Abstract
After each round of replication, new transcription initiation complexes must assemble on promoter DNA. This process may compete with packaging of the same promoter sequences into nucleosomes. To elucidate interactions between regulatory transcription factors and nucleosomes on newly replicated DNA, we asked whether heat shock factor (HSF) could be made to bind to nucleosomal DNA in vivo. A heat shock element (HSE) was embedded at either of two different sites within a DNA segment that directs the formation of a stable, positioned nucleosome. The resulting DNA segments were coupled to a reporter gene and transfected into the yeast Saccharomyces cerevisiae. Transcription from these two plasmid constructions after induction by heat shock was similar in amount to that from a control plasmid in which HSF binds to nucleosome-free DNA. High-resolution genomic footprint mapping of DNase I and micrococcal nuclease cleavage sites indicated that the HSE in these two plasmids was, nevertheless, packaged in a nucleosome. The inclusion of HSE sequences within (but relatively close to the edge of) the nucleosome did not alter the position of the nucleosome which formed with the parental DNA fragment. Genomic footprint analyses also suggested that the HSE-containing nucleosome was unchanged by the induction of transcription. Quantitative comparisons with control plasmids ruled out the possibility that HSF was bound only to a small fraction of molecules that might have escaped nucleosome assembly. Analysis of the helical orientation of HSE DNA in the nucleosome indicated that HSF contacted DNA residues that faced outward from the histone octamer. We discuss the significance of these results with regard to the role of nucleosomes in inhibiting transcription and the normal occurrence of nucleosome-free regions in promoters.
Full text
PDF

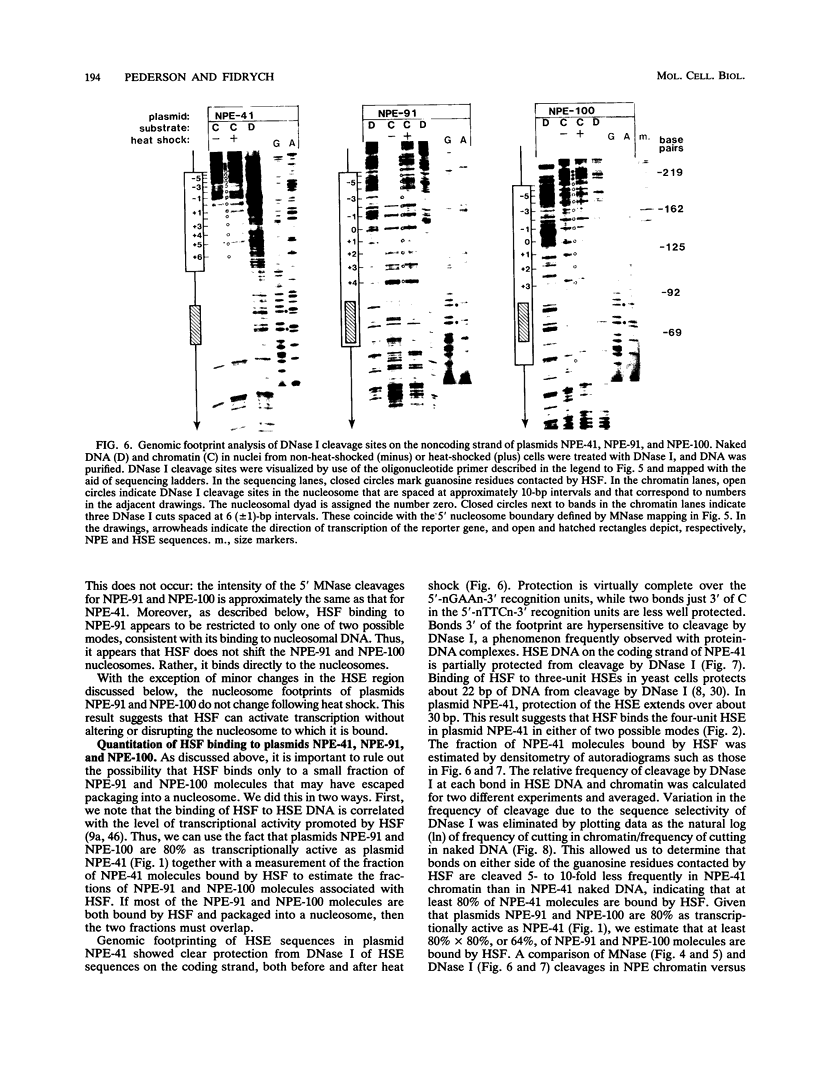
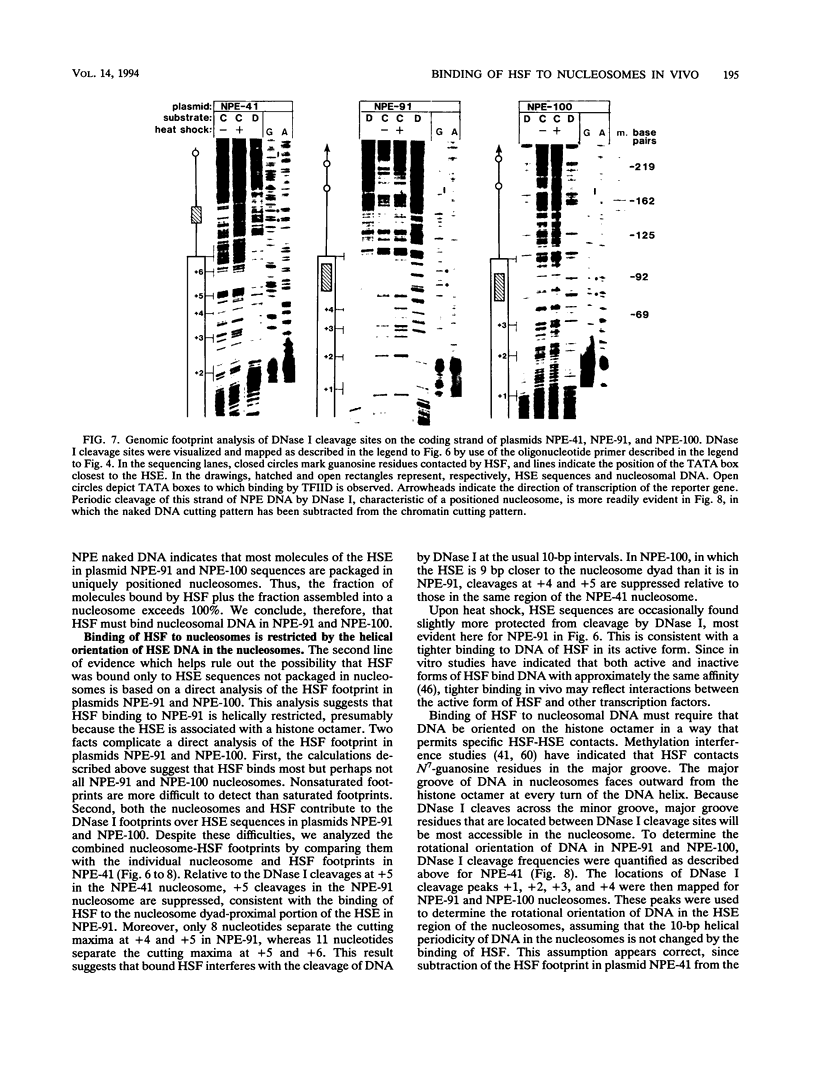
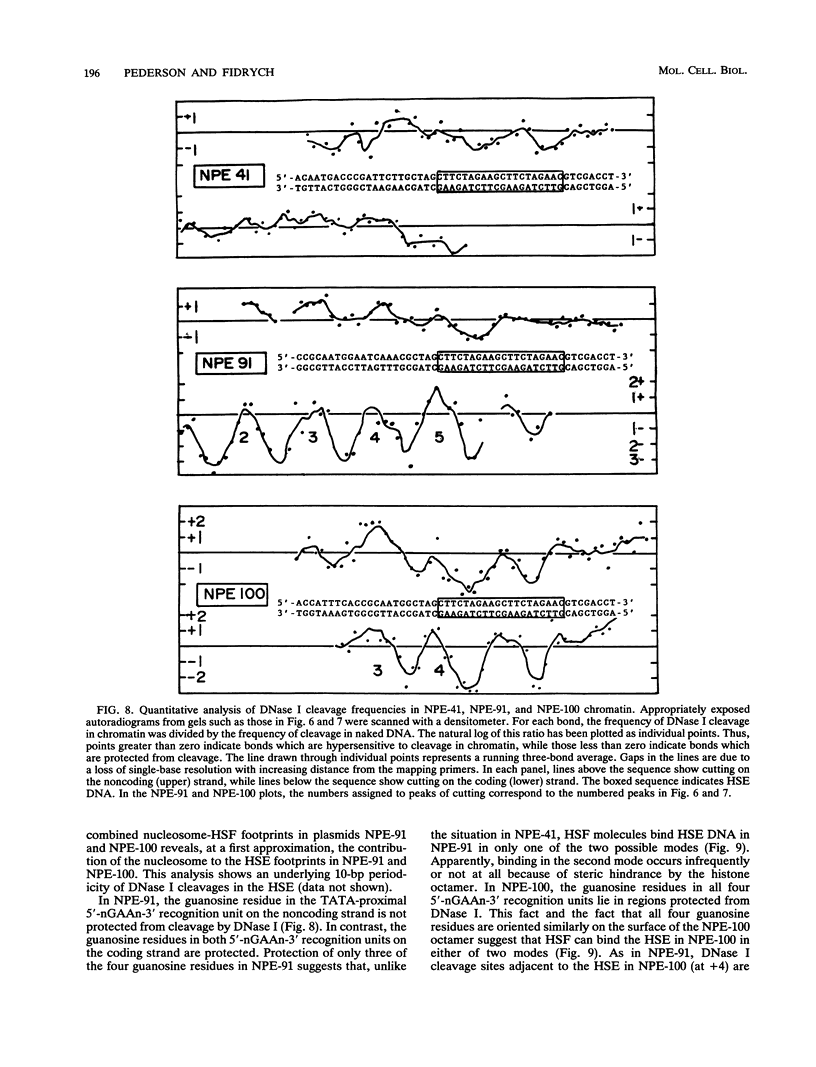
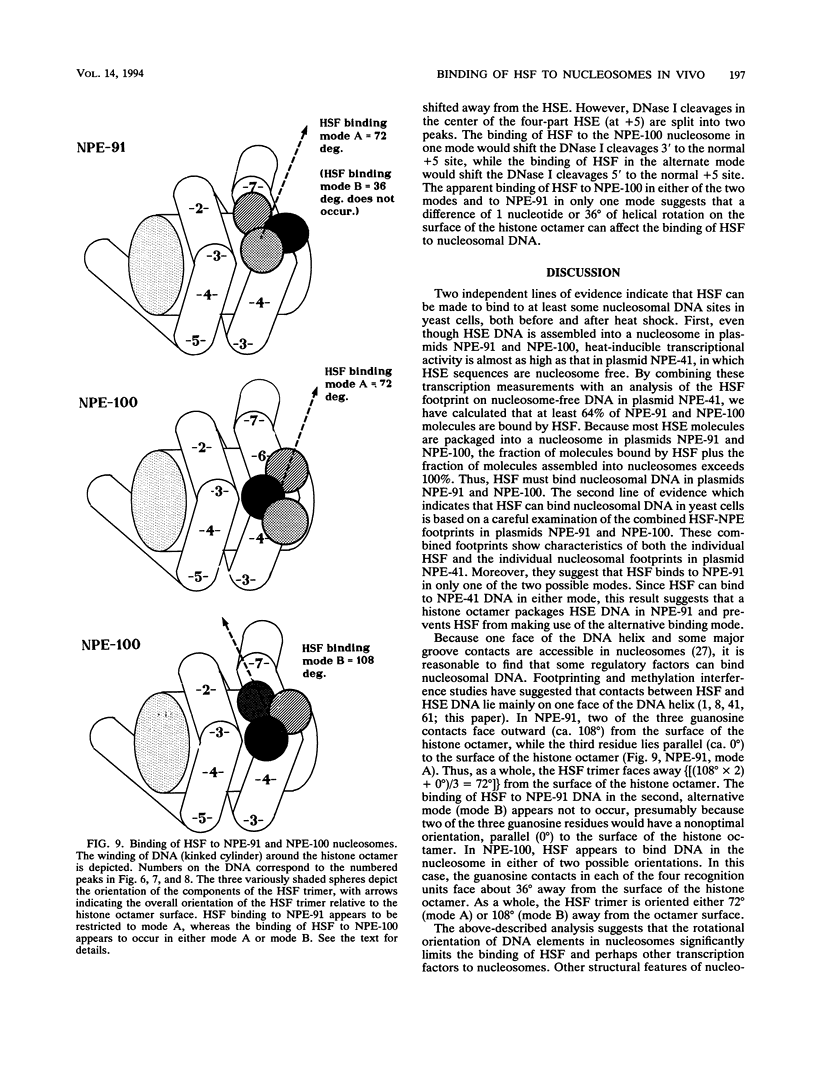


