Abstract
Pharmacokinetic analysis at the organ level provides insight into how drugs distribute throughout the body but cannot explain how drugs work at the cellular level. Here we demonstrate in vivo single cell pharmacokinetic imaging of PARP-1 inhibitors (PARPi) and model drug behavior under varying conditions. We visualize intracellular kinetics of PARPi distribution in real time, showing that PARPi reaches its cellular target compartment, the nucleus, within minutes in vivo both in cancer and normal cells in various cancer models. We also use these data to validate predictive finite element modeling. Our theoretical and experimental data indicate that tumor cells are exposed to sufficiently high PARPi concentrations in vivo and suggest that drug inefficiency is likely related to proteomic heterogeneity or insensitivity of cancer cells to DNA repair inhibition. This suggests that single cell pharmacokinetic imaging and derived modeling improves our understanding of drug action at single cell resolution in vivo.
Introduction
Drug actions on the human body depend strongly on pharmacokinetics (PK), which describe drug absorption, distribution, and elimination. Current PK models typically assume that drugs partition between organ compartments interacting by mass action transport. Some models include limited molecular and cellular detail, such as receptor occupancy, but in general, models tend to focus on coarse-grain processes, such as a single rate constant for tumor cell growth or cell death. Despite such simplifications, these models have nevertheless played a critical role in the latter stages of drug development and in clinical studies, including information on population pharmacokinetics1. Their coarse grained nature typically precludes their use to understand drug responses in terms of specific signaling pathways or cellular decision-making, and data from PK models are not easily correlated with “omic” data. Perhaps more importantly, existing models tend to neglect heterogeneity in single cell behavior. This can be especially problematic for cancer drugs, as tumors exhibit high cell-to-cell variability in both epigenetic2 and genetic3 properties. Such heterogeneity can arise from different sources4 including mutations, varying protein levels due to stochastic gene expression2, cell cycle status, tumor position, the degree of interaction with neighboring cells, as well as different stem cell characteristics5–7. Moreover, drug effects can also vary due to fluctuations in target molecules3, resistance mechanisms8; interconnected cell signaling networks within cancer cells9, and influences of the local tumor microenvironmental (such as oxygen concentration10, soluble signaling molecules, cell adhesion-mediated interactions11 and external stimuli). Given this high degree of cellular heterogeneity in tumors, PK measurements and models which account for single cell dynamics might be more predictive than classic bulk-tissue approaches. Yet, surprisingly little data exists on how specific drugs actually distribute and work in vivo. Since autoradiography of radiolabeled drugs cannot achieve the subcellular spatial and temporal resolution possible with fluorescence techniques, studying small molecule drug distribution remains difficult12–14.
Recent advances in high-resolution subcellular in vivo imaging methods have enabled more detailed analyses of single cells, and as such, are now being increasingly used to understand how cancer cells behave in in vivo environments15–18. Furthermore, parallel advances in synthetic methods have facilitated the development of small molecule fluorescent drugs with similar affinity and pharmacokinetics to their therapeutic counterparts19,20. One approach to measuring and modeling drug kinetics would thus be to use optimally labeled drugs for high resolution in vivo fluorescence imaging (Fig. 1 and S1). Single cell in vivo pharmacokinetic imaging could then be used to shed new light on several aspects of drug development in that it would allow measurement of intratumor heterogeneity, analysis of drugs that target the tumor microenvironment, mechanistic-driven optimization of drug doses (after appropriate adjustment for drug specific properties), comparison of different closely related compounds, correlation of drug concentrations to effector function, extrapolation of simulations to humans, as well as the performance of detailed failure analyses.
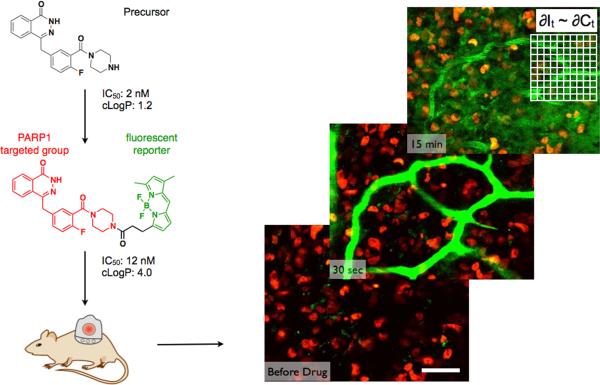
Figure 1. High spatial and temporal resolution microscopy using companion imaging drugs.
Precursor compounds are conjugated with cell permeable, small fluorophores to generate therapeutically active fluorescent companion imaging drugs (See Fig. S1 for structure of parent drug and how piperazine substitutions are tolerated48.) Using a window chamber model, in vivo microscopy enabled the detection of drugs (green) with sub-cellular resolution and frame rates of several seconds. Scale bar = 50 μm in an MDA-MB-231 breast cancer cell line expressing a fluorescent H2B protein in the nucleus (red).
Here we developed new imaging and analyses tools, validated them, and then undertook a model study using in vivo imaging to analyze drug distribution in murine cancer models. Specifically, we developed high resolution, temporal in vivo imaging of single cells in tumors to measure drug distribution, synthesized a fluorescent derivative of a PARP inhibitor that retained activity, and utilized a quantitative framework to extract and extrapolate single cell data to be used in predictive models. We chose PARP inhibitors (PARPi) as an emerging area of solid tumor pharmacology with an exciting rationale of synthetic lethality21, potent and specific compounds, and unexpectedly low success in the clinic for unexplained reasons22,23. We believe that this approach of single-cell pharmacokinetic imaging (SCPKI; Supplementary Fig. S1 could prove valuable in understanding drug action for different drugs and diseases.
Results
Development of imaging technique and PARPi characterization
During single-cell pharmacokinetic imaging (SCPKI), serial images and stacks were obtained in representative areas of the tumor (Fig. 1). Typically, 20–60× objectives were used to discern intracellular detail of ~50–300 cells per field while multiple fields were sampled in z, using objective and stage automation. Several features of the set-up were further optimized for SCPKI. Assuming that the position of single cells changes over several hours of observation, we applied robust immobilization and cell tracking techniques in addition to acquiring image stacks. Time series were initiated prior to intravenous injection, allowing dynamic scanning during partition of drug from circulation into the tumor. This allowed capture of critical early phases of drug distribution. Anesthesia was monitored and adjusted to maintain stable vital signs over a 4–6 hour imaging session. Intravenous fluids were supplied and animals were warmed appropriately to avoid hypothermia18. Fig. 2 and Movie 1 shows a representative example of a raw data set following bolus tail vein injection of the fluorescently tagged PARPi (Fig. 1, Supplementary Fig. S2). The drug filled the functional tumor vasculature within seconds after injection and extravasated within minutes, distributing non-specifically to cells and then washing out to reveal target binding in the nucleus of the vast majority of cells (Fig 3).
