Abstract
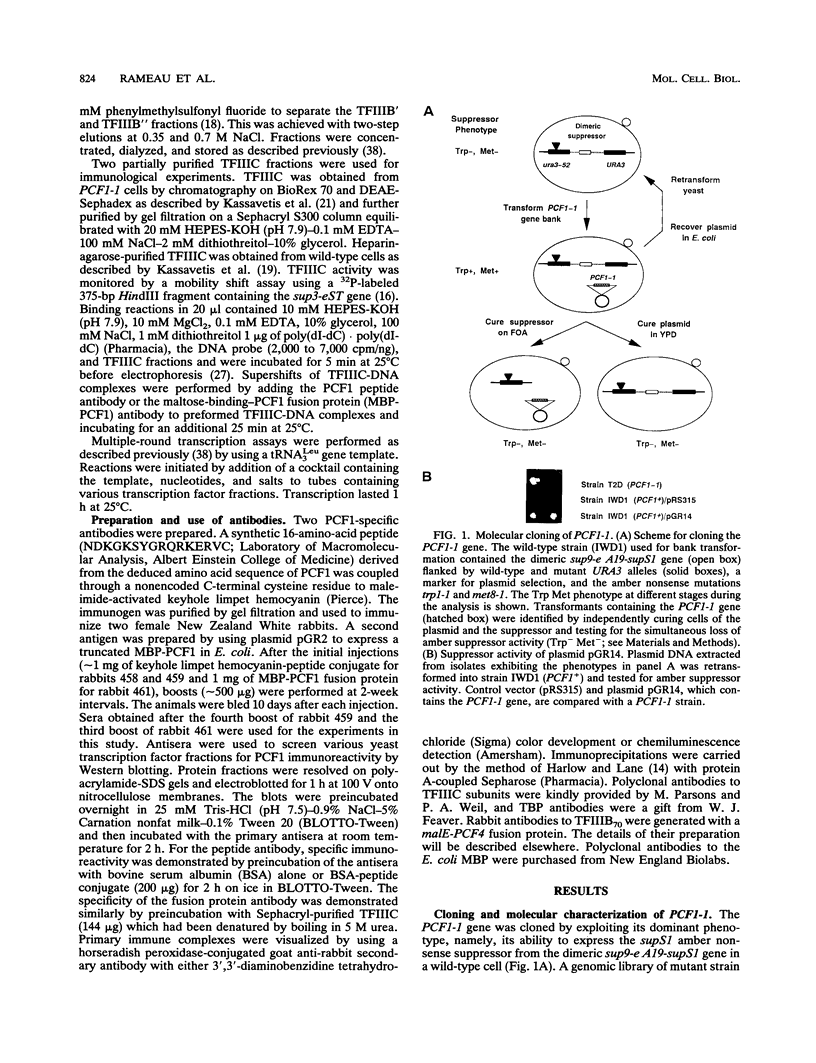
In previous studies, we have shown that the PCF1-1 mutation of Saccharomyces cerevisiae suppresses the negative effect of a tRNA gene A block promoter mutation in vivo and increases the transcription of a variety of RNA polymerase III genes in vitro. Here, we report that PCF1 encodes the second largest subunit of transcription factor IIIC (TFIIIC) and that the PCF1-1 mutation causes an amino acid substitution in a novel protein structural motif, a tetratricopeptide repeat, in this subunit. In agreement with the nature of the mutation, in vitro transcription studies with crude extracts indicate that PCF1-1 facilitates the rate-limiting step in transcription, namely, the recruitment of TFIIIB to the template. Additionally, biochemical fractionation of wild-type and mutant cell extracts shows that PCF1-1 increases the amount of the 70-kDa TFIIIB subunit detectable by Western (immunoblot) analysis in purified TFIIIB fractions and the transcription activity of a TFIIIB" fraction containing the 90-kDa subunit of this factor. We suggest that the effect of PCF1-1 on TFIIIB activity in vitro is a consequence of its increased rate of recruitment in vivo.