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- ALEXANDER M., HEPPEL L. A., HURWITZ J. The purification and properties of micrococcal nuclease. J Biol Chem. 1961 Nov;236:3014–3019. [PubMed] [Google Scholar]
- Abravaya K., Phillips B., Morimoto R. I. Heat shock-induced interactions of heat shock transcription factor and the human hsp70 promoter examined by in vivo footprinting. Mol Cell Biol. 1991 Jan;11(1):586–592. doi: 10.1128/mcb.11.1.586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Almouzni G., Clark D. J., Méchali M., Wolffe A. P. Chromatin assembly on replicating DNA in vitro. Nucleic Acids Res. 1990 Oct 11;18(19):5767–5774. doi: 10.1093/nar/18.19.5767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amin J., Ananthan J., Voellmy R. Key features of heat shock regulatory elements. Mol Cell Biol. 1988 Sep;8(9):3761–3769. doi: 10.1128/mcb.8.9.3761. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Archer T. K., Cordingley M. G., Wolford R. G., Hager G. L. Transcription factor access is mediated by accurately positioned nucleosomes on the mouse mammary tumor virus promoter. Mol Cell Biol. 1991 Feb;11(2):688–698. doi: 10.1128/mcb.11.2.688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker P. B., Rabindran S. K., Wu C. Heat shock-regulated transcription in vitro from a reconstituted chromatin template. Proc Natl Acad Sci U S A. 1991 May 15;88(10):4109–4113. doi: 10.1073/pnas.88.10.4109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown D. D. The role of stable complexes that repress and activate eucaryotic genes. Cell. 1984 Jun;37(2):359–365. doi: 10.1016/0092-8674(84)90366-0. [DOI] [PubMed] [Google Scholar]
- Chen J., Pederson D. S. A distal heat shock element promotes the rapid response to heat shock of the HSP26 gene in the yeast Saccharomyces cerevisiae. J Biol Chem. 1993 Apr 5;268(10):7442–7448. [PubMed] [Google Scholar]
- Cockell M., Rhodes D., Klug A. Location of the primary sites of micrococcal nuclease cleavage on the nucleosome core. J Mol Biol. 1983 Oct 25;170(2):423–446. doi: 10.1016/s0022-2836(83)80156-9. [DOI] [PubMed] [Google Scholar]
