hit to sequences from members of the species, the average identity within HSPs was 99.7%, whereas the average coverage by HSPs was 96.8%. Among all other species, the one yielding the highest score was 'Niablella koreensis' (DQ457019; again a misnomer, see Figure 1), which corresponded to an identity of 95.1% and an HSP coverage of 99.9%. (Note that the Greengenes database uses the INSDC (= EMBL/NCBI/DDBJ) annotation, which is not an authoritative source for nomenclature or classification.) The highest-scoring environmental sequence was JF167633 ('skin antecubital fossa clone ncd2016g05c1'), which showed an identity of 95.3% and an HSP coverage of 95.7%. The most frequently occurring keywords within the labels of all environmental samples which yielded hits were 'sludg' (3.6%), 'activ' (2.6%), 'skin' (2.3%), 'wast' (1.8%) and 'soil' (1.8%) (236 hits in total) and reveal no deeper insight into the usual habitat of close relatives of the strain. Environmental samples which yielded hits of a higher score than the highest scoring species were not found, indicating that N. soli itself is rarely found in environmental screenings.
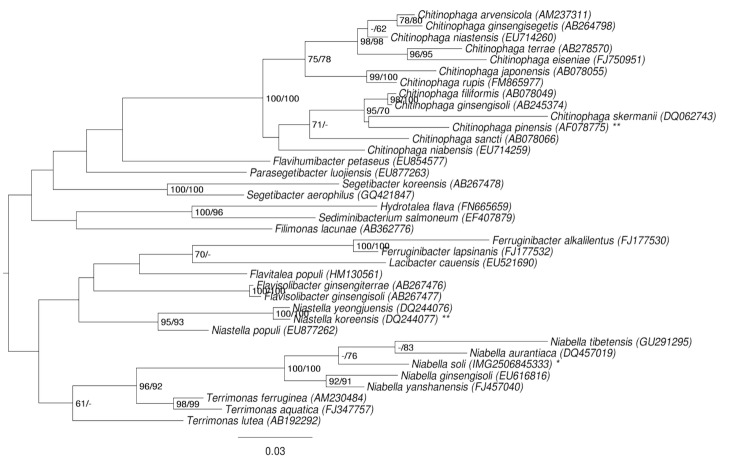
Figure 1.
Phylogenetic tree highlighting the position of N. soli relative to the type strains of the other species within the family Chitinophagaceae except for the genera Balneola and Gracilimonas. The tree was inferred from 1,395 aligned characters [8,9] of the 16S rRNA gene sequence under the maximum likelihood (ML) criterion [10]. Rooting was done initially using the midpoint method [11] and then checked for its agreement with the current classification (Table 1). The branches are scaled in terms of the expected number of substitutions per site. Numbers adjacent to the branches are support values from 950 ML bootstrap replicates [12] (left) and from 1,000 maximum-parsimony bootstrap replicates [13] (right) if larger than 60%. Lineages with type strain genome sequencing projects registered in GOLD [14] are labeled with one asterisk, those also listed as 'Complete and Published' with two asterisks [15] (for Niastella koreensis see CP003178).
Table 1. Classification and general features of N. soli JS13-8T according to the MIGS recommendations [16], List of Prokaryotic names with Standing in Nomenclature [17] and the Names for Life database [2].
| MIGS ID | Property | Term | Evidence code |
|---|---|---|---|
| Current classification | Domain Bacteria | TAS [18] | |
| Phylum Bacteroidetes | TAS [19,20] | ||
| Class Sphingobacteriia | TAS [19,21] | ||
| Order Sphingobacteriales | TAS [19,22] | ||
| Family Chitinophagaceae | TAS [23,24] | ||
| Genus Niabella | TAS [3,23,25] | ||
| Species Niabella soli | TAS [1] | ||
| Type-strain JS13-8 | TAS [1] | ||
| Gram stain | negative | TAS [1] | |
| Cell shape | short rods | TAS [1] | |
| Motility | non-motile | TAS [1] | |
| Sporulation | non-sporulating | NAS | |
| Temperature range | mesophile, 15-35°C | TAS [1] | |
| Optimum temperature | 30°C | TAS [1] | |
| Salinity | 0-1% NaCl (w/v) | TAS [3] | |
| MIGS-22 | Oxygen requirement | aerobe | TAS [1] |
| Carbon source | mono- and polysaccharides | TAS [1] | |
| Energy metabolism | chemoorganotroph | TAS [1] | |
| MIGS-6 | Habitat | soil | TAS [1] |
| MIGS-15 | Biotic relationship | free living | TAS [1] |
| MIGS-14 | Pathogenicity | none | NAS |
| Biosafety level | 1 | TAS [26] | |
| MIGS-23.1 | Isolation | soil sample | TAS [1] |
| MIGS-4 | Geographic location | Jeju Island, Republic of Korea | TAS [1] |
| MIGS-5 | Sample collection time | not reported | |
| MIGS-4.1 | Latitude | 33.37 | TAS [1] |
| Longitude | MIGS-4.2 | 126.566 | TAS [1] |
| MIGS-4.3 | Depth | not reported | |
| MIGS-4.4 | Altitude | not reported |
Evidence codes - TAS: Traceable Author Statement (i.e., a direct report exists in the literature); NAS: Non-traceable Author Statement (i.e., not directly observed for the living, isolated sample, but based on a generally accepted property for the species, or anecdotal evidence). Evidence codes are from the Gene Ontology project [27].