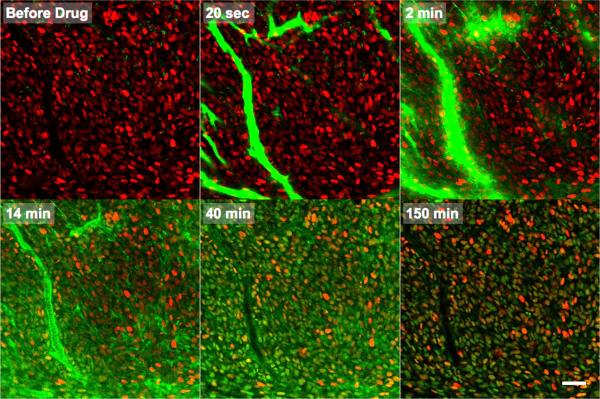
Figure 2. Real-Time In Vivo Drug Distribution of a PARP inhibitor.
Following bolus intravenous administration, the drug (green) perfused the functional tumor vasculature within seconds and extravasated within minutes (top row). The drug initially distributed non-specifically within H2B-RFP (red) expressing cells (primarily within the endoplasmic reticulum, as determined by in vitro cell imaging; Fig. S4, HT-1080 cell line shown). Rapid target binding (within minutes) combined with clearance of non-specific membrane labeling increased the specificity of target versus non-target uptake within an hour. Specific nuclear PARP targeting was observed (bottom row) and this was maintained for several hours. See Movie 1 for dynamic imaging. Scale bar = 50 μm.
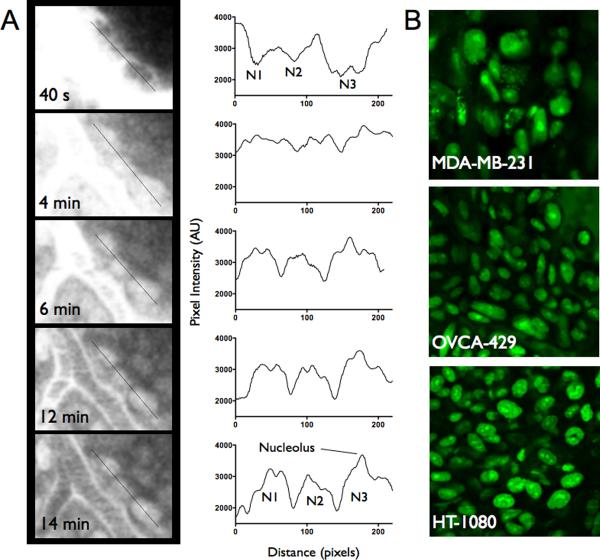
Figure 3. Subcellular spatial resolution of drug distribution.
A. A line profile through three cells adjacent to a tumor vessel shows perinuclear signal within seconds after injection (PARPi fluorescence shown in grayscale). Accumulation in the nucleus and diffusion of free drug deeper into the tumor results in nuclear specific staining with the contrast increasing over time. A high signal in the nucleolus can be observed in these HT-1080 cells with similar staining patterns in multiple cell lines. B. High resolution images of intracellular (primarily nuclear) drug distribution in three different cancer models in vivo. PARPi fluorescence shown in green.
Supplementary Fig. S2 summarizes the synthesis and physicochemical characterization of the PARP imaging agent, based on covalent modification of the olaparib (AZD2281) scaffold with a boron-dipyrrometheneN-hydroxysuccinimide (BODIPY-NHS) ester. The fluorescent companion drug (emission maximum: 525 nm) has a molecular weight of 640 Da and displays excellent characteristics as a targeted fluorescent reporter group, since its affinity is not greatly perturbed by the attachment of a fluorescent BODIPY-FL reporter. The half-maximal inhibitory concentration (IC50) was determined as 12.2±1.1 nM (Supplementary Fig. S3), which is higher than the parent drug but still in the low nanomolar range. Although olaparib has been shown to affect multiple members of the PARP family24, only PARP-1 had significant expression and drug affinity in the model systems used. PARP-2 was absent and the affinity for PARP-3 was too low for a therapeutic effect in vivo (Supplementary Fig. S5). The fluorescent compound freely permeates into cells and co-localizes with PARP immunostaining (Supplementary Figs. S4 and S5) at the single cell level, as well as in different cell lines tested including breast cancer (MDA MB231), ovarian cancer (OVCA429) and others (HT1080, A431). Supplementary Fig. S6 summarizes the cellular kinetics of target accumulation in breast cancer cells in culture where unbound compound was removed from the cells within minutes, resulting in relatively fast intracellular kinetics. To directly correlate the fluorescence signal with drug concentrations, we used phantom data (Supplementary Fig. S3), which we validated in vivo using a built-in calibration (injected dose in vessels; Fig. 2 and Supplementary Fig. S7). Together, these data show that drug concentrations can be estimated inside cells and followed over time; we observed little if any bleaching over typical observation periods and settings. At the whole body level, the imaging drug had a weighted blood half-life of 18 minutes (77% redistribution with a 5 minute half-life and a 23% clearance phase of 60 minutes; Supplementary Fig. S7); competitive inhibition of target binding could be achieved through administration of cold drug (Supplementary Fig. S3D).
Single cell distribution of PAPRi measured in vivo
To determine drug concentrations in vivo in individual tumor cells, we segmented cells in 2D images based on nuclear expression of the fluorescent protein H2B-apple and used a semiautomated algorithm to define associated cell mass (Fig. 4). Fluorescence per cell was then displayed for each individual cell as well as for populations of cells over time. A representative image is shown in Fig. 4 along with the standard deviation between cells as a function of time. Of note, the standard deviation was large immediately after drug administration due to variability in delivery (primarily in proximity to functional blood vessels). Within an hour, however, the heterogeneity decreased and continued to decrease over time as drug diffusion reduced the gradients within the tissue. 3 hours after intravenous administration, the cellular drug concentration was 1.2 ± 0.2 μM, regardless of intratumoral position. At this time point, >99% of the tumor cells are predicted to have therapeutic drug levels based on extrapolation of the dose and measured variability in drug concentrations observed in different tumor cell beds. These cell-population kinetics thus allowed the study of drug distribution to cells within different tumoral regions (i.e. to cells adjacent to vessels, to cells as a function of vascular distance or in relatively avascular regions). It also enabled measurement of the fraction of cells receiving no drug or sub-therapeutic drug levels (see below).
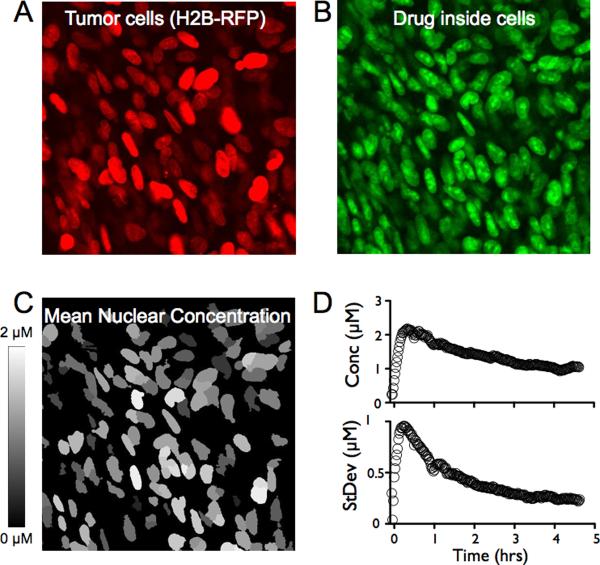
Figure 4. Cell population drug kinetics.
