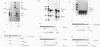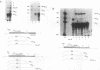Abstract
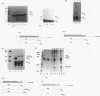
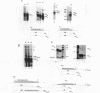
Mammalian histone mRNAs end in a highly conserved stem-loop structure, with a six-base stem and a four-base loop. We have examined the effect of mutating the stem-loop on the expression of the histone mRNA in vivo by introducing the mutated histone genes into CHO cells by stable transfection. Point mutations have been introduced into the loop sequence and into the UA base pair at the top of the stem. Changing either the first or the third base of the conserved UYUN sequence in the loop to a purine greatly reduced expression, while changing both U's to purines abolished expression. A number of alterations in the stem sequence, including reversing the stem sequence, reversing the two base pairs at the base of the stem, or destroying the UA base pair at the top of the stem, also abolished expression. Changing the UA base pair to a CG or a UG base pair also reduced expression. The loss of expression is due to inefficient processing of the pre-mRNA, as judged by the efficiency of processing in vitro. Addition of a polyadenylation site or the wild-type histone processing signal downstream of a mutant stem-loop resulted in rescuing the processing of the mutant pre-histone mRNA. These results suggest that if the histone pre-mRNA is not rapidly processed, then it is degraded.
Full text
PDF


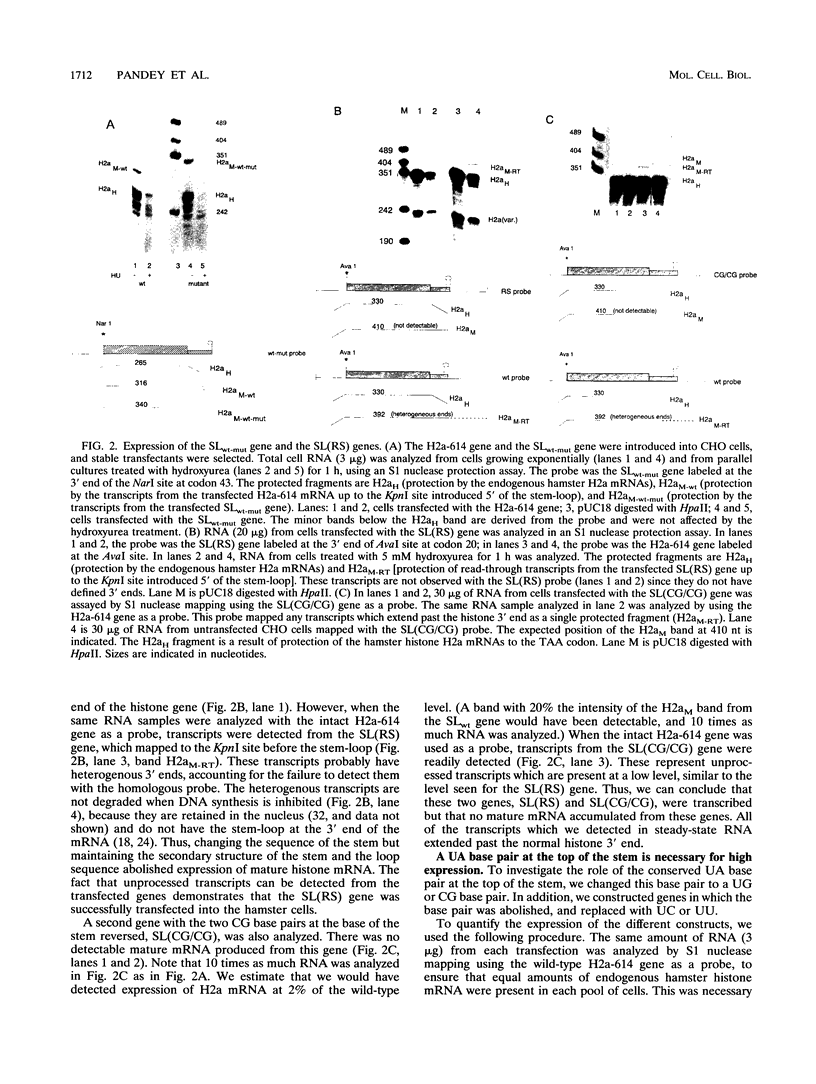
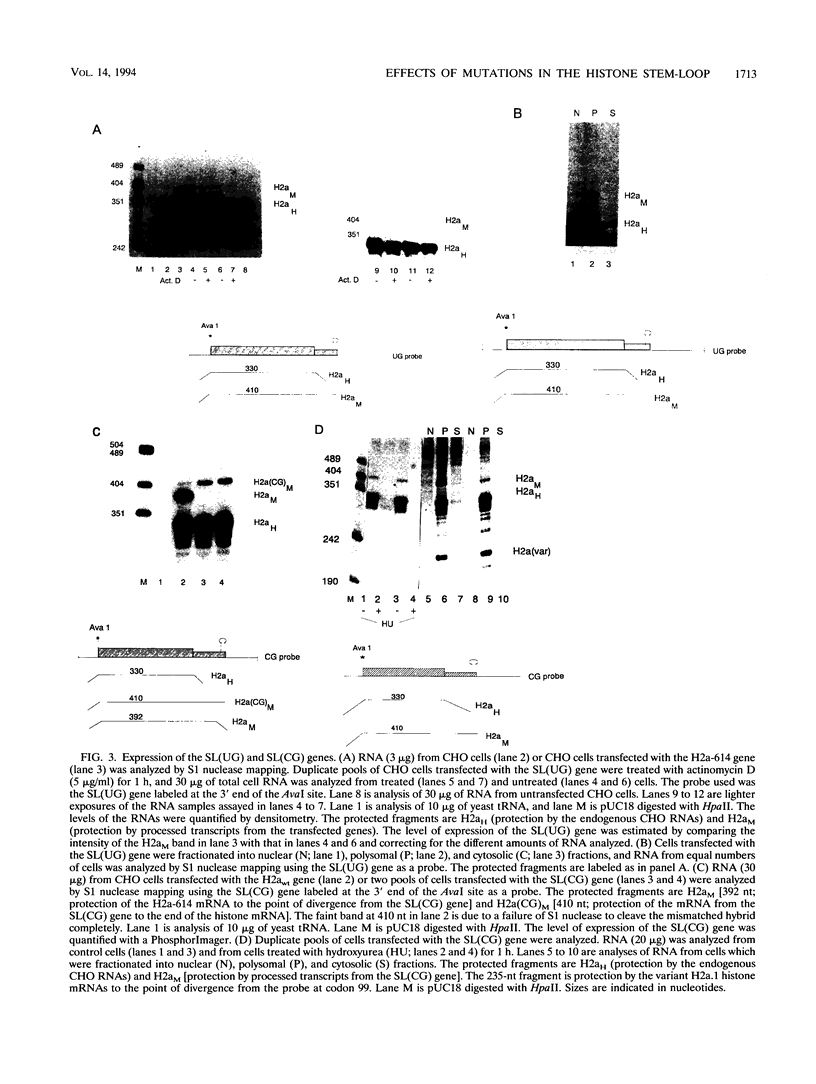

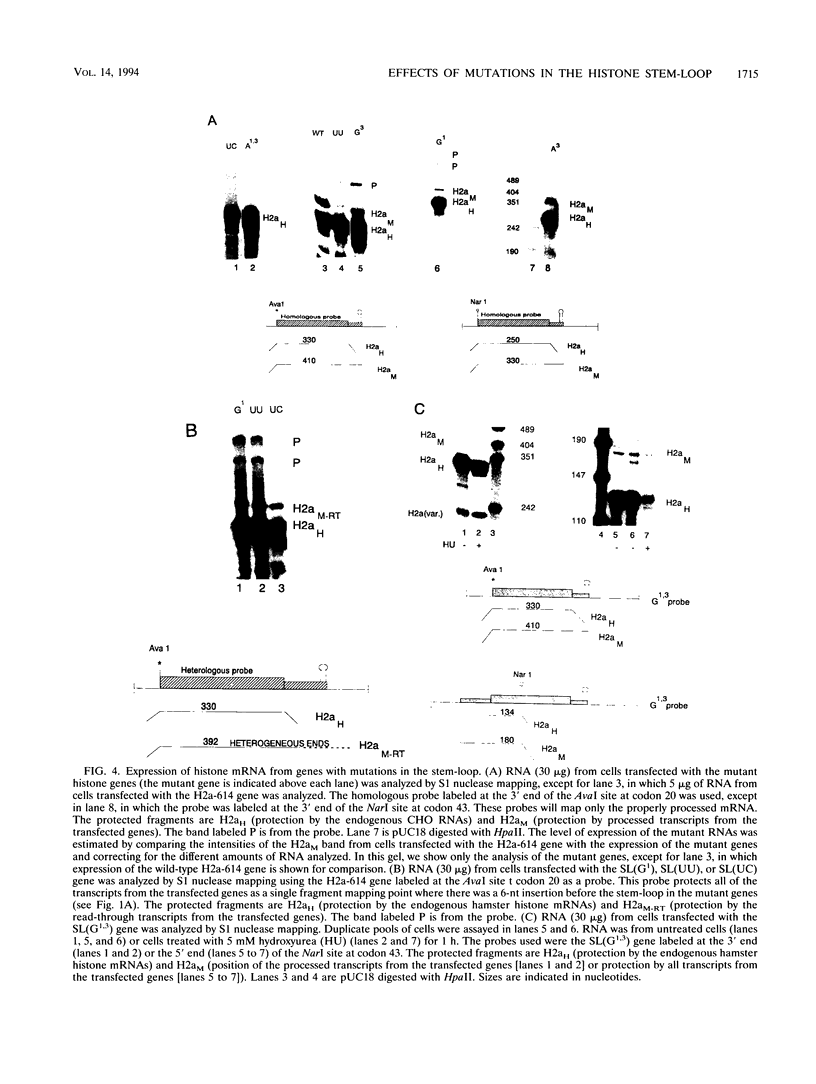

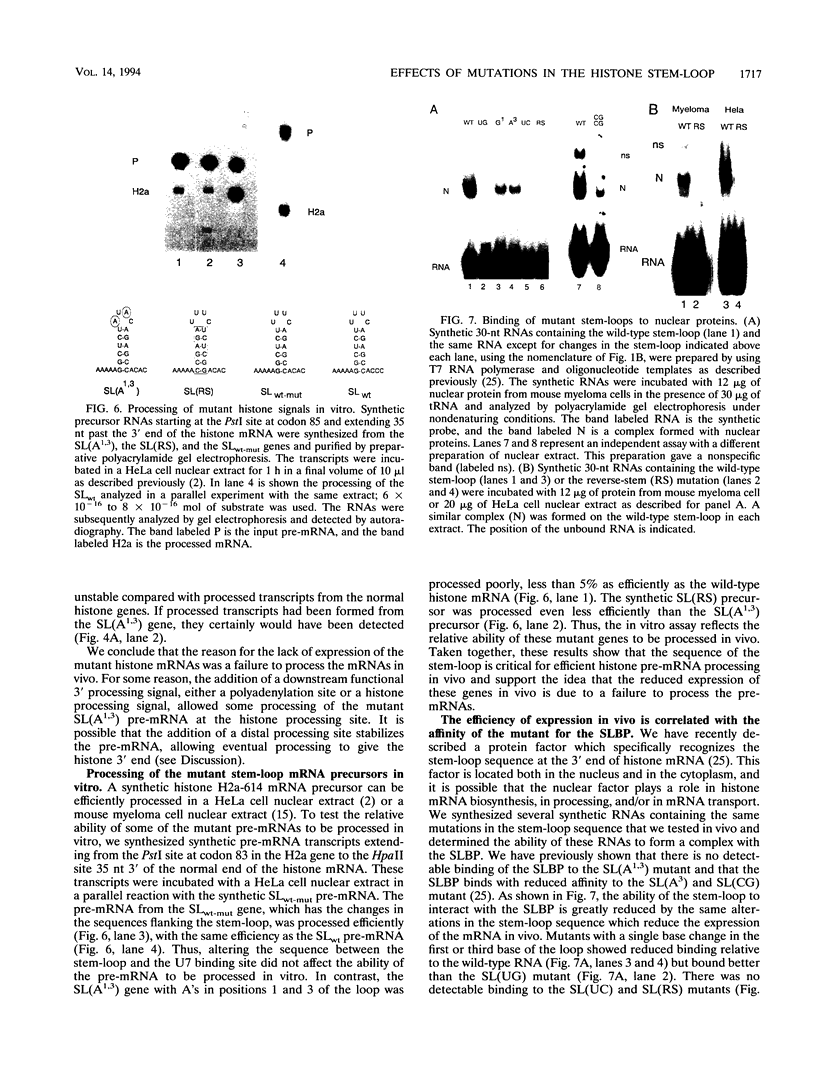



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnstiel M. L., Busslinger M., Strub K. Transcription termination and 3' processing: the end is in site! Cell. 1985 Jun;41(2):349–359. doi: 10.1016/s0092-8674(85)80007-6. [DOI] [PubMed] [Google Scholar]