Figure 1 shows the phylogenetic neighborhood of N. soli in a 16S rRNA based tree. The sequences of the two 16S rRNA gene copies in the genome differ from each other by one nucleotide, and differ by up to one nucleotide from the previously published 16S rRNA sequence (EF592608), which contains three ambiguous base calls.
In a preliminary phylogenetic analysis of the 16S rRNA sequences from the family, we observed that two genera, Balneola and Gracilimonas, listed as belonging to Chitinophagaceae by [17,28,29], formed the root of the tree and were separated from the remaining taxa by quite long branches. For this reason, they were omitted from the analysis described above, and a second phylogenetic analysis involving the type species of the type genera of all families within the phylum Bacteroidetes was conducted, either unconstrained or constrained for the monophyly of all families [30]. The alignment (inferred and filtered as described above) contained 17 operational taxonomic units and 1,384 characters. The best ML tree found had a log likelihood of -12,076.19, whereas the best trees found under the constraint had a log likelihood of -12,132.94. The constrained tree was significantly worse than the globally best one in the Shimodaira-Hasegawa test as implemented in RAxML [10] (α = 0.01). The bestMP trees found had a score of 2,432, whereas the best constrained tree found had a score of 2,485 and was significantly worse in the Kishino-Hasegawa test as implemented in PAUP* [13] (α = 0.01). (See, e.g. chapter 21 in [31] for an in-depth description of such paired-site tests.) This confirms our view that Balneola and Gracilimonas are misplaced as members of Chitinophagaceae (as all other families were represented by a single taxon only, Chitinophagaceae is the only family that might have caused conflict in this setting). Chitinophagaceae should thus be regarded to only contain the genera listed by [23] together with the more recently published genus Flavitalea [28].
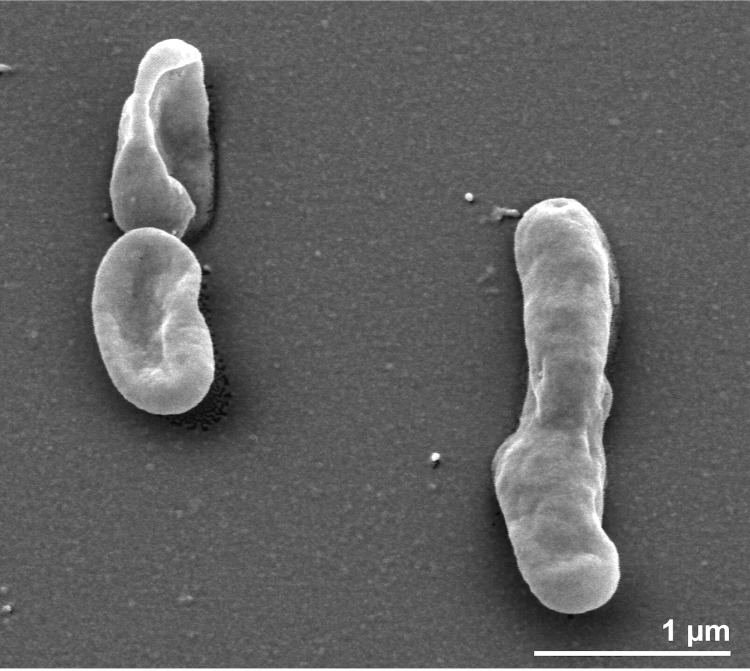
N. soli JS13-8T is a Gram-negative and non-motile aerobic bacterium [1]. Cells are short rods 0.8-1.4 μm long and with a diameter of 0.5-0.7 μm ([1], Figure 2). Colonies are dark yellow due to the pigment flexirubin [1]. Growth was observed between 15°C and 35°C with an optimum at 30°C [1]. The pH range for growth was 5.0-8.0 with 6.0-7.0 as the optimum [1]. The salinity range for growth was 0-1% NaCl [3]. N. soli JS13-8T grows on several monosaccharides, disaccharides, gluconate, and D-mannitol [1]. It produces numerous glycosyl hydrolases including α-galactosidase, β-galactosidase, β-glucuronidase, α-glucosidase, β-glucosidase, N-acetyl-β-glucosaminidase, α-mannosidase, and α-fucosidase [1]. However it did not hydrolyze starch, chitin, or carboxymethylcellulose [1].