A. H2B-Apple red fluorescent protein was used to identify the nuclei of individual cancer cells in vivo (model HT-1080 cell line shown). B. The drug (green) accumulates inside cells, specifically in the nuclei. C. To measure the average drug concentration inside each cell, nuclei were segmented (based on A) and outlines were overlaid with the companion imaging drug to yield a mean nuclear concentration (C). Note that the average cellular drug concentration in this in vivo example is 1.2 μM. While virtually all cancer cells accumulated the drug (> 99%), there was still some cell-to-cell variability. D. The average concentration rapidly increases following a bolus dose followed by a slow decay (top). Analyzing the standard deviation of 250 cells over time showed the highest deviation at early time points decreasing to a much lower level as the diffusive gradients dissipated (bottom).
To better understand the temporal changes in drug concentrations in individual cells, we performed kinetic measurements in vivo over 4 hours (Fig. 5). Interestingly, we observed some variation in maximum uptake but much less variation in cellular drug concentration at later time points. As shown in the representative kinetograms in Fig. 5, the average drug concentration within a given cell was 1.2 μM after 4 hours. We subsequently analyzed the distribution of drug at early (20 min) and late (90 min) time points in populations of cells. The log normal distributions are plotted for representative mice and tumor types (Fig. 5). At early time points, average cellular drug concentration was primarily related to vessel proximity, whereas at later time points, it appeared to be governed by stochastic distribution. Although cell movement was too great to isolate clonal populations within the tumor (with many cells migrating at 10–20 μm/hour), there was no discernible structure to the distribution. While uptake of this probe correlates with PARP-1 expression in cell culture25, where there are no transport limitations or tumor microenvironmental effects, no correlation existed when digested tumors were analyzed by flow cytometry (data not shown), although loss of drug during preparation could not be completely ruled out. In some of these experiments, we performed repeated bolus injections to determine whether drug concentrations within cells were additive and related in magnitude. The signal increase after the second injection was smaller than after the first injection, likely due to a saturation phenomenon. The heterogeneity between cells, although unexpected, could be the result of variability in target expression, the cell cycle phase (e.g. DNA content), differences in stochastic gene expression, nuclear volume, and/or genetic mutations. Variability in the competitive NAD concentration in the cell could also affect the binding, although the in vitro affinity was measured in the presence of NAD with the current probe. This method could be adopted to measure the concentration of competitive inhibitors such as NAD in vivo.
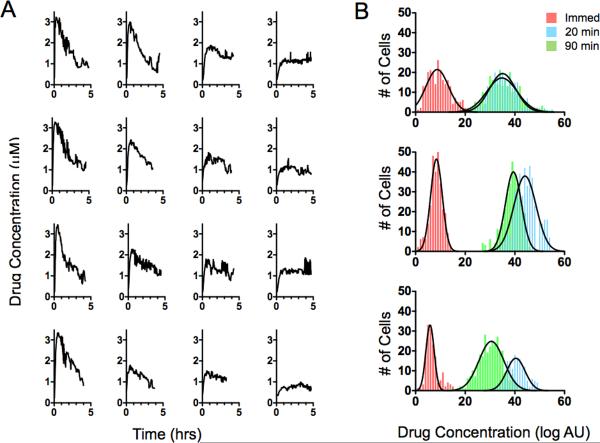
Figure 5. Single cell drug kinetics.
Individual cell profiles can be tracked by temporal single cell imaging in vivo. A. Drug concentrations inside cancer cells were plotted as a function of time following intravenous administration and according to their proximity to microvessels. Profiles in the left column are from representative single cancer cells adjacent to microvessels, profiles in the right column are from cells ~200 μm away from the nearest vessel, and the columns in the middle are from cells at intermediate distances. Note that the intracellular drug concentration in cells is similar at 4 hours after administration. B. A histogram of the log-normal population kinetics over time in different cancer cell types. Top: breast cancer (MDA-MB-231), middle: ovarian cancer (OVCA-429), and bottom: fibrosarcoma (HT-1080). Differences in heterogeneity primarily correspond to changes in vascular density.
The above methods allowed us to address another important question - what is the drug exposure in non-cancerous cells in the tumor? Tumors typically comprise a mixture of cancer cells together with varying amounts of stroma and infiltrating leukocytes. In our models, tumor cells were found to be the predominant cell type (breast cancer model: ~ 80% tumor cells, ~20 % host cells). Yet, in other models such as human cancers and murine K-ras models 26, tumor associated macrophages (TAMs) can be much more prevalent. We found that PARPi distributed non-discriminately to all cells including endothelial and host cells with similar kinetics as for cancer cells. However, the absolute magnitude of cellular accumulation was found to differ, with the highest concentrations accumulating in cancer cells (Fig. 6). This may be in part due to the fact that host cells have lower levels of PARP1 or simply because they are smaller cells.
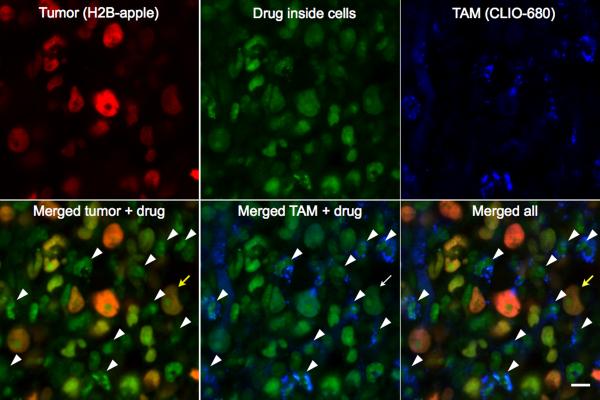
Figure 6. Mapping drug distribution to host cells.
Within a tumor, cancer cells were visualized through expression of H2B-Apple (580 nm, red) and tumor associated macrophages (TAMs) were visualized by a fluorescent nanoparticle (680 nm, blue) internalized into endosomes. The PARPi distribution to cells was visualized at 525 nm (green). The PARPi was seen to accumulate in cancer cells (yellow arrow, HT-1080 cell line) but also in TAMs (as indicated by the white arrowheads). TAM nuclei lack the H2B-apple and are surrounded by CLIO signal. Scale bar = 10 μm
Finite Element Modeling of PARPi Distribution
While the above data was informative in its own right, we hypothesized that quantitative image analysis and modeling could be used to extrapolate findings and investigate biological variables (e.g. vascularity, tumor types, mitotic rates) that would otherwise require extensive biological experiments or simply not be practical (e.g. extrapolation to humans). A major obstacle to developing these refined models has been the lack of experimental data to validate any such model. We developed a two-compartment subcellular (nuclear and cytoplasmic) non-linear partial differential equation model (Fig. 7) to simulate and display major trends in tissue distribution and directly compared modeled data with real experimentally obtained data (Fig. 8). The employed model did not represent single cells, but had unique advantages, including the ability to evaluate and refine the model based on experimental imaging data, and rapidly model in silico the distribution of any arbitrary vascular framework. This approach may be used to account for the stochastic nature of tumor vasculature 27. Once the blood clearance is known, simulation results can be adapted to different delivery routes (i.e. infusion, intraperitoneal, subcutaneous) and species, including humans; insight is also provided on how specific drug properties (permeability, non-specific uptake, diffusion) affect intratumoral distribution. Interestingly, the permeability of the PARPi did not differ statistically between a window chamber (Fig. 2 and Fig. 6), orthotopic mammary fat pad tumor (Supplementary Figs. S8 and S9), or normal vasculature of the ear (Supplementary Fig. S10). While it is known that macromolecular permeability varies depending on tumor location28, these results help validate the previous prediction that small molecule drugs are not limited by permeability29.