- Bond U. M., Yario T. A., Steitz J. A. Multiple processing-defective mutations in a mammalian histone pre-mRNA are suppressed by compensatory changes in U7 RNA both in vivo and in vitro. Genes Dev. 1991 Sep;5(9):1709–1722. doi: 10.1101/gad.5.9.1709. [DOI] [PubMed] [Google Scholar]
- Cotten M., Gick O., Vasserot A., Schaffner G., Birnstiel M. L. Specific contacts between mammalian U7 snRNA and histone precursor RNA are indispensable for the in vitro 3' RNA processing reaction. EMBO J. 1988 Mar;7(3):801–808. doi: 10.1002/j.1460-2075.1988.tb02878.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eckner R., Ellmeier W., Birnstiel M. L. Mature mRNA 3' end formation stimulates RNA export from the nucleus. EMBO J. 1991 Nov;10(11):3513–3522. doi: 10.1002/j.1460-2075.1991.tb04915.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Keller W., Birnstiel M. L. Generation of histone mRNA 3' ends by endonucleolytic cleavage of the pre-mRNA in a snRNP-dependent in vitro reaction. EMBO J. 1986 Jun;5(6):1319–1326. doi: 10.1002/j.1460-2075.1986.tb04362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Vasserot A., Birnstiel M. L. Heat-labile regulatory factor is required for 3' processing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8937–8940. doi: 10.1073/pnas.84.24.8937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves R. A., Marzluff W. F. Rapid reversible changes in the rate of histone gene transcription and histone mRNA levels in mouse myeloma cells. Mol Cell Biol. 1984 Feb;4(2):351–357. doi: 10.1128/mcb.4.2.351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves R. A., Pandey N. B., Chodchoy N., Marzluff W. F. Translation is required for regulation of histone mRNA degradation. Cell. 1987 Feb 27;48(4):615–626. doi: 10.1016/0092-8674(87)90240-6. [DOI] [PubMed] [Google Scholar]
- Graves R. A., Wellman S. E., Chiu I. M., Marzluff W. F. Differential expression of two clusters of mouse histone genes. J Mol Biol. 1985 May 25;183(2):179–194. doi: 10.1016/0022-2836(85)90211-6. [DOI] [PubMed] [Google Scholar]