Full text
PDF



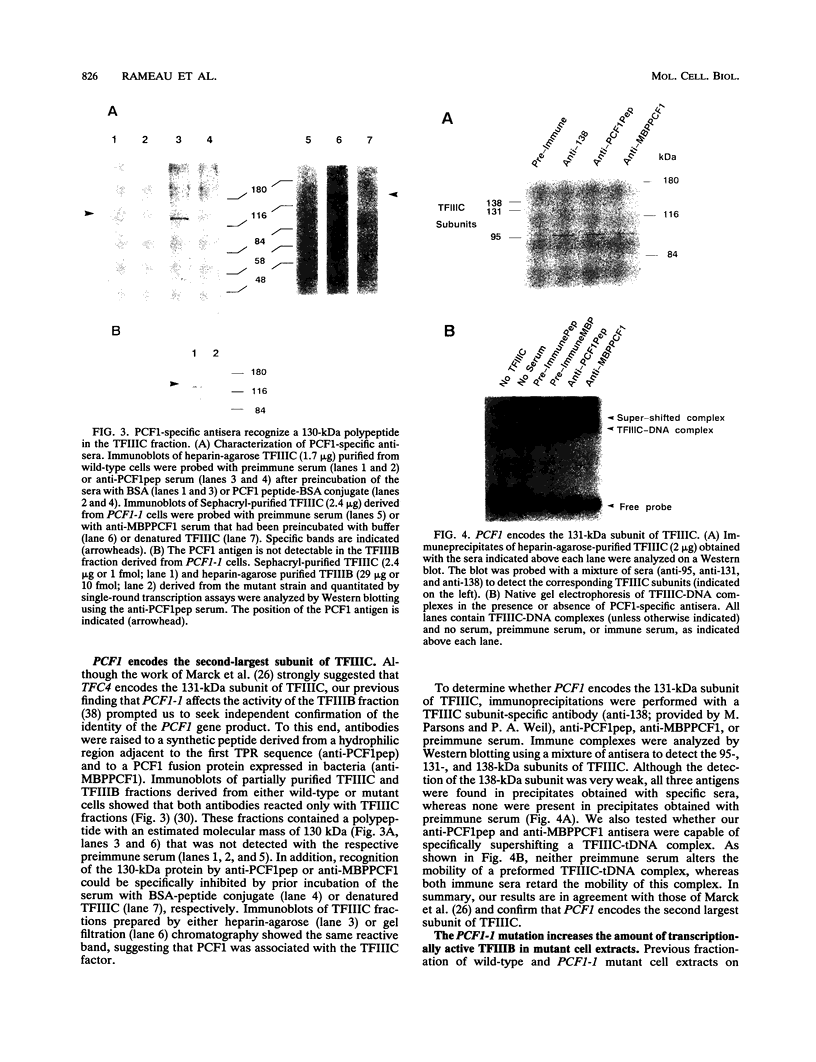

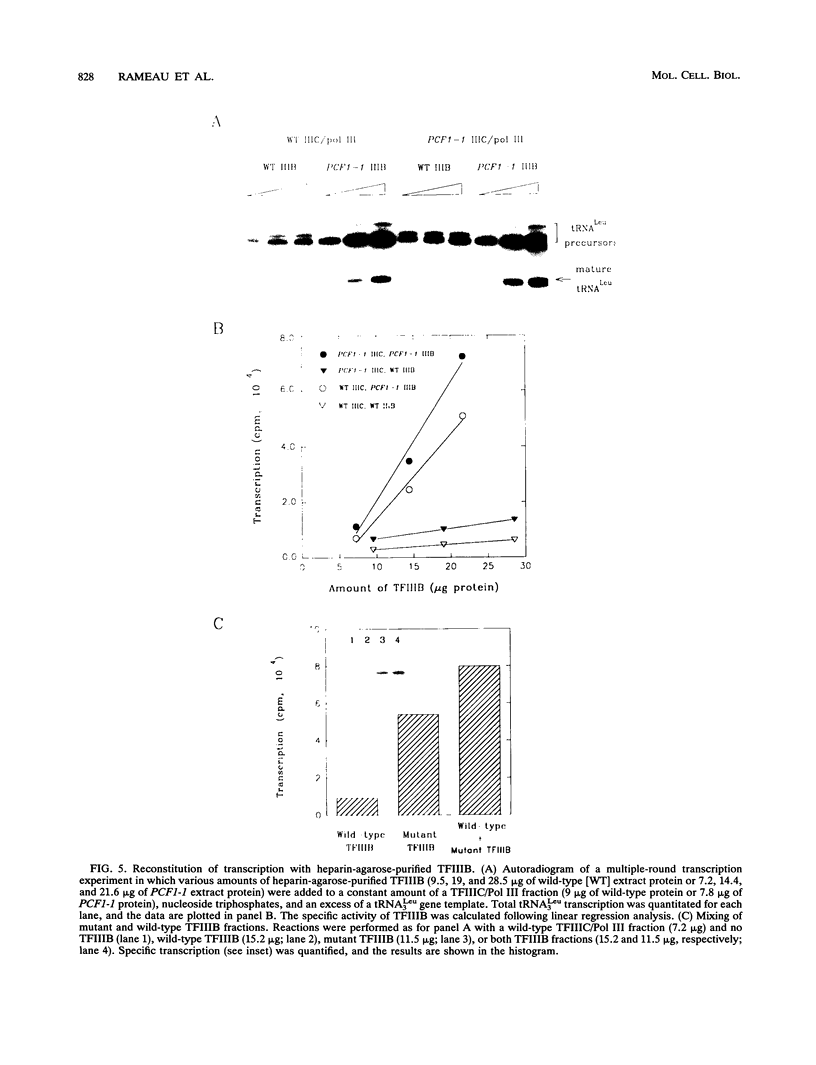
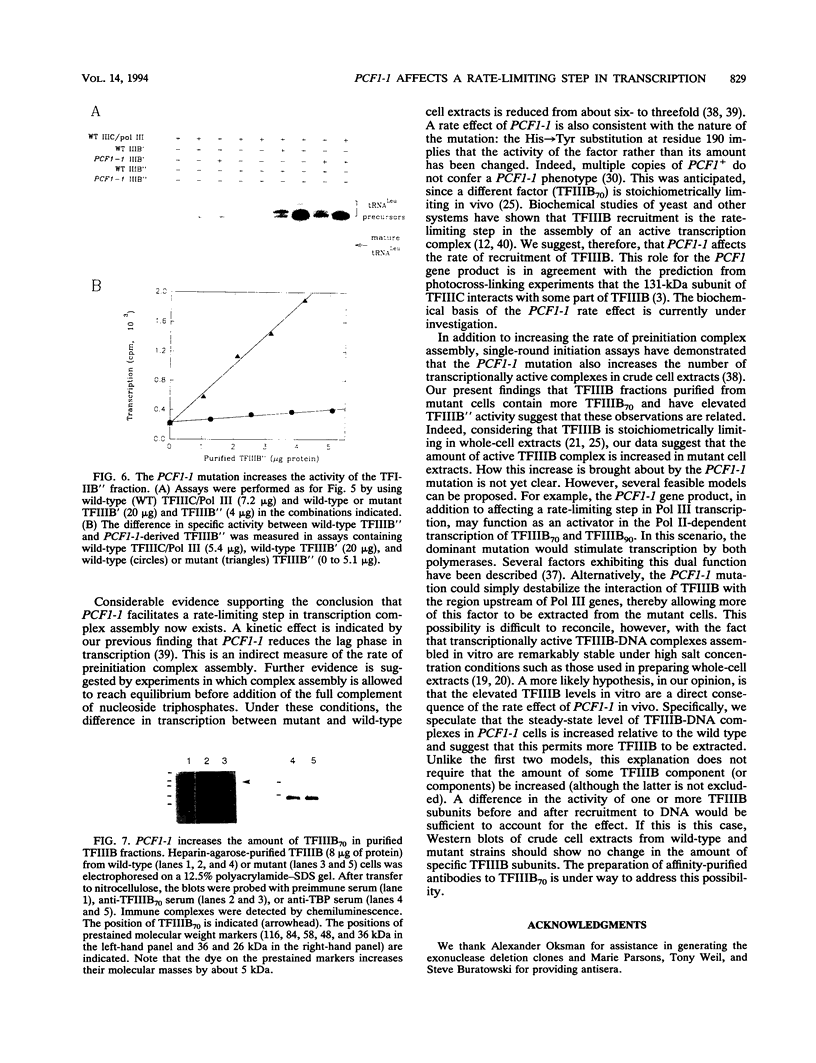

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Bartholomew B., Kassavetis G. A., Braun B. R., Geiduschek E. P. The subunit structure of Saccharomyces cerevisiae transcription factor IIIC probed with a novel photocrosslinking reagent. EMBO J. 1990 Jul;9(7):2197–2205. doi: 10.1002/j.1460-2075.1990.tb07389.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartholomew B., Kassavetis G. A., Geiduschek E. P. Two components of Saccharomyces cerevisiae transcription factor IIIB (TFIIIB) are stereospecifically located upstream of a tRNA gene and interact with the second-largest subunit of TFIIIC. Mol Cell Biol. 1991 Oct;11(10):5181–5189. doi: 10.1128/mcb.11.10.5181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Braun B. R., Bartholomew B., Kassavetis G. A., Geiduschek E. P. Topography of transcription factor complexes on the Saccharomyces cerevisiae 5 S RNA gene. J Mol Biol. 1992 Dec 20;228(4):1063–1077. doi: 10.1016/0022-2836(92)90315-b. [DOI] [PubMed] [Google Scholar]
- Buratowski S., Zhou H. A suppressor of TBP mutations encodes an RNA polymerase III transcription factor with homology to TFIIB. Cell. 1992 Oct 16;71(2):221–230. doi: 10.1016/0092-8674(92)90351-c. [DOI] [PubMed] [Google Scholar]
- Colbert T., Hahn S. A yeast TFIIB-related factor involved in RNA polymerase III transcription. Genes Dev. 1992 Oct;6(10):1940–1949. doi: 10.1101/gad.6.10.1940. [DOI] [PubMed] [Google Scholar]
- Cormack B. P., Struhl K. The TATA-binding protein is required for transcription by all three nuclear RNA polymerases in yeast cells. Cell. 1992 May 15;69(4):685–696. doi: 10.1016/0092-8674(92)90232-2. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dieci G., Duimio L., Coda-Zabetta F., Sprague K. U., Ottonello S. A novel RNA polymerase III transcription factor fraction that is not required for template commitment. J Biol Chem. 1993 May 25;268(15):11199–11207. [PubMed] [Google Scholar]