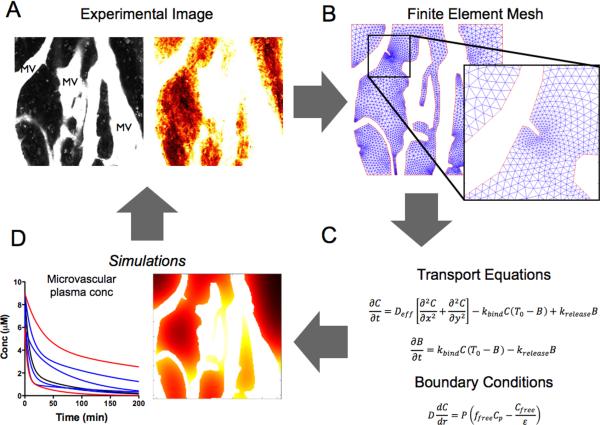
Figure 7. Simulation Development and Validation.
Small molecule drugs distribute to cancer cells from functional tumor microvasculature. A. In the model system, tortuous microvessels were mapped using lectins or vascular probes (top left). B. A finite element mesh was generated around the vessel maps. C. Transport equations and boundary conditions were applied to the simulation. D. Simulation results for time and spatially varying drug concentrations could then be compared to the intravital imaging time series (A) for validation or improvement. These simulations could likewise be extrapolated to different species and used to study the effects of other specific variables. See Supplementary Fig. S6 for specific numerical parameters.
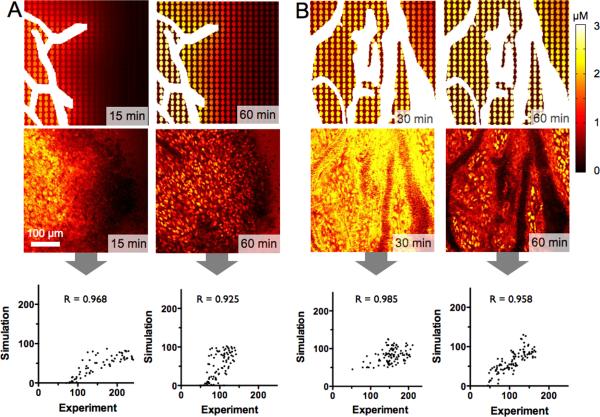
Figure 8. Predictive models of PARPi distribution.
Vessel heterogeneity can be a major determinant of drug distribution particularly early after administration. In these examples (using HT-1080 xenografts with wide variation in vascular density within the same tumor) drug gradients are modeled as a function of distance from microvessels (white); each oval shape represents a model cell. A. A poorly vascularized region shows a large gradient at early times (15 minutes; left) for both the simulation (top) and the corresponding intravital image (bottom). By 1 hour (right), the gradient is significantly reduced and intracellular drug concentrations reach ~1 μM levels. The largeMander correlation coefficient validates the model predictions based on in vitro experimental measurements and systemic clearance. B. High vessel density reduces the spatial gradients, and the cellular accumulation reaches 2–3 μM following administration of the imaging dose. These types of simulations can be used to determine the impact of changing the drug physiochemical properties on distribution for both therapeutics and imaging agents. See Supplementary Fig. S8 for additional modeling including human extrapolation.
We first validated the model by directly comparing outputs to experimentally acquired data. We initially focused on vascular density (Fig. 8) and later expanded the predictions to other variables such as intermittent flow (Supplementary Fig. S11A) and human dosing (Supplementary Fig. S11B). Experimental data showed that poorly vascularized regions showed a strong spatial gradient at 15 minutes that started to dissipate after 1 hour (Fig. 8A). The non-specific tumoral distribution at early time points cleared to reveal nuclear specific staining at later time points. Interestingly there was remarkably good agreement between purely model data and real experimental data. Fig. 8 shows the overall agreement and the calculated Mander's correlation coefficients for individual datasets. It is important to note that the simulations only use the vascular architecture for experimental input; the curves are not fit to the microscopy data. Rather, independent in vitro experiments are used to estimate the driving forces for transient drug distribution (Supplementary Table S1) to allow true predictions of drug uptake in vivo. This also allows data from in vitro experiments on non-fluorescent drug variants to be utilized in the simulations. For regions with much higher vascular density (Fig. 8B), increased delivery resulted in higher drug concentrations and a more homogeneous distribution.
The model was subsequently used to predict the distribution profile around vessels with intermittent blood flow 30. While intermittent blood flow caused transient changes in drug distribution (in addition to a prediction of acute hypoxia) at early time points, the diffusion from surrounding vessels and the time-averaged delivery of the drug by the vessel of interest resulted in a low impact at later time points (Supplementary Fig. S11A). Perhaps one of the most useful aspects of modeling is the ability to extend the data to other species (human) by scaling the plasma clearance rates and adjusting drug specific parameters (molecular weight, oral bioavailability, drug transporter affinity, etc.). As an example, the slower absorption and clearance of drugs after oral delivery in humans 31 resulted in the prediction of a lower drug gradient for this compound in the clinic. This represents one of the most important strengths of developing predictive simulations - the ability to provide insight into situations that are difficult to measure experimentally (e.g. the temporal tissue, cellular, and subcellular distribution of drugs in human tumors). These models can then be used for hypothesis driven research that will either validate or refute the predictions in human tumors when future improvements in technology allow such measurements. Supplementary Fig. S11B shows the maximum predicted gradient following orally administered drug (BID) at steady state compared to the concentration prior to the next dose in a poorly vascularized region. The oral availability was assumed the same as the clinical drug, although the simulation allows arbitrary adjustment of this parameter to investigate the impact of bioavailability on distribution. The relatively quick absorption after oral dosing (< 2 hours) resulted in gradients within the tissue for regions that were poorly vascularized. A small gradient in the opposite direction was found at time points prior to dosing as the drug slowly washed out from the tissue. Notably, cells adjacent to vessels had larger temporal changes in concentration while those further away were buffered from these changes by a depot effect. Overall, these gradients were much lower than in mouse models due to slower systemic absorption and clearance in humans.
Our results indicate that at current clinical doses, virtually all cells in a human tumor (>99% including cancer and non-cancer cells) should attain therapeutic drug concentrations and maintain them over the entire treatment period with this compound. Using the model, generic scenarios in which sub-therapeutic concentrations might occur in clinical settings can be predicted and differentiated, including the following scenarios with the potential to cause drug failure: 1) low systemic doses (e.g. as a result of poor oral availability); 2) rapid clearance or metabolism where trough plasma concentrations are low enough to wash out the drug between doses; 3) non-continuous dosing where the drug fails to reach distant cells before therapy is discontinued (this is more likely when there is significant drug binding to tissue e.g. with drugs targeting DNA, microtubules, or high target density molecules); 4) local metabolism within the tumor tissue, 5) local active transport out of the target region (e.g. when there is high p-glycoprotein expression); 6) changes in effective therapeutic concentrations such as might occur when there is increased expression of competitors (e.g. ATP for kinase inhibitors); or 7) mutations in the binding pocket reducing drug affinity.