- Cunniff N. F., Morgan W. D. Analysis of heat shock element recognition by saturation mutagenesis of the human HSP70.1 gene promoter. J Biol Chem. 1993 Apr 15;268(11):8317–8324. [PubMed] [Google Scholar]
- Drew H. R. Structural specificities of five commonly used DNA nucleases. J Mol Biol. 1984 Jul 15;176(4):535–557. doi: 10.1016/0022-2836(84)90176-1. [DOI] [PubMed] [Google Scholar]
- Drew H. R., Travers A. A. DNA bending and its relation to nucleosome positioning. J Mol Biol. 1985 Dec 20;186(4):773–790. doi: 10.1016/0022-2836(85)90396-1. [DOI] [PubMed] [Google Scholar]
- Elgin S. C. The formation and function of DNase I hypersensitive sites in the process of gene activation. J Biol Chem. 1988 Dec 25;263(36):19259–19262. [PubMed] [Google Scholar]
- Felsenfeld G. Chromatin as an essential part of the transcriptional mechanism. Nature. 1992 Jan 16;355(6357):219–224. doi: 10.1038/355219a0. [DOI] [PubMed] [Google Scholar]
- Gill G., Ptashne M. Mutants of GAL4 protein altered in an activation function. Cell. 1987 Oct 9;51(1):121–126. doi: 10.1016/0092-8674(87)90016-x. [DOI] [PubMed] [Google Scholar]
- Gross D. S., English K. E., Collins K. W., Lee S. W. Genomic footprinting of the yeast HSP82 promoter reveals marked distortion of the DNA helix and constitutive occupancy of heat shock and TATA elements. J Mol Biol. 1990 Dec 5;216(3):611–631. doi: 10.1016/0022-2836(90)90387-2. [DOI] [PubMed] [Google Scholar]
- Gross D. S., Garrard W. T. Nuclease hypersensitive sites in chromatin. Annu Rev Biochem. 1988;57:159–197. doi: 10.1146/annurev.bi.57.070188.001111. [DOI] [PubMed] [Google Scholar]
- Grunstein M. Histone function in transcription. Annu Rev Cell Biol. 1990;6:643–678. doi: 10.1146/annurev.cb.06.110190.003235. [DOI] [PubMed] [Google Scholar]
- Guarente L., Ptashne M. Fusion of Escherichia coli lacZ to the cytochrome c gene of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1981 Apr;78(4):2199–2203. doi: 10.1073/pnas.78.4.2199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Tullius T. D., Wolffe A. P. The structure of DNA in a nucleosome. Proc Natl Acad Sci U S A. 1990 Oct;87(19):7405–7409. doi: 10.1073/pnas.87.19.7405. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayes J. J., Wolffe A. P. The interaction of transcription factors with nucleosomal DNA. Bioessays. 1992 Sep;14(9):597–603. doi: 10.1002/bies.950140905. [DOI] [PubMed] [Google Scholar]