Figure 2.
Scanning electron micrograph of N. soli JS13-8T
Chemotaxonomy
The major respiratory quinone found in N. soli JS13-8T was MK-7, and the major fatty acids identified were iso-C15:0 (29.2%), iso-C15:1 G (18.4%), iso-C17:0 3-OH (11.8%), and summed feature 3 (11.1%), which is generally reported to include iso-C15:0 2-OH and/or C16:1 ω7c, although careful examination of the MIDI fatty acid reports generally allow a more precise identification [1]. Smaller amounts of anteiso-C15 : 0 (1.2%), iso-C15:0 3-OH (2.2%), C16:0 (6.8%), C16:0 2-OH (1.3%), C16:0 3-OH (2.2%), C18:0 (3.8%), C18:1 ω7c (1.5%), C18:1 ω9c (1.0%), Summed feature 5 (comprising anteiso-C18:0 and/or C18:2 ω6,9c 3.4%) and an unknown peak with an equivalent chain length of 13.565 (1.1%) were also detected. The presence of major amounts of branched chain saturated and unsaturated fatty acids, together with significant amounts of 3-OH and 2-OH fatty acids is characteristic of members of this evolutionary group and also points to the presence of characteristic lipids, for which data is missing from this strain.
Genome sequencing and annotation
Genome project history
This organism was selected for sequencing on the basis of its phylogenetic position [32], and is part of the Genomic Encyclopedia of Bacteria and Archaea project [33]. The genome project is deposited in the Genomes On Line Database [14] and the complete genome sequence is deposited in GenBank. Sequencing, finishing and annotation were performed by the DOE Joint Genome Institute (JGI). A summary of the project information is shown in Table 2.
Table 2. Genome sequencing project information.
| MIGS ID | Property | Term |
|---|---|---|
| MIGS-31 | Finishing quality | Finished |
| MIGS-28 | Libraries used | Five genomic libraries: two 454 pyrosequence standard libraries, two 454 PE library (13 kb and 20 kb insert size), one Illumina library |
| MIGS-29 | Sequencing platforms | Illumina GAii, 454 GS FLX Titanium |
| MIGS-31.2 | Sequencing coverage | 113.0 × Illumina; 23.6 × pyrosequence |
| MIGS-30 | Assemblers | Newbler version 2.3, Velvet version 1.0.13, phrap version SPS - 4.24 |
| MIGS-32 | Gene calling method | Prodigal |
| INSDC ID | AGSA00000000 | |
| GenBank Date of Release | January 19, 2012 | |
| GOLD ID | Gi04680 | |
| NCBI project ID | 61269 | |
| Database: IMG | 2506783006 | |
| MIGS-13 | Source material identifier | DSM 19437 |
| Project relevance | Tree of Life, GEBA |
Growth conditions and DNA isolation
N. soli strain JS13-8T, DSM 19437, was grown in DSMZ medium 830 (R2A medium) [34] at 37°C. DNA was isolated from 0.5-1 g of cell paste using MasterPure Gram-positive DNA purification kit (Epicentre MGP04100) following the standard protocol as recommended by the manufacturer with modification st/DL for cell lysis as described in Wu et al. 2009 [33]. DNA is available through the DNA Bank Network [35].
Genome sequencing and assembly
The genome was sequenced using a combination of Illumina and 454 sequencing platforms. All general aspects of library construction and sequencing can be found at the JGI website [36]. Pyrosequencing reads were assembled using the Newbler assembler (Roche). The initial Newbler assembly consisting of 15 contigs in one scaffold was converted into a phrap [37] assembly by making fake reads from the consensus, to collect the read pairs in the 454 paired end library. Illumina GAii sequencing data (1,116.9 Mb) was assembled with Velvet [38] and the consensus sequences were shredded into 1.5 kb overlapped fake reads and assembled together with the 454 data. The 454 draft assembly was based on 158.8 Mb of 454 draft data and all of the 454 paired end data. Newbler parameters are -consed -a 50 -l 350 -g -m -ml 20. The Phred/Phrap/Consed software package [37] was used for sequence assembly and quality assessment in the subsequent finishing process. After the shotgun stage, reads were assembled with parallel phrap (High Performance Software, LLC). Possible mis-assemblies were corrected with gapResolution [36], Dupfinisher [39], or sequencing cloned bridging PCR fragments with subcloning. Gaps between contigs were closed by editing in Consed, by PCR and by Bubble PCR primer walks (J.-F. Chang, unpublished). A total of 45 additional reactions were necessary to close gaps and to raise the quality of the finished sequence. Illumina reads were also used to correct potential base errors and increase consensus quality using a software Polisher developed at JGI [40]. The error rate of the completed genome sequence is less than 1 in 100,000. Together, the combination of the Illumina and 454 sequencing platforms provided 136.6 × coverage of the genome. The final assembly contained 354,991 pyrosequence and 14,750,629 Illumina reads.