- Gabrielsen O. S., Marzouki N., Ruet A., Sentenac A., Fromageot P. Two polypeptide chains in yeast transcription factor tau interact with DNA. J Biol Chem. 1989 May 5;264(13):7505–7511. [PubMed] [Google Scholar]
- Gabrielsen O. S., Sentenac A. RNA polymerase III (C) and its transcription factors. Trends Biochem Sci. 1991 Nov;16(11):412–416. doi: 10.1016/0968-0004(91)90166-s. [DOI] [PubMed] [Google Scholar]
- Geiduschek E. P., Tocchini-Valentini G. P. Transcription by RNA polymerase III. Annu Rev Biochem. 1988;57:873–914. doi: 10.1146/annurev.bi.57.070188.004301. [DOI] [PubMed] [Google Scholar]
- Goebl M., Yanagida M. The TPR snap helix: a novel protein repeat motif from mitosis to transcription. Trends Biochem Sci. 1991 May;16(5):173–177. doi: 10.1016/0968-0004(91)90070-c. [DOI] [PubMed] [Google Scholar]
- Hill J., Donald K. A., Griffiths D. E., Donald G. DMSO-enhanced whole cell yeast transformation. Nucleic Acids Res. 1991 Oct 25;19(20):5791–5791. doi: 10.1093/nar/19.20.5791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hottinger-Werlen A., Schaack J., Lapointe J., Mao J., Nichols M., Söll D. Dimeric tRNA gene arrangement in Schizosaccharomyces pombe allows increased expression of the downstream gene. Nucleic Acids Res. 1985 Dec 20;13(24):8739–8747. doi: 10.1093/nar/13.24.8739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones J. S., Prakash L. Yeast Saccharomyces cerevisiae selectable markers in pUC18 polylinkers. Yeast. 1990 Sep-Oct;6(5):363–366. doi: 10.1002/yea.320060502. [DOI] [PubMed] [Google Scholar]
