Abstract
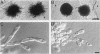
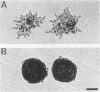
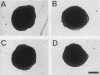
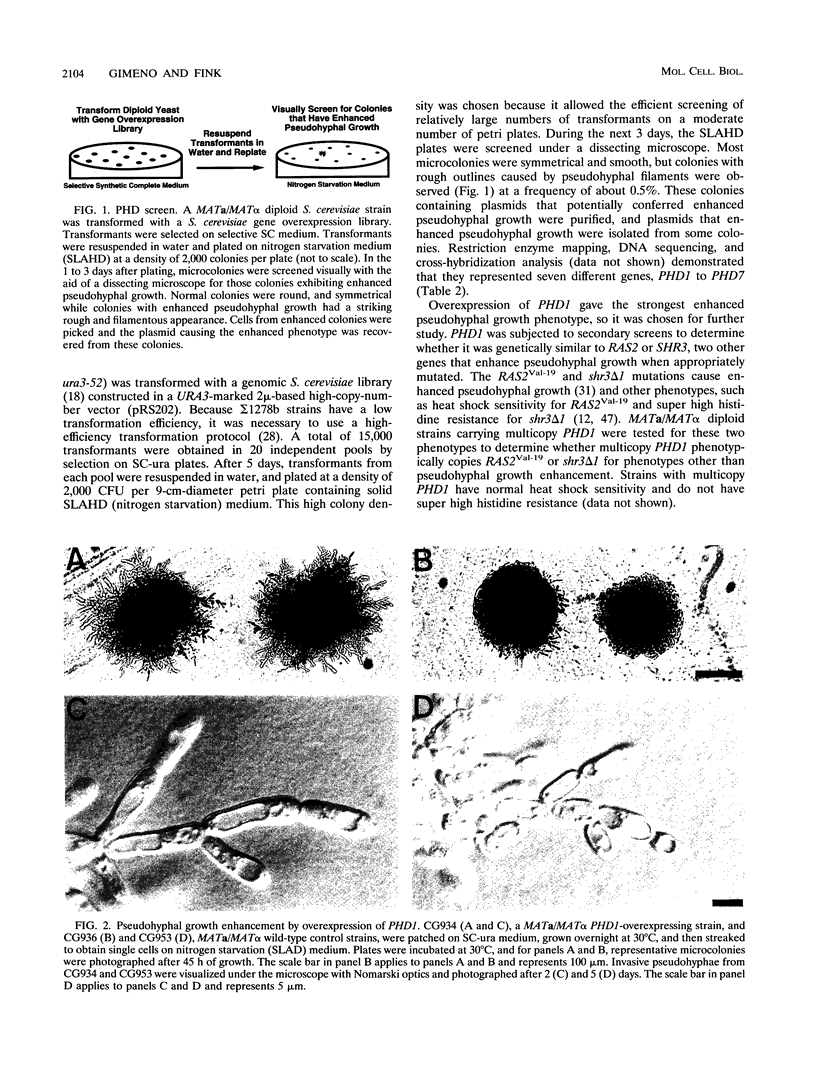
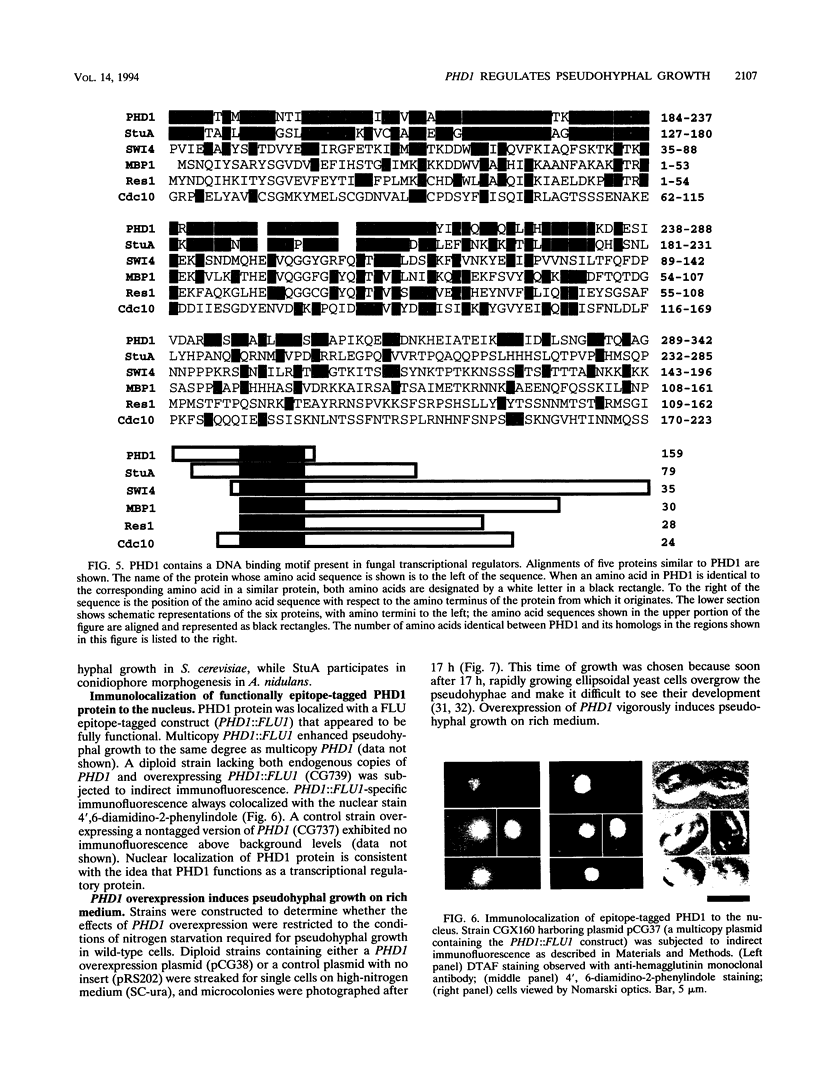
When starved for nitrogen, MATa/MAT alpha cells of the budding yeast Saccharomyces cerevisiae undergo a dimorphic transition to pseudohyphal growth. A visual genetic screen, called PHD (pseudohyphal determinant), for S. cerevisiae pseudohyphal growth mutants was developed. The PHD screen was used to identify seven S. cerevisiae genes that when overexpressed in MATa/MAT alpha cells growing on nitrogen starvation medium cause precocious and unusually vigorous pseudohyphal growth. PHD1, a gene whose overexpression induced invasive pseudohyphal growth on a nutritionally rich medium, was characterized. PHD1 maps to chromosome XI and is predicted to encode a 366-amino-acid protein. PHD1 has a SWI4- and MBP1-like DNA binding motif that is 73% identical over 100 amino acids to a region of Aspergillus nidulans StuA. StuA regulates two pseudohyphal growth-like cell divisions during conidiophore morphogenesis. Epitope-tagged PHD1 was localized to the nucleus by indirect immunofluorescence. These facts suggest that PHD1 may function as a transcriptional regulatory protein. Overexpression of PHD1 in wild-type haploid strains does not induce pseudohyphal growth. Interestingly, PHD1 overexpression enhances pseudohyphal growth in a haploid strain that has the diploid polar budding pattern because of a mutation in the BUD4 gene. In addition, wild-type diploid strains lacking PHD1 undergo pseudohyphal growth when starved for nitrogen. The possible functions of PHD1 in pseudohyphal growth and the uses of the PHD screen to identify morphogenetic regulatory genes from heterologous organisms are discussed.