Genome annotation
Genes were identified using Prodigal [41] as part of the Oak Ridge National Laboratory genome annotation pipeline, followed by a round of manual curation using the JGI GenePRIMP pipeline [42]. The predicted CDSs were translated and used to search the National Center for Biotechnology Information (NCBI) non-redundant database, UniProt, TIGRFam, Pfam, PRIAM, KEGG, COG, and InterPro databases. These data sources were combined to assert a product description for each predicted protein. Non-coding genes and miscellaneous features were predicted using tRNAscan-SE [43], RNAmmer [44], Rfam [45], TMHMM [46], and signalP [47].
Genome properties
The genome consists of one circular chromosome of 4,697,343 bp length with a 45.2% G+C content (Table 3 and Figure 3). Of the 3,932 genes predicted, 3,882 were protein-coding genes, and 49 RNAs; 34 pseudogenes were also identified. The majority of the protein-coding genes (71.9%) were assigned a putative function while the remaining ones were annotated as hypothetical proteins. The distribution of genes into COGs functional categories is presented in Table 4.
Table 3. Genome Statistics.
| Attribute | Value | % of Total |
|---|---|---|
| Genome size (bp) | 4,697,343 | 100.00% |
| DNA coding region (bp) | 4,154,623 | 88.45% |
| DNA G+C content (bp) | 2,124,959 | 45.24% |
| Number of replicons | 1 | |
| Extrachromosomal elements | 0 | |
| Total genes | 3,931 | 100.00% |
| RNA genes | 6 | 0.15% |
| rRNA operons | 2 | |
| tRNA genes | 49 | 1.25% |
| Protein-coding genes | 3,882 | 98.75% |
| Pseudo genes | 34 | 0.86% |
| Genes with function prediction (proteins) | 2,827 | 71.92% |
| Genes in paralog clusters | 1,833 | 46.63% |
| Genes assigned to COGs | 2,734 | 69.55% |
| Genes assigned Pfam domains | 2,915 | 74.15% |
| Genes with signal peptides | 1,273 | 32.38% |
| Genes with transmembrane helices | 924 | 23.51% |
| CRISPR repeats | 1 |
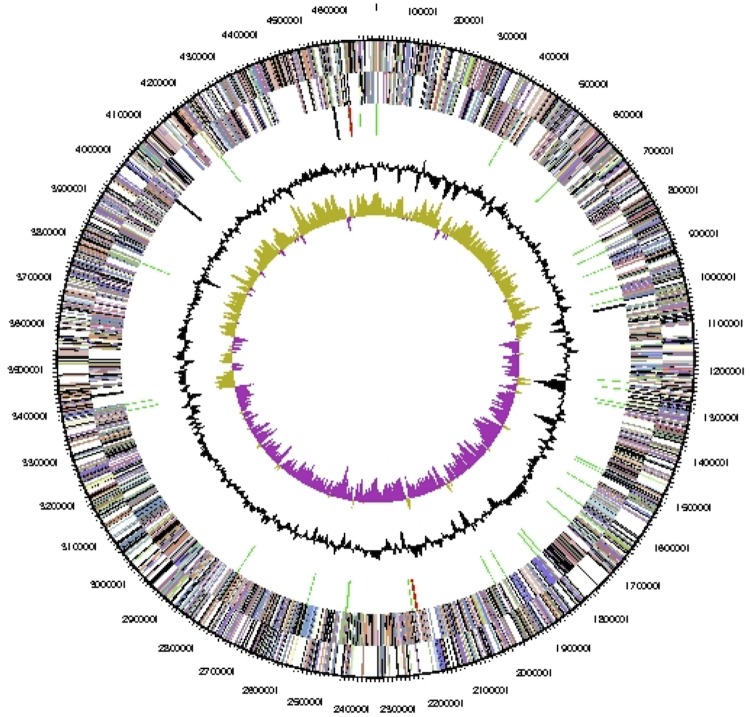
Figure 3.