- Kassavetis G. A., Bartholomew B., Blanco J. A., Johnson T. E., Geiduschek E. P. Two essential components of the Saccharomyces cerevisiae transcription factor TFIIIB: transcription and DNA-binding properties. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7308–7312. doi: 10.1073/pnas.88.16.7308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kassavetis G. A., Braun B. R., Nguyen L. H., Geiduschek E. P. S. cerevisiae TFIIIB is the transcription initiation factor proper of RNA polymerase III, while TFIIIA and TFIIIC are assembly factors. Cell. 1990 Jan 26;60(2):235–245. doi: 10.1016/0092-8674(90)90739-2. [DOI] [PubMed] [Google Scholar]
- Kassavetis G. A., Joazeiro C. A., Pisano M., Geiduschek E. P., Colbert T., Hahn S., Blanco J. A. The role of the TATA-binding protein in the assembly and function of the multisubunit yeast RNA polymerase III transcription factor, TFIIIB. Cell. 1992 Dec 11;71(6):1055–1064. doi: 10.1016/0092-8674(92)90399-w. [DOI] [PubMed] [Google Scholar]
- Kassavetis G. A., Riggs D. L., Negri R., Nguyen L. H., Geiduschek E. P. Transcription factor IIIB generates extended DNA interactions in RNA polymerase III transcription complexes on tRNA genes. Mol Cell Biol. 1989 Jun;9(6):2551–2566. doi: 10.1128/mcb.9.6.2551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klekamp M. S., Weil P. A. Partial purification and characterization of the Saccharomyces cerevisiae transcription factor TFIIIB. J Biol Chem. 1986 Feb 25;261(6):2819–2827. [PubMed] [Google Scholar]
- Lefebvre O., Carles C., Conesa C., Swanson R. N., Bouet F., Riva M., Sentenac A. TFC3: gene encoding the B-block binding subunit of the yeast transcription factor IIIC. Proc Natl Acad Sci U S A. 1992 Nov 1;89(21):10512–10516. doi: 10.1073/pnas.89.21.10512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lobo S. M., Tanaka M., Sullivan M. L., Hernandez N. A TBP complex essential for transcription from TATA-less but not TATA-containing RNA polymerase III promoters is part of the TFIIIB fraction. Cell. 1992 Dec 11;71(6):1029–1040. doi: 10.1016/0092-8674(92)90397-u. [DOI] [PubMed] [Google Scholar]
