Abstract
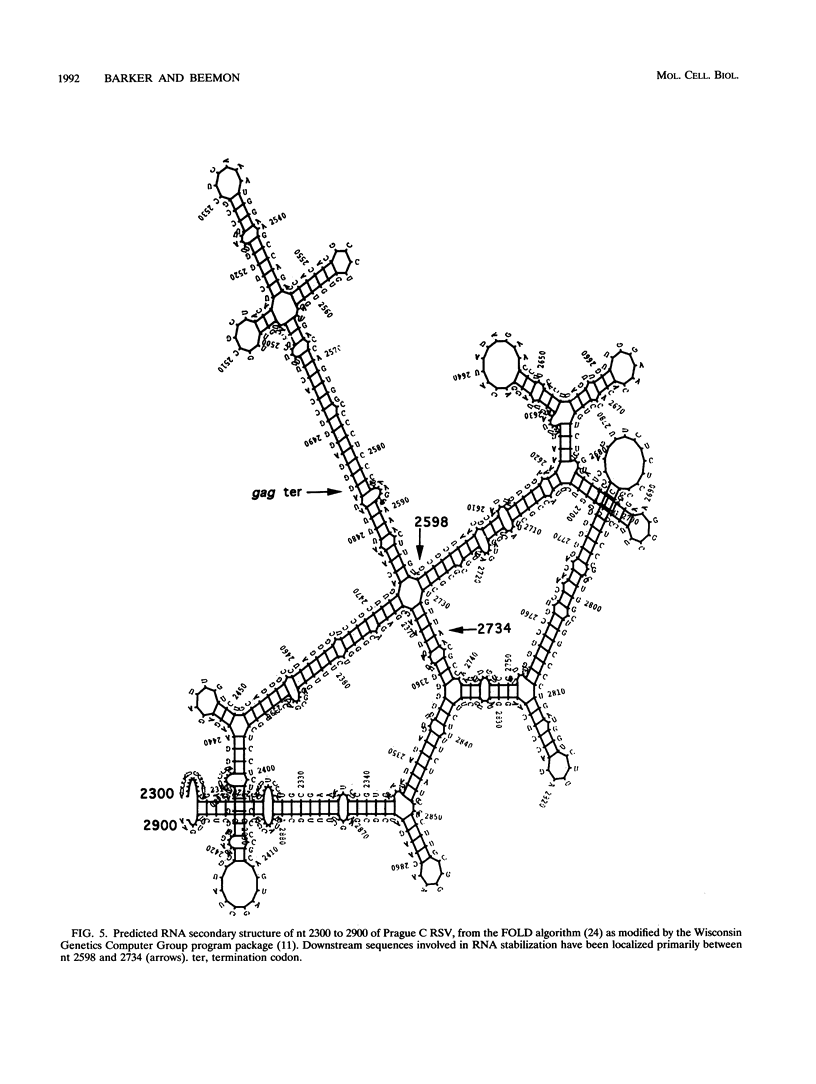
The intracellular accumulation of the unspliced RNA of Rous sarcoma virus was decreased when translation was prematurely terminated by the introduction of nonsense codons within its 5' proximal gene, the gag gene. Subcellular fractionation of transfected cells suggested that nonsense codon-mediated instability occurred in the cytoplasm. Analysis of constructs containing an in-frame deletion in the nucleocapsid domain of gag, which prevents interaction between the Gag protein and viral RNA, showed that an open reading frame extending to approximately 30 nucleotides from the natural gag termination codon was needed for RNA stability. Sequences at the gag-pol junction necessary for ribosomal frameshifting were not required for RNA stability; however, sequences located 100 to 200 nucleotides downstream of the natural gag termination codon were found to be necessary for stable RNA. The stability of RNAs lacking this downstream sequence was not markedly affected by premature termination codons. We propose that this downstream RNA sequence may interact with ribosomes translating gag to stabilize the RNA.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arrigo S., Beemon K. Regulation of Rous sarcoma virus RNA splicing and stability. Mol Cell Biol. 1988 Nov;8(11):4858–4867. doi: 10.1128/mcb.8.11.4858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barker G. F., Beemon K. Nonsense codons within the Rous sarcoma virus gag gene decrease the stability of unspliced viral RNA. Mol Cell Biol. 1991 May;11(5):2760–2768. doi: 10.1128/mcb.11.5.2760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baserga S. J., Benz E. J., Jr Beta-globin nonsense mutation: deficient accumulation of mRNA occurs despite normal cytoplasmic stability. Proc Natl Acad Sci U S A. 1992 Apr 1;89(7):2935–2939. doi: 10.1073/pnas.89.7.2935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnie G. D. Isolation of nuclei from animal cells in culture. Methods Cell Biol. 1978;17:13–26. doi: 10.1016/s0091-679x(08)61131-0. [DOI] [PubMed] [Google Scholar]
- Cheng J., Fogel-Petrovic M., Maquat L. E. Translation to near the distal end of the penultimate exon is required for normal levels of spliced triosephosphate isomerase mRNA. Mol Cell Biol. 1990 Oct;10(10):5215–5225. doi: 10.1128/mcb.10.10.5215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chomczynski P., Sacchi N. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal Biochem. 1987 Apr;162(1):156–159. doi: 10.1006/abio.1987.9999. [DOI] [PubMed] [Google Scholar]