Graphical map of the chromosome. From outside to the center: Genes on forward strand (colored by COG categories), Genes on reverse strand (colored by COG categories), RNA genes (tRNAs green, rRNAs red, other RNAs black), GC content (black), GC skew (purple/olive).
Table 4. Number of genes associated with the general COG functional categories.
| Code | value | %age | Description |
|---|---|---|---|
| J | 156 | 5.2 | Translation, ribosomal structure and biogenesis |
| A | 0 | 0.0 | RNA processing and modification |
| K | 204 | 6.8 | Transcription |
| L | 115 | 3.9 | Replication, recombination and repair |
| B | 0 | 0.0 | Chromatin structure and dynamics |
| D | 21 | 0.7 | Cell cycle control, cell division, chromosome partitioning |
| Y | 0 | 0.0 | Nuclear structure |
| V | 68 | 2.3 | Defense mechanisms |
| T | 132 | 4.4 | Signal transduction mechanisms |
| M | 239 | 8.0 | Cell wall/membrane biogenesis |
| N | 5 | 0.2 | Cell motility |
| Z | 1 | 0.0 | Cytoskeleton |
| W | 0 | 0.0 | Extracellular structures |
| U | 49 | 1.6 | Intracellular trafficking and secretion, and vesicular transport |
| O | 123 | 4.1 | Posttranslational modification, protein turnover, chaperones |
| C | 151 | 5.1 | Energy production and conversion |
| G | 293 | 9.8 | Carbohydrate transport and metabolism |
| E | 231 | 7.7 | Amino acid transport and metabolism |
| F | 69 | 2.3 | Nucleotide transport and metabolism |
| H | 143 | 4.8 | Coenzyme transport and metabolism |
| I | 104 | 3.5 | Lipid transport and metabolism |
| P | 178 | 6.0 | Inorganic ion transport and metabolism |
| Q | 51 | 1.7 | Secondary metabolites biosynthesis, transport and catabolism |
| R | 383 | 12.8 | General function prediction only |
| S | 268 | 9.0 | Function unknown |
| - | 1,197 | 30.5 | Not in COGs |
Insights into the genome sequence
Two other complete genomes are available in GenBank from the family Chitinophagaceae – Chitinophaga pinensis [15] and N. koreensis (unpublished) – and the permanent draft genome of Sediminibacterium sp. OR43 is available from the IMG/GEBA website [48]. Of these three organisms, N. soli is most closely related to N. koreensis (Figure 1). The genome size of N. soli is much smaller than those of N. koreensis and C. pinensis (9.0-9.1 Mbp) but larger than that of Sediminibacterium sp. OR43 (3.8 Mbp). Using the genome-to-genome distance calculator [49,50] version 2.0 revealed that 83.72% of all positions within HSPs are identical between the type-strain genomes of N. soli and C. pinensis, which corresponds to a DNA-DNA hybridization value of 26.60±2.42%. For N. koreensis, these values were 78.29% and 20.20±2.31%, respectively.
A major feature of the previously sequenced genomes from this family is the presence of large numbers of glycosyl hydrolases. N. koreensis has 228 glycosyl hydrolases, while C. pinensis has 187 [51]. We analyzed the genomes of N. soli and strain OR43 and found that they encode 164 and 86 glycosyl hydrolases, respectively. When viewed as a percentage of the total protein-coding sequences, glycosyl hydrolases constitute 4.2% of the N. soli genome and 3.1% of the N. koreensis genome. In the C. pinensis and OR43 genomes, glycosyl hydrolases account for 2.6% of the protein-coding genes. Thus N. soli has the highest density of glycosyl hydrolases in this family examined to date. In addition N. koreensis has 28 polysaccharide lyases while C. pinensis has only six [51]. We found that N. soli has 15 polysaccharide lyases and OR43 has only two. Thus N. soli also has a substantial number of polysaccharide lyases in addition to glycosyl hydrolases.
Of the glycosyl hydrolase families with many members in N. soli, some are also prevalent in N. koreensis and C. pinensis, for example families GH2, GH28, GH29, GH43, and GH78. However, there are GH families in which N. soli has a greater number of members than the genomes from other Chitinophagaceae – GH20 and GH106. N. soli also has enzymes from GH116 and GH123, which are not found in the other three genomes. There is also one GH family (GH92) for which N. soli has only two members, while N. koreensis and C. pinensis have 10 and 9, respectively.