- Jakobsen B. K., Pelham H. R. Constitutive binding of yeast heat shock factor to DNA in vivo. Mol Cell Biol. 1988 Nov;8(11):5040–5042. doi: 10.1128/mcb.8.11.5040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornberg R. D., Lorch Y. Chromatin structure and transcription. Annu Rev Cell Biol. 1992;8:563–587. doi: 10.1146/annurev.cb.08.110192.003023. [DOI] [PubMed] [Google Scholar]
- Kornberg R. D., Lorch Y. Irresistible force meets immovable object: transcription and the nucleosome. Cell. 1991 Nov 29;67(5):833–836. doi: 10.1016/0092-8674(91)90354-2. [DOI] [PubMed] [Google Scholar]
- Laskey R. A., Leno G. H. Assembly of the cell nucleus. Trends Genet. 1990 Dec;6(12):406–410. doi: 10.1016/0168-9525(90)90301-l. [DOI] [PubMed] [Google Scholar]
- Lee D. Y., Hayes J. J., Pruss D., Wolffe A. P. A positive role for histone acetylation in transcription factor access to nucleosomal DNA. Cell. 1993 Jan 15;72(1):73–84. doi: 10.1016/0092-8674(93)90051-q. [DOI] [PubMed] [Google Scholar]
- Li W. Z., Sherman F. Two types of TATA elements for the CYC1 gene of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1991 Feb;11(2):666–676. doi: 10.1128/mcb.11.2.666. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McGhee J. D., Felsenfeld G. Reaction of nucleosome DNA with dimethyl sulfate. Proc Natl Acad Sci U S A. 1979 May;76(5):2133–2137. doi: 10.1073/pnas.76.5.2133. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morse R. H., Pederson D. S., Dean A., Simpson R. T. Yeast nucleosomes allow thermal untwisting of DNA. Nucleic Acids Res. 1987 Dec 23;15(24):10311–10330. doi: 10.1093/nar/15.24.10311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morse R. H., Roth S. Y., Simpson R. T. A transcriptionally active tRNA gene interferes with nucleosome positioning in vivo. Mol Cell Biol. 1992 Sep;12(9):4015–4025. doi: 10.1128/mcb.12.9.4015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pederson D. S., Morse R. H. Effect of transcription of yeast chromatin on DNA topology in vivo. EMBO J. 1990 Jun;9(6):1873–1881. doi: 10.1002/j.1460-2075.1990.tb08313.