- Desjardins P., Morais R. Sequence and gene organization of the chicken mitochondrial genome. A novel gene order in higher vertebrates. J Mol Biol. 1990 Apr 20;212(4):599–634. doi: 10.1016/0022-2836(90)90225-B. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacks T., Madhani H. D., Masiarz F. R., Varmus H. E. Signals for ribosomal frameshifting in the Rous sarcoma virus gag-pol region. Cell. 1988 Nov 4;55(3):447–458. doi: 10.1016/0092-8674(88)90031-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacks T. Translational suppression in gene expression in retroviruses and retrotransposons. Curr Top Microbiol Immunol. 1990;157:93–124. doi: 10.1007/978-3-642-75218-6_4. [DOI] [PubMed] [Google Scholar]
- Loeb D. M., Woolford J., Beemon K. pp60c-src has less affinity for the detergent-insoluble cellular matrix than do pp60v-src and other viral protein-tyrosine kinases. J Virol. 1987 Aug;61(8):2420–2427. doi: 10.1128/jvi.61.8.2420-2427.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marzluff W. F. Preparation of active nuclei. Methods Enzymol. 1990;181:30–36. doi: 10.1016/0076-6879(90)81109-8. [DOI] [PubMed] [Google Scholar]
- McNally M. T., Gontarek R. R., Beemon K. Characterization of Rous sarcoma virus intronic sequences that negatively regulate splicing. Virology. 1991 Nov;185(1):99–108. doi: 10.1016/0042-6822(91)90758-4. [DOI] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Méric C., Spahr P. F. Rous sarcoma virus nucleic acid-binding protein p12 is necessary for viral 70S RNA dimer formation and packaging. J Virol. 1986 Nov;60(2):450–459. doi: 10.1128/jvi.60.2.450-459.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz D. E., Tizard R., Gilbert W. Nucleotide sequence of Rous sarcoma virus. Cell. 1983 Mar;32(3):853–869. doi: 10.1016/0092-8674(83)90071-5. [DOI] [PubMed] [Google Scholar]
- Stoltzfus C. M. Synthesis and processing of avian sarcoma retrovirus RNA. Adv Virus Res. 1988;35:1–38. doi: 10.1016/s0065-3527(08)60707-1. [DOI] [PubMed] [Google Scholar]
- Stone E. M., Rothblum K. N., Alevy M. C., Kuo T. M., Schwartz R. J. Complete sequence of the chicken glyceraldehyde-3-phosphate dehydrogenase gene. Proc Natl Acad Sci U S A. 1985 Mar;82(6):1628–1632. doi: 10.1073/pnas.82.6.1628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urlaub G., Mitchell P. J., Ciudad C. J., Chasin L. A. Nonsense mutations in the dihydrofolate reductase gene affect RNA processing. Mol Cell Biol. 1989 Jul;9(7):2868–2880. doi: 10.1128/mcb.9.7.2868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wuarin J., Schibler U. Expression of the liver-enriched transcriptional activator protein DBP follows a stringent circadian rhythm. Cell. 1990 Dec 21;63(6):1257–1266. doi: 10.1016/0092-8674(90)90421-a. [DOI] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]