In Bacteroides thetaiotaomicron, the SusC and SusD outer membrane proteins are required for starch utilization [52] and the B. thetaiotaomicron genome contains many proteins related to SusC and SusD [53]. The genomes from the family Chitinophagaceae also contain large numbers of these proteins. N. soli has 60 SusC family and 50 SusD family proteins, which is about half as many as in the larger N. koreensis and C. pinensis genomes.
The Chitinophagaceae appear to rely mainly on symporters for sugar transport. Only two sugar ABC transporters were found in N. soli, one in N. koreensis, and none in the other two genomes. The phosphotransferase system is not found in any of the four genomes. In contrast N. soli has 23 sugar symporters, N. koreensis has 27, C. pinensis has 14, and OR43 has 12. The sugar symporters belong to several families, with the most prevalent being the Major Facilitator Superfamily (TC 2.A.1) and the Solute:Sodium Symporter Family (TC 2.A.21).
Acknowledgements
We would like to gratefully acknowledge the help of Regine Fähnrich for growing N. soli cultures and Evelyne-Marie Brambilla for DNA extraction and quality control (both at DSMZ). This work was performed under the auspices of the US Department of Energy Office of Science, Biological and Environmental Research Program, and by the University of California, Lawrence Berkeley National Laboratory under contract No. DE-AC02-05CH11231, Lawrence Livermore National Laboratory under Contract No. DE-AC52-07NA27344, and Los Alamos National Laboratory under contract No. DE-AC02-06NA25396, UT-Battelle and Oak Ridge National Laboratory under contract DE-AC05-00OR22725, as well as German Research Foundation (DFG) INST 599/1-2.
References
- 1.Weon HY, Kim BY, Joa JH, Kwon SW, Kim WG, Koo BS. Niabella soli sp. nov., isolated from soil from Jeju Island, Korea. Int J Syst Evol Microbiol 2008; 58:467-6469 10.1099/ijs.0.65304-0 [DOI] [PubMed] [Google Scholar]
- 2.Garrity G. NamesforLife. BrowserTool takes expertise out of the database and puts it right in the browser. Microbiol Today 2010; 37:9 [Google Scholar]
- 3.Dai J, Jiang F, Wang Y, Yu B, Qi H, Fang C, Zheng C. Niabella tibetensis sp, nov., isolated from soil, and emended description of the genus Niabella. Int J Syst Evol Microbiol 2011; 61:1201-1205 10.1099/ijs.0.022103-0 [DOI] [PubMed] [Google Scholar]
- 4.Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990; 215:403-410 [DOI] [PubMed] [Google Scholar]
- 5.Korf I, Yandell M, Bedell J. BLAST, O'Reilly, Sebastopol, 2003. [Google Scholar]
- 6.DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 2006; 72:5069-5072 10.1128/AEM.03006-05 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Porter MF. An algorithm for suffix stripping. Program: electronic library and information systems 1980; 14:130-137.
- 8.Lee C, Grasso C, Sharlow MF. Multiple sequence alignment using partial order graphs. Bioinformatics 2002; 18:452-464 10.1093/bioinformatics/18.3.452 [DOI] [PubMed] [Google Scholar]
- 9.Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 2000; 17:540-552 10.1093/oxfordjournals.molbev.a026334 [DOI] [PubMed] [Google Scholar]
- 10.Stamatakis A, Hoover P, Rougemont J. A rapid bootstrap algorithm for the RAxML web servers. Syst Biol 2008; 57:758-771 10.1080/10635150802429642 [DOI] [PubMed] [Google Scholar]