Discussion
Our study demonstrates that robust high-resolution profiles of drug distribution over time in single cells in model tumors can be obtained and that by examining multiple cells from the same mouse tumor, inferences about global effects can be made. Moreover, quantitative data can be either modeled to predict cumulative drug effects or extrapolated to human settings to provide insights into drug action previously undetectable by other methods. The findings with PARPi in the present study demonstrate the drug's ubiquitous distribution to virtually all cells in a tumor, including non-cancerous cells, rapid permeation to the nucleus, and sustained binding to target in the nucleus. Given the lower molecular weight and reduced lipophilicity (logP) of the clinical compound, olaparib is predicted to distribute slightly faster than the fluorescent derivative analyzed here, leading to conservative estimates with these predictions. These observations suggest that clinically observed failures with olaparib are less likely attributable to delivery failures. Rather, our findings are more consistent with recent mechanistic studies32,33, which have all indicated that BRCA (but not triple negative breast cancer/TNBC) mutations are required for drug efficacy.
There are a number of different global models describing drug distribution and response; these include population PK models1, physiologically-based pharmacokinetic (PBPK) models34–36, empirical PK/PD models, and mechanistic (e.g. receptor binding) PK/PD models, among others37. While these models have been developed to incorporate delivery limitations or more complex pharmacodynamic (PD) measures, there is currently no method that incorporates the known cell-to-cell variability in cancer phenotypes, the multivariate cell decision inputs, the heterogeneous nature of drug delivery, and the variability of cellular responses into a comprehensive framework. To reconcile the gap between these global models and molecular pathways at the cellular level38,39, we have been working on developing intravital imaging techniques that are capable of providing insight into drug dose-response relationships at the single cell level. Supplementary Fig. S1 summarizes our approach and compares it to global measurements and related modeling methods. A central aspect of the quantitative measurements obtained using the described technique is the development and use of companion imaging drugs that display similar efficacy and in vivo behaviors to the parent drug19. The magnitude and evolution of drug gradients can thus be measured and subsequently modeled using finite element based simulations. These results can also be extrapolated to other species based on plasma concentration time profiles. We examined only two cancer types in depth; both exhibited ubiquitous distribution of the fluorescent PARP inhibitor. Using image quantification and numerical simulations, we found that intracellular drug concentrations will reach sufficiently high levels (above the IC50) in most breast and ovarian cancer cells lines and exceeding that in BRCA−/− models by an order of magnitude (Katy S. Yang, unpublished data) with current preclinical dosing levels. Using a model-based extrapolation of clinical physiological and drug parameters to humans, we infer that the qualified failure of PARP inhibitors in recent trials are likely failures of efficacy rather than failures of delivery22.
Measuring drug concentrations by intravital imaging at the single cell level could be further expanded to other drug classes (e.g. siRNA, antibodies, nanocarriers), other engineered cell and tumor models. First, the experimental window chamber set-up enables rapid analysis and screening of different cancer types; indeed, this set-up is currently being used to evaluate several drugs of interest. Of note, the permeability of this small molecule drug was not statistically different in the window chamber, an orthotopic model, or the normal vasculature of the ear (Supplementary Figs. S8 and S10) as also predicted by transport models29, enabling the use of the window chamber for most experiments with this class of agent. Second, the method allows drug response to be tested in the presence of key mutations (e.g. BRCA status, oncogene mutations). Third, it is possible to conduct mixing experiments where mosaic cells from different backgrounds (encoded by different fluorescent proteins) are studied in parallel to determine differences in drug effects in vivo (e.g. using FUCCI reporters to detect cell cycle effects40, using resistant and sensitive cell lines encoded by different fluorescent proteins, or using clonal cell populations41). Fourth, through the use of reporter genes, it is possible to link kinetic data to PD data. While in the present study we used morphologic nuclear features as a predictor of apoptosis42, fluorescent reporter genes such as 53BP1-XFP43, Ki67-XFP44 or p21-XFP45 could also be used. Fifth, the model enables in vivo synergy and distribution to be determined by using combinations of differently tagged drugs. BODIPY fluorochromes in particular have a relatively small footprint and can be synthesized in a variety of excitation/emission ranges46. Finally, due to advances in optics and objectives47, and through the use of newer immobilization and gating techniques, it has recently become possible to study drug distribution directly in orthotopic cancers without the use of window chambers. Together, these advances will enable the field of single cell PK and PD imaging/modeling to directly connect information on cancer cell signaling networks to therapeutic efficacy.
Methods
Synthesis
Synthesis of the companion imaging drug AZD2281-(BODIPY-Fl) was carried out as described in Supplementary Fig. S2A. Specifically, we reacted a solution of BODIPY-FL succinimidyl ester (5.0 mg, 12.8 μmol; Invitrogen, Carlsbad, CA) in acetonitrile (250 μL), with a solution of 4.7 mg (12.8 μmol, dissolved in 250 μL acetonitrile) of 4-4-fluoro-3-(piperazine-1-carbonyl)phenylmethyl]-2H-phthalazin-1-one48 in the presence triethylamine (4.6 μL, 64.2 μmol). The two components reacted within 4 hours at room temperature. The product AZD2281-BODIPY FL was then isolated using standard high-performance liquid chromatography (HPLC) techniques performed on a Waters (Milford, MA) liquid chromatography–mass spectrometry (LC-MS) system and using a Waters XTerra® C18 5 μm column. The title compound was isolated in 72% yield and its identity confirmed using LC/MS, HPLC, nuclear magnetic resonance (NMR) and high resolution mass spectrometry (HRMS) techniques (Supplementary Fig. S1B–C). The compound had a maximum absorbance at 507 nm and a maximum emission at 527 nm (Supplementary Fig. S2A).
Tumor models
The human breast cancer cell line MDA-MB-231 was purchased from ATCC (Manassas, VA), and OVCA429 and A2780 cell lines were generously provided by Dr. Michael Birrer (Massachusetts General Hospital, Boston, MA). These cell lines were grown in RPMI media supplemented with 10% fetal bovine serum, and 1% penicillin-streptomycin. HT-1080 and A431 cells from ATCC were grown in Dulbecco's Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS) and 1% penicillin-streptomycin.
Reporter cell lines
Histone 2B-red fluorescent protein (H2B-apple) was used to identify the nuclei of individual cancer cells since this model has previously been shown to be robust in vivo42. pmApple-N1 (Myo1E-pmApple-C1, Addgene, Prof. Christien Merrifield49) was cloned by ligating Apple into pmCherry-N1 (Clontech) using AfeI and BsrG1 restriction enzymes. The pTag-H2B-Apple construct was generated by subcloning mApple from pmApple-N1 into pTag-H2B-BFP (Evrogen) using AgeI and NotI. Correct insertion of Apple was confirmed by sequencing the insert in its entirety.