- López-De-León A., Librizzi M., Puglia K., Willis I. M. PCF4 encodes an RNA polymerase III transcription factor with homology to TFIIB. Cell. 1992 Oct 16;71(2):211–220. doi: 10.1016/0092-8674(92)90350-l. [DOI] [PubMed] [Google Scholar]
- Marck C., Lefebvre O., Carles C., Riva M., Chaussivert N., Ruet A., Sentenac A. The TFIIIB-assembling subunit of yeast transcription factor TFIIIC has both tetratricopeptide repeats and basic helix-loop-helix motifs. Proc Natl Acad Sci U S A. 1993 May 1;90(9):4027–4031. doi: 10.1073/pnas.90.9.4027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nichols M., Söll D., Willis I. Yeast RNase P: catalytic activity and substrate binding are separate functions. Proc Natl Acad Sci U S A. 1988 Mar;85(5):1379–1383. doi: 10.1073/pnas.85.5.1379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parsons M. C., Weil P. A. Cloning of TFC1, the Saccharomyces cerevisiae gene encoding the 95-kDa subunit of transcription factor TFIIIC. J Biol Chem. 1992 Feb 15;267(5):2894–2901. [PubMed] [Google Scholar]
- Parsons M. C., Weil P. A. Purification and characterization of Saccharomyces cerevisiae transcription factor TFIIIC. Polypeptide composition defined with polyclonal antibodies. J Biol Chem. 1990 Mar 25;265(9):5095–5103. [PubMed] [Google Scholar]
- Rockmill B., Lambie E. J., Roeder G. S. Spore enrichment. Methods Enzymol. 1991;194:146–149. doi: 10.1016/0076-6879(91)94012-2. [DOI] [PubMed] [Google Scholar]
- Schultz M. C., Reeder R. H., Hahn S. Variants of the TATA-binding protein can distinguish subsets of RNA polymerase I, II, and III promoters. Cell. 1992 May 15;69(4):697–702. doi: 10.1016/0092-8674(92)90233-3. [DOI] [PubMed] [Google Scholar]