- Harris M. E., Böhni R., Schneiderman M. H., Ramamurthy L., Schümperli D., Marzluff W. F. Regulation of histone mRNA in the unperturbed cell cycle: evidence suggesting control at two posttranscriptional steps. Mol Cell Biol. 1991 May;11(5):2416–2424. doi: 10.1128/mcb.11.5.2416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffmann I., Birnstiel M. L. Cell cycle-dependent regulation of histone precursor mRNA processing by modulation of U7 snRNA accessibility. Nature. 1990 Aug 16;346(6285):665–668. doi: 10.1038/346665a0. [DOI] [PubMed] [Google Scholar]
- Jackson R. J., Standart N. Do the poly(A) tail and 3' untranslated region control mRNA translation? Cell. 1990 Jul 13;62(1):15–24. doi: 10.1016/0092-8674(90)90235-7. [DOI] [PubMed] [Google Scholar]
- Levine B. J., Liu T. J., Marzluff W. F., Skoultchi A. I. Differential expression of individual members of the histone multigene family due to sequences in the 5' and 3' regions of the genes. Mol Cell Biol. 1988 May;8(5):1887–1895. doi: 10.1128/mcb.8.5.1887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu T. J., Levine B. J., Skoultchi A. I., Marzluff W. F. The efficiency of 3'-end formation contributes to the relative levels of different histone mRNAs. Mol Cell Biol. 1989 Aug;9(8):3499–3508. doi: 10.1128/mcb.9.8.3499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lüscher B., Schümperli D. RNA 3' processing regulates histone mRNA levels in a mammalian cell cycle mutant. A processing factor becomes limiting in G1-arrested cells. EMBO J. 1987 Jun;6(6):1721–1726. doi: 10.1002/j.1460-2075.1987.tb02423.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marzluff W. F. Histone 3' ends: essential and regulatory functions. Gene Expr. 1992;2(2):93–97. [PMC free article] [PubMed] [Google Scholar]
- Marzluff W. F., Pandey N. B. Multiple regulatory steps control histone mRNA concentrations. Trends Biochem Sci. 1988 Feb;13(2):49–52. doi: 10.1016/0968-0004(88)90027-8. [DOI] [PubMed] [Google Scholar]