Full text
PDF









Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Andrews B. J. Gene expression. Dialogue with the cell cycle. Nature. 1992 Jan 30;355(6359):393–394. doi: 10.1038/355393a0. [DOI] [PubMed] [Google Scholar]
- Andrews B. J., Herskowitz I. The yeast SWI4 protein contains a motif present in developmental regulators and is part of a complex involved in cell-cycle-dependent transcription. Nature. 1989 Dec 14;342(6251):830–833. doi: 10.1038/342830a0. [DOI] [PubMed] [Google Scholar]
- Andrews B. J., Mason S. W. Gene expression and the cell cycle: a family affair. Science. 1993 Sep 17;261(5128):1543–1544. doi: 10.1126/science.8372349. [DOI] [PubMed] [Google Scholar]
- Aves S. J., Durkacz B. W., Carr A., Nurse P. Cloning, sequencing and transcriptional control of the Schizosaccharomyces pombe cdc10 'start' gene. EMBO J. 1985 Feb;4(2):457–463. doi: 10.1002/j.1460-2075.1985.tb03651.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banuett F. Ustilago maydis, the delightful blight. Trends Genet. 1992 May;8(5):174–180. doi: 10.1016/0168-9525(92)90220-x. [DOI] [PubMed] [Google Scholar]
- Bender A., Pringle J. R. Multicopy suppression of the cdc24 budding defect in yeast by CDC42 and three newly identified genes including the ras-related gene RSR1. Proc Natl Acad Sci U S A. 1989 Dec;86(24):9976–9980. doi: 10.1073/pnas.86.24.9976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blacketer M. J., Koehler C. M., Coats S. G., Myers A. M., Madaule P. Regulation of dimorphism in Saccharomyces cerevisiae: involvement of the novel protein kinase homolog Elm1p and protein phosphatase 2A. Mol Cell Biol. 1993 Sep;13(9):5567–5581. doi: 10.1128/mcb.13.9.5567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brandriss M. C., Magasanik B. Genetics and physiology of proline utilization in Saccharomyces cerevisiae: enzyme induction by proline. J Bacteriol. 1979 Nov;140(2):498–503. doi: 10.1128/jb.140.2.498-503.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broach J. R., Deschenes R. J. The function of ras genes in Saccharomyces cerevisiae. Adv Cancer Res. 1990;54:79–139. doi: 10.1016/s0065-230x(08)60809-x. [DOI] [PubMed] [Google Scholar]
- Caligiuri M., Beach D. Sct1 functions in partnership with Cdc10 in a transcription complex that activates cell cycle START and inhibits differentiation. Cell. 1993 Feb 26;72(4):607–619. doi: 10.1016/0092-8674(93)90079-6. [DOI] [PubMed] [Google Scholar]
- Chant J., Herskowitz I. Genetic control of bud site selection in yeast by a set of gene products that constitute a morphogenetic pathway. Cell. 1991 Jun 28;65(7):1203–1212. doi: 10.1016/0092-8674(91)90015-q. [DOI] [PubMed] [Google Scholar]
- Chant J., Pringle J. R. Budding and cell polarity in Saccharomyces cerevisiae. Curr Opin Genet Dev. 1991 Oct;1(3):342–350. doi: 10.1016/s0959-437x(05)80298-9. [DOI] [PubMed] [Google Scholar]
- Ciriacy M., Freidel K., Löhning C. Characterization of trans-acting mutations affecting Ty and Ty-mediated transcription in Saccharomyces cerevisiae. Curr Genet. 1991 Dec;20(6):441–448. doi: 10.1007/BF00334769. [DOI] [PubMed] [Google Scholar]
- Clutterbuck A. J. A mutational analysis of conidial development in Aspergillus nidulans. Genetics. 1969 Oct;63(2):317–327. doi: 10.1093/genetics/63.2.317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis L. I., Fink G. R. The NUP1 gene encodes an essential component of the yeast nuclear pore complex. Cell. 1990 Jun 15;61(6):965–978. doi: 10.1016/0092-8674(90)90062-j. [DOI] [PubMed] [Google Scholar]