For the MDA-MB-231 and OVCA429 cells, H2B-Apple was subcloned from the pTag H2B-Apple vector into the lentiviral vector pLVX (Clontech) containing H2B-FP using the BamHI and NotI restriction enzymes. Infectious lentiviral particles were produced in Lenti-X 293T cells (Clontech) by transfecting cells with pLVX-H2B-Apple using the Xfect transfection reagent (Clontech). The medium (DMEM, 10% Tet-Approved FBS, 100 I.U. penicillin, 100 μg/ml streptomycin, and 2mM L-glutamine) was replaced 6 hours after transfection. The lentivirus was harvested 72 hours after transfection by collecting and filtering supernatant through a 0.45 μm cellulose acetate filter. Lentiviral particles were concentrated using the Lenti-X concentrator (Clontech) and incubated overnight at 4°C. The supernatant was centrifuged at 1500rpm for 45 minutes at 4°C. To infect MDA-MB-231 and OVCA429 cells, 5 × 104 cells were plated into a 12-well plate and allowed to adhere overnight. Medium was replaced with 1ml complete growth medium (RPMI, 10% FBS, 100 I.U. penicillin, 100 μg/ml streptomycin, 2mM L-glutamine) and 50 μl lentivirus was added to the medium along with 4 μg/ml polybrene. After overnight incubation, the medium was replaced with fresh growth medium. 48 hours post-infection, cells were trypsinized and expanded in a 10 cm dish with 3 μg/ml puromycin for selection.
pTag-H2B-Apple was transfected into HT1080 cells using the X-tremeGENE HP transfection reagent (Roche), followed by selection in 500 μg/ml G418. Using a fluorescent microscope, one clone was selected for fluorescence activated cell sorting, and the brightest 5% of cells were isolated for expansion. Cells were maintained in complete growth medium with 100 μg/ml G418.
Intravital microscopic imaging
All animal experiments were carried out in accordance with guidelines from the Institutional Subcommittee on Research Animal Care. Nude mice (Cox7, Massachusetts General Hospital) were surgically implanted with a dorsal skin window chamber. Approximately 3–4 × 106 cells suspended in 1:1 phosphate buffered saline (PBS) and Matrigel (BD Biosciences, Franklin Lakes, NJ) were implanted under the fascia and allowed to grow for ~2 weeks. When the tumors became vascularized and reached 1–2 mm in size, mice were anesthetized with 2% isoflurane in 2 L/minute oxygen on a heated microscope stage and injected via tail vein catheter with either Angiosense-680 (Perkin Elmer, Waltham, MA) or 250 μg of a 500 kDa amino-dextran labeled with Pacific Blue N-Hydroxysuccinimide (NHS) ester (Invitrogen, Grand Island, NY), according to the manufacturer's instructions. Vascularized regions of interest in the tumor were identified by the vessel probe and H2B-Apple tumor cell signal; regions with minimal out-of-plane vessels were chosen for imaging. Pre-treatment images were collected and time-lapse imaging initiated before injection of the drug via the tail vein catheter. For experiments in which tumor associated macrophages (TAMs) were identified, 20–40 mg/kg of VT680-labeled dextran nanoparticle (CLIO) was injected 24 hours prior to imaging. A time lapse imaging series was initiated, and the AZD2281-BODIDY-Fl was injected during image sequence acquisition. The drug was formulated by dissolving 7.5 μL of a 1 mM solution in dimethyl sulfoxide (DMSO) along with 30 μL of a 1:1 dimethylacetamide (DMAc):solutol solution. 112.5 μl PBS was then slowly added with sonication to obtain a final injection volume of 150 μL.
Static and time series images were collected using a customized Olympus FV1000 confocal microscope (Olympus America). A XLUMPLFLN 20× water immersion objective (NA 1.0) and a 60× LUMFLN (NA 1.10) water immersion objective were used for data collection (both Olympus America). The imaging drug, H2B-Apple, and vascular probes were scanned and excited sequentially using a 405nm, a 473-nm, a 559-nm and/or a 633 nm diode laser, respectively, in combination with a DM405/488/559/635-nm dichroic beam splitter. Emitted light was then separated and collected using appropriate combinations of beam splitters (SDM473, SDM560, and/or SDM 640) and emission filters BA430–455, BA490–540, BA575–620, BA575–675, and/or BA655–755 (all Olympus America). Control tumors were used to determine settings for voltage and laser power and to optimize imaging conditions by ensuring that no photobleaching or phototoxicity occurred.
Mammary fat pad MDA-MB-231 tumors were grown and imaged as described54. MDA-MB-231 cells were injected subcutaneously at the second lowest nipple and grown for 4 weeks. Tumors were exposed for imaging by a medial skin incision. The skin was stretched outward and fixed on a heated cork surface with needles without interfering with blood supply.
Image analysis
Time lapse images were analyzed using Cell Profiler software50. Briefly, cell nuclei were identified by the presence of the fluorescent marker H2B-Apple, which outlined the nucleus. The mean drug signal in the AZD2281-BODIPY-Fl channel was then measured, and the cell trafficking function was used to track cells over time. Any overlapping cells were eliminated from the final data set.
Histology
Five million MDA-MB-231 cells were injected subcutaneously into nude mice and harvested after 4 weeks. The tumors were flash-frozen in O.C.T. compound (Sakura Finetek) with isopentane on dry ice and serial 5 μm thick sections were prepared for immunohistochemistry. Serial sections were stained with PARP-1 (E102: abcam), PARP-2 (A-18: Santa Cruz Biotechnology, Inc.), and PARP-3 (Sigma-Aldrich) followed by biotinylated secondary antibodies (Vector Laboratories. Inc.). The sections were incubated with 0.3% hydrogen peroxide to block endogenous peroxidase activity and blocked with 4% normal goat or rabbit serum to reduce non-specific background. ABC kit (Vector Laboratories, Inc.) and AEC substrate (Dako) were used for the color development, and all the sections were counterstained with Harris hematoxylin (Sigma-Aldrich). The slides were scanned and digitally processed by Nanozoomer 2.0RS (Hamamatsu, Japan).
Mathematical model
To model drug distribution around tumor blood vessels, a finite element method (FEM) was implemented using Matlab (Natick, MA). An image of the vascular probe was used to define a two-dimensional layout of microvessels in the field of view. Following automatic mesh generation using a Delaunay triangulation algorithm, the appropriate transport equations and boundary conditions were specified. A mixed (Robin) boundary condition was applied to all vessels, with the plasma concentration set to the experimentally measured biexponential decay. With high plasma protein binding (> 99% bound), the concentration of drug is not depleted along the length of the vessel29. The boundary conditions were therefore identical throughout the image. No-flux conditions were specified at the image edge. A two-compartment pseudo-homogeneous tissue model was used, where non-specific uptake was assumed to be instantaneous (thereby reducing the effective diffusion coefficient)51. Binding to the nuclear target immobilized the drug. Binding and dissociation rates were fit to experimental kinetic rates measured in cell culture and in vitro affinity measurements (Supplementary Fig. S4). Other parameters are listed in Supplementary Fig. S7. An explicit time step algorithm was implemented, where the step size was decreased until no over-shoot in bound drug concentration occurred. To visualize the results, a nuclear `mask' was employed where the nuclear concentration was shown within the nuclear region and the non-specific drug concentration was shown outside. This was done for visualization only since both compartments were simulated in a pseudo-homogeneous tissue.