- Melin L., Soldati D., Mital R., Streit A., Schümperli D. Biochemical demonstration of complex formation of histone pre-mRNA with U7 small nuclear ribonucleoprotein and hairpin binding factors. EMBO J. 1992 Feb;11(2):691–697. doi: 10.1002/j.1460-2075.1992.tb05101.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan J. F., Groebe D. R., Witherell G. W., Uhlenbeck O. C. Oligoribonucleotide synthesis using T7 RNA polymerase and synthetic DNA templates. Nucleic Acids Res. 1987 Nov 11;15(21):8783–8798. doi: 10.1093/nar/15.21.8783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Oh R., Steitz J. A. Each of the conserved sequence elements flanking the cleavage site of mammalian histone pre-mRNAs has a distinct role in the 3'-end processing reaction. Mol Cell Biol. 1989 Jul;9(7):3105–3108. doi: 10.1128/mcb.9.7.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Both conserved signals on mammalian histone pre-mRNAs associate with small nuclear ribonucleoproteins during 3' end formation in vitro. Mol Cell Biol. 1987 May;7(5):1663–1672. doi: 10.1128/mcb.7.5.1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey N. B., Chodchoy N., Liu T. J., Marzluff W. F. Introns in histone genes alter the distribution of 3' ends. Nucleic Acids Res. 1990 Jun 11;18(11):3161–3170. doi: 10.1093/nar/18.11.3161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey N. B., Marzluff W. F. The stem-loop structure at the 3' end of histone mRNA is necessary and sufficient for regulation of histone mRNA stability. Mol Cell Biol. 1987 Dec;7(12):4557–4559. doi: 10.1128/mcb.7.12.4557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pandey N. B., Sun J. H., Marzluff W. F. Different complexes are formed on the 3' end of histone mRNA with nuclear and polyribosomal proteins. Nucleic Acids Res. 1991 Oct 25;19(20):5653–5659. doi: 10.1093/nar/19.20.5653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pilch D. R., Marzluff W. F. Expression of histone-U1 snRNA chimeric genes: U1 promoters are compatible with histone 3' end formation. Gene Expr. 1991 Apr;1(1):41–53. [PMC free article] [PubMed] [Google Scholar]