- Dohrmann P. R., Butler G., Tamai K., Dorland S., Greene J. R., Thiele D. J., Stillman D. J. Parallel pathways of gene regulation: homologous regulators SWI5 and ACE2 differentially control transcription of HO and chitinase. Genes Dev. 1992 Jan;6(1):93–104. doi: 10.1101/gad.6.1.93. [DOI] [PubMed] [Google Scholar]
- Donahue T. F., Cigan A. M. Sequence and structural requirements for efficient translation in yeast. Methods Enzymol. 1990;185:366–372. doi: 10.1016/0076-6879(90)85032-j. [DOI] [PubMed] [Google Scholar]
- Eide D., Guarente L. Increased dosage of a transcriptional activator gene enhances iron-limited growth of Saccharomyces cerevisiae. J Gen Microbiol. 1992 Feb;138(2):347–354. doi: 10.1099/00221287-138-2-347. [DOI] [PubMed] [Google Scholar]
- Epstein C. B., Cross F. R. CLB5: a novel B cyclin from budding yeast with a role in S phase. Genes Dev. 1992 Sep;6(9):1695–1706. doi: 10.1101/gad.6.9.1695. [DOI] [PubMed] [Google Scholar]
- Errede B., Ammerer G. STE12, a protein involved in cell-type-specific transcription and signal transduction in yeast, is part of protein-DNA complexes. Genes Dev. 1989 Sep;3(9):1349–1361. doi: 10.1101/gad.3.9.1349. [DOI] [PubMed] [Google Scholar]
- Estruch F., Carlson M. Increased dosage of the MSN1 gene restores invertase expression in yeast mutants defective in the SNF1 protein kinase. Nucleic Acids Res. 1990 Dec 11;18(23):6959–6964. doi: 10.1093/nar/18.23.6959. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foiani M., Santocanale C., Plevani P., Lucchini G. A single essential gene, PRI2, encodes the large subunit of DNA primase in Saccharomyces cerevisiae. Mol Cell Biol. 1989 Jul;9(7):3081–3087. doi: 10.1128/mcb.9.7.3081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gietz D., St Jean A., Woods R. A., Schiestl R. H. Improved method for high efficiency transformation of intact yeast cells. Nucleic Acids Res. 1992 Mar 25;20(6):1425–1425. doi: 10.1093/nar/20.6.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gimeno C. J., Fink G. R. The logic of cell division in the life cycle of yeast. Science. 1992 Jul 31;257(5070):626–626. doi: 10.1126/science.1496375. [DOI] [PubMed] [Google Scholar]
- Gimeno C. J., Ljungdahl P. O., Styles C. A., Fink G. R. Unipolar cell divisions in the yeast S. cerevisiae lead to filamentous growth: regulation by starvation and RAS. Cell. 1992 Mar 20;68(6):1077–1090. doi: 10.1016/0092-8674(92)90079-r. [DOI] [PubMed] [Google Scholar]
- Grenson M., Mousset M., Wiame J. M., Bechet J. Multiplicity of the amino acid permeases in Saccharomyces cerevisiae. I. Evidence for a specific arginine-transporting system. Biochim Biophys Acta. 1966 Oct 31;127(2):325–338. doi: 10.1016/0304-4165(66)90387-4. [DOI] [PubMed] [Google Scholar]
- Haltiner M., Kempe T., Tjian R. A novel strategy for constructing clustered point mutations. Nucleic Acids Res. 1985 Feb 11;13(3):1015–1025. doi: 10.1093/nar/13.3.1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartwell L. H. Saccharomyces cerevisiae cell cycle. Bacteriol Rev. 1974 Jun;38(2):164–198. doi: 10.1128/br.38.2.164-198.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hoffman C. S., Winston F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene. 1987;57(2-3):267–272. doi: 10.1016/0378-1119(87)90131-4. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koch C., Moll T., Neuberg M., Ahorn H., Nasmyth K. A role for the transcription factors Mbp1 and Swi4 in progression from G1 to S phase. Science. 1993 Sep 17;261(5128):1551–1557. doi: 10.1126/science.8372350. [DOI] [PubMed] [Google Scholar]