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pederson D. S., Thoma F., Simpson R. T. Core particle, fiber, and transcriptionally active chromatin structure. Annu Rev Cell Biol. 1986;2:117–147. doi: 10.1146/annurev.cb.02.110186.001001. [DOI] [PubMed] [Google Scholar]
- Pederson D. S., Venkatesan M., Thoma F., Simpson R. T. Isolation of an episomal yeast gene and replication origin as chromatin. Proc Natl Acad Sci U S A. 1986 Oct;83(19):7206–7210. doi: 10.1073/pnas.83.19.7206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perisic O., Xiao H., Lis J. T. Stable binding of Drosophila heat shock factor to head-to-head and tail-to-tail repeats of a conserved 5 bp recognition unit. Cell. 1989 Dec 1;59(5):797–806. doi: 10.1016/0092-8674(89)90603-x. [DOI] [PubMed] [Google Scholar]
- Perlmann T., Wrange O. Specific glucocorticoid receptor binding to DNA reconstituted in a nucleosome. EMBO J. 1988 Oct;7(10):3073–3079. doi: 10.1002/j.1460-2075.1988.tb03172.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piña B., Brüggemeier U., Beato M. Nucleosome positioning modulates accessibility of regulatory proteins to the mouse mammary tumor virus promoter. Cell. 1990 Mar 9;60(5):719–731. doi: 10.1016/0092-8674(90)90087-u. [DOI] [PubMed] [Google Scholar]
- Ramsay N., Felsenfeld G., Rushton B. M., McGhee J. D. A 145-base pair DNA sequence that positions itself precisely and asymmetrically on the nucleosome core. EMBO J. 1984 Nov;3(11):2605–2611. doi: 10.1002/j.1460-2075.1984.tb02181.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richmond T. J., Finch J. T., Rushton B., Rhodes D., Klug A. Structure of the nucleosome core particle at 7 A resolution. Nature. 1984 Oct 11;311(5986):532–537. doi: 10.1038/311532a0. [DOI] [PubMed] [Google Scholar]
- Saavedra R. A., Huberman J. A. Both DNA topoisomerases I and II relax 2 micron plasmid DNA in living yeast cells. Cell. 1986 Apr 11;45(1):65–70. doi: 10.1016/0092-8674(86)90538-6. [DOI] [PubMed] [Google Scholar]