- Roberts S. B., Emmons S. W., Childs G. Nucleotide sequences of Caenorhabditis elegans core histone genes. Genes for different histone classes share common flanking sequence elements. J Mol Biol. 1989 Apr 20;206(4):567–577. doi: 10.1016/0022-2836(89)90566-4. [DOI] [PubMed] [Google Scholar]
- Sittman D. B., Graves R. A., Marzluff W. F. Histone mRNA concentrations are regulated at the level of transcription and mRNA degradation. Proc Natl Acad Sci U S A. 1983 Apr;80(7):1849–1853. doi: 10.1073/pnas.80.7.1849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stauber C., Schümperli D. 3' processing of pre-mRNA plays a major role in proliferation-dependent regulation of histone gene expression. Nucleic Acids Res. 1988 Oct 25;16(20):9399–9414. doi: 10.1093/nar/16.20.9399. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streit A., Koning T. W., Soldati D., Melin L., Schümperli D. Variable effects of the conserved RNA hairpin element upon 3' end processing of histone pre-mRNA in vitro. Nucleic Acids Res. 1993 Apr 11;21(7):1569–1575. doi: 10.1093/nar/21.7.1569. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Birnstiel M. L. Genetic complementation in the Xenopus oocyte: co-expression of sea urchin histone and U7 RNAs restores 3' processing of H3 pre-mRNA in the oocyte. EMBO J. 1986 Jul;5(7):1675–1682. doi: 10.1002/j.1460-2075.1986.tb04411.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun J., Pilch D. R., Marzluff W. F. The histone mRNA 3' end is required for localization of histone mRNA to polyribosomes. Nucleic Acids Res. 1992 Nov 25;20(22):6057–6066. doi: 10.1093/nar/20.22.6057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasserot A. P., Schaufele F. J., Birnstiel M. L. Conserved terminal hairpin sequences of histone mRNA precursors are not involved in duplex formation with the U7 RNA but act as a target site for a distinct processing factor. Proc Natl Acad Sci U S A. 1989 Jun;86(12):4345–4349. doi: 10.1073/pnas.86.12.4345. [DOI] [PMC free article] [PubMed] [Google Scholar]