- Swanson R. N., Conesa C., Lefebvre O., Carles C., Ruet A., Quemeneur E., Gagnon J., Sentenac A. Isolation of TFC1, a gene encoding one of two DNA-binding subunits of yeast transcription factor tau (TFIIIC). Proc Natl Acad Sci U S A. 1991 Jun 1;88(11):4887–4891. doi: 10.1073/pnas.88.11.4887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taggart A. K., Fisher T. S., Pugh B. F. The TATA-binding protein and associated factors are components of pol III transcription factor TFIIIB. Cell. 1992 Dec 11;71(6):1015–1028. doi: 10.1016/0092-8674(92)90396-t. [DOI] [PubMed] [Google Scholar]
- White R. J., Jackson S. P. Mechanism of TATA-binding protein recruitment to a TATA-less class III promoter. Cell. 1992 Dec 11;71(6):1041–1053. doi: 10.1016/0092-8674(92)90398-v. [DOI] [PubMed] [Google Scholar]
- Willis I. M. RNA polymerase III. Genes, factors and transcriptional specificity. Eur J Biochem. 1993 Feb 15;212(1):1–11. doi: 10.1111/j.1432-1033.1993.tb17626.x. [DOI] [PubMed] [Google Scholar]
- Willis I., Oksman A., López-De-León A. The PCF1-1 mutation increases the activity of the transcription factor (TF) IIIB fraction from Saccharomyces cerevisiae. Nucleic Acids Res. 1992 Jul 25;20(14):3725–3730. doi: 10.1093/nar/20.14.3725. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willis I., Schmidt P., Söll D. A selection for mutants of the RNA polymerase III transcription apparatus: PCF1 stimulates transcription of tRNA and 5S RNA genes. EMBO J. 1989 Dec 20;8(13):4281–4288. doi: 10.1002/j.1460-2075.1989.tb08614.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Brown D. D. Developmental regulation of two 5S ribosomal RNA genes. Science. 1988 Sep 23;241(4873):1626–1632. doi: 10.1126/science.241.4873.1626. [DOI] [PubMed] [Google Scholar]