- Shuey D. J., Parker C. S. Binding of Drosophila heat-shock gene transcription factor to the hsp 70 promoter. Evidence for symmetric and dynamic interactions. J Biol Chem. 1986 Jun 15;261(17):7934–7940. [PubMed] [Google Scholar]
- Simpson R. T. Nucleosome positioning can affect the function of a cis-acting DNA element in vivo. Nature. 1990 Jan 25;343(6256):387–389. doi: 10.1038/343387a0. [DOI] [PubMed] [Google Scholar]
- Simpson R. T., Stafford D. W. Structural features of a phased nucleosome core particle. Proc Natl Acad Sci U S A. 1983 Jan;80(1):51–55. doi: 10.1073/pnas.80.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slater M. R., Craig E. A. Transcriptional regulation of an hsp70 heat shock gene in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1987 May;7(5):1906–1916. doi: 10.1128/mcb.7.5.1906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith S., Stillman B. Stepwise assembly of chromatin during DNA replication in vitro. EMBO J. 1991 Apr;10(4):971–980. doi: 10.1002/j.1460-2075.1991.tb08031.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sorger P. K., Lewis M. J., Pelham H. R. Heat shock factor is regulated differently in yeast and HeLa cells. Nature. 1987 Sep 3;329(6134):81–84. doi: 10.1038/329081a0. [DOI] [PubMed] [Google Scholar]
- Sorger P. K., Nelson H. C. Trimerization of a yeast transcriptional activator via a coiled-coil motif. Cell. 1989 Dec 1;59(5):807–813. doi: 10.1016/0092-8674(89)90604-1. [DOI] [PubMed] [Google Scholar]
- Sorger P. K., Pelham H. R. Yeast heat shock factor is an essential DNA-binding protein that exhibits temperature-dependent phosphorylation. Cell. 1988 Sep 9;54(6):855–864. doi: 10.1016/s0092-8674(88)91219-6. [DOI] [PubMed] [Google Scholar]
- Suck D., Lahm A., Oefner C. Structure refined to 2A of a nicked DNA octanucleotide complex with DNase I. Nature. 1988 Mar 31;332(6163):464–468. doi: 10.1038/332464a0. [DOI] [PubMed] [Google Scholar]