- 11.Hess PN, De Moraes Russo CA. An empirical test of the midpoint rooting method. Biol J Linn Soc Lond 2007; 92:669-674 10.1111/j.1095-8312.2007.00864.x [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pattengale ND, Alipour M, Bininda-Emonds ORP, Moret BME, Stamatakis A. How many bootstrap replicates are necessary? Lect Notes Comput Sci 2009; 5541:184-200 10.1007/978-3-642-02008-7_13 [DOI] [PubMed] [Google Scholar]
- 13.Swofford DL. PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods), Version 4.0 b10. Sinauer Associates, Sunderland, 2002. [Google Scholar]
- 14.Pagani I, Liolios K, Jansson J, Chen IM, Smirnova T, Nosrat B, Markowitz VM, Kyrpides NC. The Genomes OnLine Database (GOLD) v.4: status of genomic and metagenomic projects and their associated metadata. Nucleic Acids Res 2012; 40:D571-D579 10.1093/nar/gkr1100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Glavina Del Rio T, Abt B, Spring S, Lapidus A, Nolan M, Tice H, Copeland A, Cheng JF, Chen F, Bruce D, et al. Complete genome sequence of Chitinophaga pinensis type strain (UQM 2034T). Stand Genomic Sci 2010; 2:87-95 10.4056/sigs.661199 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Field D, Garrity G, Gray T, Morrison N, Selengut J, Sterk P, Tatusova T, Thomson N, Allen MJ, Angiuoli SV, et al. The minimum information about a genome sequence (MIGS) specification. Nat Biotechnol 2008; 26:541-547 10.1038/nbt1360 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Euzéby JP. List of bacterial names with standing in nomenclature: A folder available on the Internet. Int J Syst Bacteriol 1997; 47:590-592 10.1099/00207713-47-2-590 [DOI] [PubMed] [Google Scholar]
- 18.Woese CR, Kandler O, Wheelis ML. Towards a natural system of organisms. Proposal for the domains Archaea and Bacteria. Proc Natl Acad Sci USA 1990; 87:4576-4579 10.1073/pnas.87.12.4576 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Editor L. Validation List No. 143. [PubMed]. Int J Syst Evol Microbiol 2012; 62:1-4; . 10.1099/ijs.0.039487-0 [DOI] [Google Scholar]
- 20.Krieg NR, Ludwig W, Euzéby J, Whitman WB. Phylum XIV. Bacteroidetes phyl. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 25. [Google Scholar]
- 21.Kämpfer P. Class III. Sphingobacteriia class. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011, p. 330. [Google Scholar]
- 22.Kämpfer P. Order I. Sphingobacteriales ord. nov. In: Krieg NR, Staley JT, Brown DR, Hedlund BP, Paster BJ, Ward NL, Ludwig W, Whitman WB (eds), Bergey's Manual of Systematic Bacteriology, Second Edition, Volume 4, Springer, New York, 2011. [Google Scholar]
- 23.Kämpfer P, Lodders N, Falsen E. Hydrotalea flava gen. nov., sp. nov., a new member of the phylum Bacteroidetes and allocation of the genera Chitinophaga, Sediminibacterium, Lacibacter, Flavihumibacter, Flavisolibacter, Niabella, Niastella, Segetibacter, Terrimonas, Ferruginibacter, Filimonas and Hydrotalea to the family Chitinophagaceae fam. nov. Int J Syst Evol Microbiol 2011; 61:518-523 10.1099/ijs.0.023002-0 [DOI] [PubMed] [Google Scholar]
- 24.Ludwig W, Euzeby J, Whitman WG. Draft taxonomic outline of the Bacteroidetes, Planctomycetes, Clamydiae, Spirochaetes, Fibrobacteres, Fusobacteria, Acidobacteria, Verrucomicrobia, Dictyoglomi, and Gemmatimonades, http//www.bergeys.org/outlines/ Bergeys_Vol4_Outline.pdf Taxonomic Outline 2008.
- 25.Kim BY, Weon HY, Yoo SH, Hong SB, Kwon SW, Stackebrandt E, Go SJ. Niabella aurantiaca gen. nov., sp. nov., isolated from a greenhouse soil in Korea. Int J Syst Evol Microbiol 2007; 57:538-541 10.1099/ijs.0.64614-0 [DOI] [PubMed] [Google Scholar]
- 26.BAuA. 2010, Classification of bacteria and archaea in risk groups. http://www.baua.de TRBA 466, p. 150.