- Kolodziej P. A., Young R. A. Epitope tagging and protein surveillance. Methods Enzymol. 1991;194:508–519. doi: 10.1016/0076-6879(91)94038-e. [DOI] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Kuranda M. J., Robbins P. W. Chitinase is required for cell separation during growth of Saccharomyces cerevisiae. J Biol Chem. 1991 Oct 15;266(29):19758–19767. [PubMed] [Google Scholar]
- Lee K. S., Hines L. K., Levin D. E. A pair of functionally redundant yeast genes (PPZ1 and PPZ2) encoding type 1-related protein phosphatases function within the PKC1-mediated pathway. Mol Cell Biol. 1993 Sep;13(9):5843–5853. doi: 10.1128/mcb.13.9.5843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lew D. J., Reed S. I. Morphogenesis in the yeast cell cycle: regulation by Cdc28 and cyclins. J Cell Biol. 1993 Mar;120(6):1305–1320. doi: 10.1083/jcb.120.6.1305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H., Styles C. A., Fink G. R. Elements of the yeast pheromone response pathway required for filamentous growth of diploids. Science. 1993 Dec 10;262(5140):1741–1744. doi: 10.1126/science.8259520. [DOI] [PubMed] [Google Scholar]
- Ljungdahl P. O., Gimeno C. J., Styles C. A., Fink G. R. SHR3: a novel component of the secretory pathway specifically required for localization of amino acid permeases in yeast. Cell. 1992 Oct 30;71(3):463–478. doi: 10.1016/0092-8674(92)90515-e. [DOI] [PubMed] [Google Scholar]
- Maresca B., Kobayashi G. S. Dimorphism in Histoplasma capsulatum: a model for the study of cell differentiation in pathogenic fungi. Microbiol Rev. 1989 Jun;53(2):186–209. doi: 10.1128/mr.53.2.186-209.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medoff G., Sacco M., Maresca B., Schlessinger D., Painter A., Kobayashi G. S., Carratu L. Irreversible block of the mycelial-to-yeast phase transition of Histoplasma capsulatum. Science. 1986 Jan 31;231(4737):476–479. doi: 10.1126/science.3001938. [DOI] [PubMed] [Google Scholar]
- Mermod N., O'Neill E. A., Kelly T. J., Tjian R. The proline-rich transcriptional activator of CTF/NF-I is distinct from the replication and DNA binding domain. Cell. 1989 Aug 25;58(4):741–753. doi: 10.1016/0092-8674(89)90108-6. [DOI] [PubMed] [Google Scholar]
- Miller K. Y., Toennis T. M., Adams T. H., Miller B. L. Isolation and transcriptional characterization of a morphological modifier: the Aspergillus nidulans stunted (stuA) gene. Mol Gen Genet. 1991 Jun;227(2):285–292. doi: 10.1007/BF00259682. [DOI] [PubMed] [Google Scholar]
- Miller K. Y., Wu J., Miller B. L. StuA is required for cell pattern formation in Aspergillus. Genes Dev. 1992 Sep;6(9):1770–1782. doi: 10.1101/gad.6.9.1770. [DOI] [PubMed] [Google Scholar]
- Mitchell P. J., Tjian R. Transcriptional regulation in mammalian cells by sequence-specific DNA binding proteins. Science. 1989 Jul 28;245(4916):371–378. doi: 10.1126/science.2667136. [DOI] [PubMed] [Google Scholar]
- Molenaar C. M., Woudt L. P., Jansen A. E., Mager W. H., Planta R. J., Donovan D. M., Pearson N. J. Structure and organization of two linked ribosomal protein genes in yeast. Nucleic Acids Res. 1984 Oct 11;12(19):7345–7358. doi: 10.1093/nar/12.19.7345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nasmyth K. Control of the yeast cell cycle by the Cdc28 protein kinase. Curr Opin Cell Biol. 1993 Apr;5(2):166–179. doi: 10.1016/0955-0674(93)90099-c. [DOI] [PubMed] [Google Scholar]
- Primig M., Sockanathan S., Auer H., Nasmyth K. Anatomy of a transcription factor important for the start of the cell cycle in Saccharomyces cerevisiae. Nature. 1992 Aug 13;358(6387):593–597. doi: 10.1038/358593a0. [DOI] [PubMed] [Google Scholar]