Supplementary Material
Acknowledgments
We would like to thank Yoshiko Iwamoto for expert histopathological analysis, Dr. Claudio Vinegoni for intravital imaging, Dr. Michael Birrer for cell lines, Dr. GaudenzDanuser for many helpful discussions, Drs. Mikael Pittet, John Higgins, Yvonna Fisher-Jeffes, Filip Swirski and Matthias Nahrendorf for critical review. The work was funded in parts by 1R01CA164448 (RW, PS, TM), PO1CA139980 (TM, PS, RW) and P50CA086355 (RW). GT was supported by K01DK093766.
Footnotes
Author contributions G.M.T., K.S.Y., P.S., T.M., and R.W. developed the concepts and designed the experiments, G.M.T., K.S.Y., T.R., and R.H.K. performed the experiments, G.M.T., K.S.Y., P.S., T.M., and R.W. analyzed the results and wrote the paper.
Competing financial interests The authors declare no competing financial interests.
References
- 1.Sun H, et al. Population pharmacokinetics. A regulatory perspective. Clin Pharmacokinet. 1999;37:41–58. doi: 10.2165/00003088-199937010-00003. [DOI] [PubMed] [Google Scholar]
- 2.Spencer SL, Gaudet S, Albeck JG, Burke JM, Sorger PK. Non-genetic origins of cell-to-cell variability in TRAIL-induced apoptosis. Nature. 2009;459:428–432. doi: 10.1038/nature08012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Gerlinger M, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012;366:883–892. doi: 10.1056/NEJMoa1113205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Niepel M, Spencer SL, Sorger PK. Non-genetic cell-to-cell variability and the consequences for pharmacology. Curr Opin Chem Biol. 2009;13:556–561. doi: 10.1016/j.cbpa.2009.09.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Roesch A, et al. A temporarily distinct subpopulation of slow-cycling melanoma cells is required for continuous tumor growth. Cell. 2010;141:583–594. doi: 10.1016/j.cell.2010.04.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hemmings C. The elaboration of a critical framework for understanding cancer: the cancer stem cell hypothesis. Pathology. 2010;42:105–112. doi: 10.3109/00313020903488773. [DOI] [PubMed] [Google Scholar]
- 7.Besancon R, Valsesia-Wittmann S, Puisieux A, Caron de Fromentel C, Maguer-Satta V. Cancer stem cells: the emerging challenge of drug targeting. Curr Med Chem. 2009;16:394–416. doi: 10.2174/092986709787315531. [DOI] [PubMed] [Google Scholar]
- 8.Zander SA, et al. Sensitivity and acquired resistance of BRCA1;p53-deficient mouse mammary tumors to the topoisomerase I inhibitor topotecan. Cancer Res. 2010;70:1700–1710. doi: 10.1158/0008-5472.CAN-09-3367. [DOI] [PubMed] [Google Scholar]
- 9.Joslin EJ, et al. Structure of the EGF receptor transactivation circuit integrates multiple signals with cell context. Mol Biosyst. 2010;6:1293–1306. doi: 10.1039/c003921g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Palmer GM, et al. Optical imaging of tumor hypoxia dynamics. J Biomed Opt. 2010;15:066021. doi: 10.1117/1.3523363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Cavallaro U, Christofori G. Cell adhesion and signalling by cadherins and Ig-CAMs in cancer. Nat Rev Cancer. 2004;4:118–132. doi: 10.1038/nrc1276. [DOI] [PubMed] [Google Scholar]
- 12.Kyle AH, Huxham LA, Yeoman DM, Minchinton AI. Limited tissue penetration of taxanes: a mechanism for resistance in solid tumors. Clin Cancer Res. 2007;13:2804–2810. doi: 10.1158/1078-0432.CCR-06-1941. [DOI] [PubMed] [Google Scholar]
- 13.McKillop D, et al. Tumor penetration of gefitinib (Iressa), an epidermal growth factor receptor tyrosine kinase inhibitor. Mol Cancer Ther. 2005;4:641–649. doi: 10.1158/1535-7163.MCT-04-0329. [DOI] [PubMed] [Google Scholar]
- 14.Doubrovin M, et al. (124)I-iodopyridopyrimidinone for PET of Abl kinase-expressing tumors in vivo. J Nucl Med. 2010;51:121–129. doi: 10.2967/jnumed.109.066126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kedrin D, et al. Intravital imaging of metastatic behavior through a mammary imaging window. Nat Methods. 2008;5:1019–1021. doi: 10.1038/nmeth.1269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Vakoc BJ, Fukumura D, Jain RK, Bouma BE. Cancer imaging by optical coherence tomography: preclinical progress and clinical potential. Nat Rev Cancer. 2012;12:363–368. doi: 10.1038/nrc3235. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Fukumura D, Duda DG, Munn LL, Jain RK. Tumor microvasculature and microenvironment: novel insights through intravital imaging in pre-clinical models. Microcirculation. 2010;17:206–225. doi: 10.1111/j.1549-8719.2010.00029.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Pittet MJ, Weissleder R. Intravital imaging. Cell. 2011;147:983–991. doi: 10.1016/j.cell.2011.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Budin G, Yang KS, Reiner T, Weissleder R. Bioorthogonal probes for polo-like kinase 1 imaging and quantification. Angew Chem Int Ed Engl. 2011;50:9378–9381. doi: 10.1002/anie.201103273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Yang KS, Budin G, Reiner T, Vinegoni C, Weissleder R. Bioorthogonal imaging of aurora kinase A in live cells. Angew Chem Int Ed Engl. 2012;51:6598–6603. doi: 10.1002/anie.201200994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rouleau M, Patel A, Hendzel MJ, Kaufmann SH, Poirier GG. PARP inhibition: PARP1 and beyond. Nat Rev Cancer. 2010;10:293–301. doi: 10.1038/nrc2812. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kaye SB, et al. Phase II, open-label, randomized, multicenter study comparing the efficacy and safety of olaparib, a poly (ADP-ribose) polymerase inhibitor, and pegylated liposomal doxorubicin in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer. J Clin Oncol. 2012;30:372–379. doi: 10.1200/JCO.2011.36.9215. [DOI] [PubMed] [Google Scholar]
- 23.Gelmon KA, et al. Olaparib in patients with recurrent high-grade serous or poorly differentiated ovarian carcinoma or triple-negative breast cancer: a phase 2, multicentre, open-label, non-randomised study. Lancet Oncol. 2011;12:852–861. doi: 10.1016/S1470-2045(11)70214-5. [DOI] [PubMed] [Google Scholar]
- 24.Wahlberg E, et al. Family-wide chemical profiling and structural analysis of PARP and tankyrase inhibitors. Nat Biotechnol. 2012;30:283–288. doi: 10.1038/nbt.2121. [DOI] [PubMed] [Google Scholar]