- 27.Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT, et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat Genet 2000; 25:25-29 10.1038/75556 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Wang Y, Cai F, Tang Y, Dai J, Qi H, Rahman E, Peng F, Fang C. Flavitalea populi gen. nov., sp. nov., isolated from soil of a Euphrates poplar (Populus euphratica) forest. Int J Syst Evol Microbiol 2011; 61:1554-1560 10.1099/ijs.0.025221-0 [DOI] [PubMed] [Google Scholar]
- 29.Yarza P, Ludwig W, Euzéby J, Amann R, Schleifer KH, Glöckner FO, Rosselló-Móra R. Update of the All-Species Living Tree Project based on 16S and 23S rRNA sequence analyses. Syst Appl Microbiol 2010; 33:291-299 10.1016/j.syapm.2010.08.001 [DOI] [PubMed] [Google Scholar]
- 30.Abt B, Han C, Scheuner C, Lu M, Lapidus A, Nolan M, Lucas S, Hammon N, Deshpande S, Cheng JF, et al. Complete genome sequence of the termite hindgut bacterium Spirochaeta coccoides type strain (SPN1T), reclassification in the genus Sphaerochaeta as Sphaerochaeta coccoides comb. nov. and emendations of the family Spirochaetaceae and the genus Sphaerochaeta. Stand Genomic Sci 2012; 6:194-209 10.4056/sigs.2796069 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Felsenstein J. Inferring phylogenies. Sinauer Associates Inc., Sunderland, Massachusetts, 2004. [Google Scholar]
- 32.Klenk HP, Göker M. En route to a genome-based classification of Archaea and Bacteria? Syst Appl Microbiol 2010; 33:175-182 10.1016/j.syapm.2010.03.003 [DOI] [PubMed] [Google Scholar]
- 33.Wu D, Hugenholtz P, Mavromatis K, Pukall R, Dalin E, Ivanova NN, Kunin V, Goodwin L, Wu M, Tindall BJ, et al. A phylogeny-driven Genomic Encyclopaedia of Bacteria and Archaea. Nature 2009; 462:1056-1060 10.1038/nature08656 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.List of growth media used at DSMZ: http://www.dsmz.de/catalogues/catalogue-microorganisms/culture-technology/list-of-media-for-microorganisms.html
- 35.Gemeinholzer B, Dröge G, Zetzsche H, Haszprunar G, Klenk HP, Güntsch A, Berendsohn WG, Wägele JW. The DNA Bank Network: the start from a German initiative. Biopreserv Biobank 2011; 9:51-55 10.1089/bio.2010.0029 [DOI] [PubMed] [Google Scholar]
- 36.The DOE Joint Genome Institute www.jgi.doe.gov
- 37.Phrap and Phred for Windows. MacOS, Linux, and Unix. www.phrap.com
- 38.Zerbino DR, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res 2008; 18:821-829 10.1101/gr.074492.107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Han C, Chain P. Finishing repeat regions automatically with Dupfinisher. In: Proceeding of the 2006 international conference on bioinformatics & computational biology. Arabnia HR, Valafar H (eds), CSREA Press. June 26-29, 2006: 141-146. [Google Scholar]
- 40.Lapidus A, LaButti K, Foster B, Lowry S, Trong S, Goltsman E. POLISHER: An effective tool for using ultra short reads in microbial genome assembly and finishing. AGBT, Marco Island, FL, 2008. [Google Scholar]
- 41.Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal Prokaryotic Dynamic Programming Genefinding Algorithm. BMC Bioinformatics 2010; 11:119 10.1186/1471-2105-11-119 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pati A, Ivanova N, Mikhailova N, Ovchinikova G, Hooper SD, Lykidis A, Kyrpides NC. GenePRIMP: A Gene Prediction Improvement Pipeline for microbial genomes. Nat Methods 2010; 7:455-457 10.1038/nmeth.1457 [DOI] [PubMed] [Google Scholar]
- 43.Lowe TM, Eddy SR. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res 1997; 25:955-964 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lagesen K, Hallin PF, Rødland E, Stærfeldt HH, Rognes T, Ussery DW. RNAmmer: consistent annotation of rRNA genes in genomic sequences. Nucleic Acids Res 2007; 35:3100-3108 10.1093/nar/gkm160 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Griffiths-Jones S, Bateman A, Marshall M, Khanna A, Eddy SR. Rfam: an RNA family database. Nucleic Acids Res 2003; 31:439-441 10.1093/nar/gkg006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Krogh A, Larsson B, von Heijne G, Sonnhammer ELL. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J Mol Biol 2001; 305:567-580 10.1006/jmbi.2000.4315 [DOI] [PubMed] [Google Scholar]
- 47.Bendtsen JD, Nielsen H, von Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol 2004; 340:783-795 10.1016/j.jmb.2004.05.028 [DOI] [PubMed] [Google Scholar]
- 48.IMG/GEBA img.jgi.doe.gov/geba
- 49.Auch AF, Klenk HP, Göker M. Standard operating procedure for calculating genome-to-genome distances based on high-scoring segment pairs. Stand Genomic Sci 2010; 2:142-148 10.4056/sigs.541628 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Auch AF, von Jan M, Klenk HP, Göker M. Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2010; 2:117-134 10.4056/sigs.531120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Carbohydrate-Active Enzymes Database www.cazy.org
- 52.Cho KH, Salyers AA. Biochemical analysis of interactions between outer membrane proteins that contribute to starch utilization by Bacteroides thetaiotaomicron. J Bacteriol 2001; 183:7224-7230 10.1128/JB.183.24.7224-7230.2001 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Xu J, Bjursell MK, Himrod J, Deng S, Carmichael LK, Chiang HC, Hooper LV, Gordon JI. A genomic view of the human- Bacteroides thetaiotaomicron symbiosis. Science 2003; 299:2074-2076 10.1126/science.1080029 [DOI] [PubMed] [Google Scholar]