- Ramer S. W., Elledge S. J., Davis R. W. Dominant genetics using a yeast genomic library under the control of a strong inducible promoter. Proc Natl Acad Sci U S A. 1992 Dec 1;89(23):11589–11593. doi: 10.1073/pnas.89.23.11589. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riles L., Dutchik J. E., Baktha A., McCauley B. K., Thayer E. C., Leckie M. P., Braden V. V., Depke J. E., Olson M. V. Physical maps of the six smallest chromosomes of Saccharomyces cerevisiae at a resolution of 2.6 kilobase pairs. Genetics. 1993 May;134(1):81–150. doi: 10.1093/genetics/134.1.81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rine J. Gene overexpression in studies of Saccharomyces cerevisiae. Methods Enzymol. 1991;194:239–251. doi: 10.1016/0076-6879(91)94019-9. [DOI] [PubMed] [Google Scholar]
- Ruggieri R., Bender A., Matsui Y., Powers S., Takai Y., Pringle J. R., Matsumoto K. RSR1, a ras-like gene homologous to Krev-1 (smg21A/rap1A): role in the development of cell polarity and interactions with the Ras pathway in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Feb;12(2):758–766. doi: 10.1128/mcb.12.2.758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SCHERR G. H., WEAVER R. H. The dimorphism phenomenon in yeasts. Bacteriol Rev. 1953 Mar;17(1):51–92. doi: 10.1128/br.17.1.51-92.1953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwob E., Nasmyth K. CLB5 and CLB6, a new pair of B cyclins involved in DNA replication in Saccharomyces cerevisiae. Genes Dev. 1993 Jul;7(7A):1160–1175. doi: 10.1101/gad.7.7a.1160. [DOI] [PubMed] [Google Scholar]
- Shepherd M. G. Morphogenetic transformation of fungi. Curr Top Med Mycol. 1988;2:278–304. doi: 10.1007/978-1-4612-3730-3_8. [DOI] [PubMed] [Google Scholar]
- Siddiqui A. H., Brandriss M. C. A regulatory region responsible for proline-specific induction of the yeast PUT2 gene is adjacent to its TATA box. Mol Cell Biol. 1988 Nov;8(11):4634–4641. doi: 10.1128/mcb.8.11.4634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka K., Okazaki K., Okazaki N., Ueda T., Sugiyama A., Nojima H., Okayama H. A new cdc gene required for S phase entry of Schizosaccharomyces pombe encodes a protein similar to the cdc 10+ and SWI4 gene products. EMBO J. 1992 Dec;11(13):4923–4932. doi: 10.1002/j.1460-2075.1992.tb05599.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Timberlake W. E. Molecular genetics of Aspergillus development. Annu Rev Genet. 1990;24:5–36. doi: 10.1146/annurev.ge.24.120190.000253. [DOI] [PubMed] [Google Scholar]
- Timberlake W. E. Temporal and spatial controls of Aspergillus development. Curr Opin Genet Dev. 1991 Oct;1(3):351–357. doi: 10.1016/s0959-437x(05)80299-0. [DOI] [PubMed] [Google Scholar]
- Van Arsdell S. W., Stetler G. L., Thorner J. The yeast repeated element sigma contains a hormone-inducible promoter. Mol Cell Biol. 1987 Feb;7(2):749–759. doi: 10.1128/mcb.7.2.749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. Production of single-stranded plasmid DNA. Methods Enzymol. 1987;153:3–11. doi: 10.1016/0076-6879(87)53044-0. [DOI] [PubMed] [Google Scholar]
- Wada K., Wada Y., Ishibashi F., Gojobori T., Ikemura T. Codon usage tabulated from the GenBank genetic sequence data. Nucleic Acids Res. 1992 May 11;20 (Suppl):2111–2118. doi: 10.1093/nar/20.suppl.2111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson I. A., Niman H. L., Houghten R. A., Cherenson A. R., Connolly M. L., Lerner R. A. The structure of an antigenic determinant in a protein. Cell. 1984 Jul;37(3):767–778. doi: 10.1016/0092-8674(84)90412-4. [DOI] [PubMed] [Google Scholar]
- Wright R. M., Repine T., Repine J. E. Reversible pseudohyphal growth in haploid Saccharomyces cerevisiae is an aerobic process. Curr Genet. 1993 May-Jun;23(5-6):388–391. doi: 10.1007/BF00312623. [DOI] [PubMed] [Google Scholar]