- Susek R. E., Lindquist S. L. hsp26 of Saccharomyces cerevisiae is related to the superfamily of small heat shock proteins but is without a demonstrable function. Mol Cell Biol. 1989 Nov;9(11):5265–5271. doi: 10.1128/mcb.9.11.5265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taylor I. C., Workman J. L., Schuetz T. J., Kingston R. E. Facilitated binding of GAL4 and heat shock factor to nucleosomal templates: differential function of DNA-binding domains. Genes Dev. 1991 Jul;5(7):1285–1298. doi: 10.1101/gad.5.7.1285. [DOI] [PubMed] [Google Scholar]
- Thoma F., Simpson R. T. Local protein-DNA interactions may determine nucleosome positions on yeast plasmids. Nature. 1985 May 16;315(6016):250–252. doi: 10.1038/315250a0. [DOI] [PubMed] [Google Scholar]
- Travers A. A. The reprogramming of transcriptional competence. Cell. 1992 May 15;69(4):573–575. doi: 10.1016/0092-8674(92)90218-2. [DOI] [PubMed] [Google Scholar]
- Wolffe A. P. Implications of DNA replication for eukaryotic gene expression. J Cell Sci. 1991 Jun;99(Pt 2):201–206. doi: 10.1242/jcs.99.2.201. [DOI] [PubMed] [Google Scholar]
- Wolffe A. P. New approaches to chromatin function. New Biol. 1990 Mar;2(3):211–218. [PubMed] [Google Scholar]
- Workman J. L., Buchman A. R. Multiple functions of nucleosomes and regulatory factors in transcription. Trends Biochem Sci. 1993 Mar;18(3):90–95. doi: 10.1016/0968-0004(93)90160-o. [DOI] [PubMed] [Google Scholar]
- Workman J. L., Kingston R. E. Nucleosome core displacement in vitro via a metastable transcription factor-nucleosome complex. Science. 1992 Dec 11;258(5089):1780–1784. doi: 10.1126/science.1465613. [DOI] [PubMed] [Google Scholar]
- Xiao H., Perisic O., Lis J. T. Cooperative binding of Drosophila heat shock factor to arrays of a conserved 5 bp unit. Cell. 1991 Feb 8;64(3):585–593. doi: 10.1016/0092-8674(91)90242-q. [DOI] [PubMed] [Google Scholar]
- Zimarino V., Tsai C., Wu C. Complex modes of heat shock factor activation. Mol Cell Biol. 1990 Feb;10(2):752–759. doi: 10.1128/mcb.10.2.752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Holde K. Transcription. The omnipotent nucleosome. Nature. 1993 Mar 11;362(6416):111–112. doi: 10.1038/362111a0. [DOI] [PubMed] [Google Scholar]