- 25.Reiner T, et al. Imaging therapeutic PARP inhibition in vivo through bioorthogonally developed companion imaging agents. Neoplasia. 2012;14:169–177. doi: 10.1593/neo.12414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Grimm J, et al. Use of gene expression profiling to direct in vivo molecular imaging of lung cancer. Proc Natl Acad Sci U S A. 2005;102:14404–14409. doi: 10.1073/pnas.0503920102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pries AR, et al. Structural adaptation and heterogeneity of normal and tumor microvascular networks. PLoS Comput Biol. 2009;5:e1000394. doi: 10.1371/journal.pcbi.1000394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yuan F, et al. Vascular permeability and microcirculation of gliomas and mammary carcinomas transplanted in rat and mouse cranial windows. Cancer Res. 1994;54:4564–4568. [PubMed] [Google Scholar]
- 29.Thurber GM, Weissleder R. A systems approach for tumor pharmacokinetics. PLoS One. 2011;6:e24696. doi: 10.1371/journal.pone.0024696. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Minchinton AI, Durand RE, Chaplin DJ. Intermittent blood flow in the KHT sarcoma--flow cytometry studies using Hoechst 33342. Br J Cancer. 1990;62:195–200. doi: 10.1038/bjc.1990.259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Yamamoto N, et al. A Phase I, dose-finding and pharmacokinetic study of olaparib (AZD2281) in Japanese patients with advanced solid tumors*,dagger. Cancer Sci. 2012;103:504–509. doi: 10.1111/j.1349-7006.2011.02179.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kortmann U, et al. Tumor growth inhibition by olaparib in BRCA2 germline-mutated patient-derived ovarian cancer tissue xenografts. Clin Cancer Res. 2011;17:783–791. doi: 10.1158/1078-0432.CCR-10-1382. [DOI] [PubMed] [Google Scholar]
- 33.Brenner JC, et al. Mechanistic rationale for inhibition of poly(ADP-ribose) polymerase in ETS gene fusion-positive prostate cancer. Cancer Cell. 2011;19:664–678. doi: 10.1016/j.ccr.2011.04.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Bischoff KB, Dedrick RL. Thiopental pharmacokinetics. J Pharm Sci. 1968;57:1346–1351. doi: 10.1002/jps.2600570814. [DOI] [PubMed] [Google Scholar]
- 35.Baxter LT, Zhu H, Mackensen DG, Butler WF, Jain RK. Biodistribution of monoclonal antibodies: scale-up from mouse to human using a physiologically based pharmacokinetic model. Cancer Res. 1995;55:4611–4622. [PubMed] [Google Scholar]
- 36.Ferl GZ, Kenanova V, Wu AM, DiStefano J. J. r. A two-tiered physiologically based model for dually labeled single-chain Fv-Fc antibody fragments. Mol Cancer Ther. 2006;5:1550–1558. doi: 10.1158/1535-7163.MCT-06-0072. [DOI] [PubMed] [Google Scholar]
- 37.Mager DE, Wyska E, Jusko WJ. Diversity of mechanism-based pharmacodynamic models. Drug Metab Dispos. 2003;31:510–518. doi: 10.1124/dmd.31.5.510. [DOI] [PubMed] [Google Scholar]
- 38.Kleiman LB, Maiwald T, Conzelmann H, Lauffenburger DA, Sorger PK. Rapid phospho-turnover by receptor tyrosine kinases impacts downstream signaling and drug binding. Mol Cell. 2011;43:723–737. doi: 10.1016/j.molcel.2011.07.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Ventura AC, Jackson TL, Merajver SD. On the role of cell signaling models in cancer research. Cancer Res. 2009;69:400–402. doi: 10.1158/0008-5472.CAN-08-4422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sakaue-Sawano A, et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell. 2008;132:487–498. doi: 10.1016/j.cell.2007.12.033. [DOI] [PubMed] [Google Scholar]
- 41.Weber K, Thomaschewski M, Benten D, Fehse B. RGB marking with lentiviral vectors for multicolor clonal cell tracking. Nat Protoc. 2012;7:839–849. doi: 10.1038/nprot.2012.026. [DOI] [PubMed] [Google Scholar]
- 42.Orth JD, et al. Analysis of mitosis and antimitotic drug responses in tumors by in vivo microscopy and single-cell pharmacodynamics. Cancer Res. 2011;71:4608–4616. doi: 10.1158/0008-5472.CAN-11-0412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Bekker-Jensen S, Lukas C, Melander F, Bartek J, Lukas J. Dynamic assembly and sustained retention of 53BP1 at the sites of DNA damage are controlled by Mdc1/NFBD1. J Cell Biol. 2005;170:201–211. doi: 10.1083/jcb.200503043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Zambon AC. Use of the Ki67 promoter to label cell cycle entry in living cells. Cytometry A. 2010;77:564–570. doi: 10.1002/cyto.a.20890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Loewer A, Batchelor E, Gaglia G, Lahav G. Basal dynamics of p53 reveal transcriptionally attenuated pulses in cycling cells. Cell. 2010;142:89–100. doi: 10.1016/j.cell.2010.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Descalzo AB, Xu HJ, Shen Z, Rurack K. Red/near-infrared boron-dipyrromethene dyes as strongly emitting fluorophores. Ann N Y Acad Sci. 2008;1130:164–171. doi: 10.1196/annals.1430.016. [DOI] [PubMed] [Google Scholar]
- 47.Alencar H, Mahmood U, Kawano Y, Hirata T, Weissleder R. Novel multiwavelength microscopic scanner for mouse imaging. Neoplasia. 2005;7:977–983. doi: 10.1593/neo.05376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Menear KA, et al. 4-[3-(4-cyclopropanecarbonylpiperazine-1-carbonyl)-4-fluorobenzyl]-2H-phthalazin- 1-one: a novel bioavailable inhibitor of poly(ADP-ribose) polymerase-1. J Med Chem. 2008;51:6581–6591. doi: 10.1021/jm8001263. [DOI] [PubMed] [Google Scholar]
- 49.Taylor MJ, Perrais D, Merrifield CJ. A high precision survey of the molecular dynamics of mammalian clathrin-mediated endocytosis. PLoS Biol. 2011;9:e1000604. doi: 10.1371/journal.pbio.1000604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Lamprecht MR, Sabatini DM, Carpenter AE. CellProfiler: free, versatile software for automated biological image analysis. Biotechniques. 2007;42:71–75. doi: 10.2144/000112257. [DOI] [PubMed] [Google Scholar]
- 51.Crank J. The mathematics of diffusion. Clarendon Press; Oxford, [Eng]: 1975. [Google Scholar]
- 52.Gerlowski LE, Jain RK. Microvascular permeability of ormal and neoplastic tissues. Microvasc Res. 1986;31:288–305. doi: 10.1016/0026-2862(86)90018-x. [DOI] [PubMed] [Google Scholar]
- 53.Smith DA, Di L, Kerns EH. The effect of plasma protein binding on in vivo efficacy: misconceptions in drug discovery. Nat Rev Drug Discov. 2010;9:929–939. doi: 10.1038/nrd3287. [DOI] [PubMed] [Google Scholar]
- 54.Kedrin D, Wyckoff J, Sahai E, Condeelis J, Segall JE. Imaging tumor cell movement in vivo. Curr Protoc Cell Biol. 2007;Chapter 19(Unit 19.7) doi: 10.1002/0471143030.cb1907s35. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.