Abstract
Xenopus UBF (xUBF) is a transcription factor for RNA polymerase I which contains multiple DNA-binding motifs. These include a short basic region adjacent to a dimer motif plus five high-mobility-group (HMG) boxes. All of these DNA-binding motifs exhibit low sequence specificity, whether assayed singly or together. In contrast, the HMG boxes recognize DNA structure that is formed when two double helices are crossed over each other. HMG box 1, in particular, requires association of two double helices before it will bind and, either by itself or in the context of the intact protein, will loop DNA and organize it into higher-order structures. We discuss how this mode of binding affects the function of xUBF as a transcription factor.
Full text
PDF


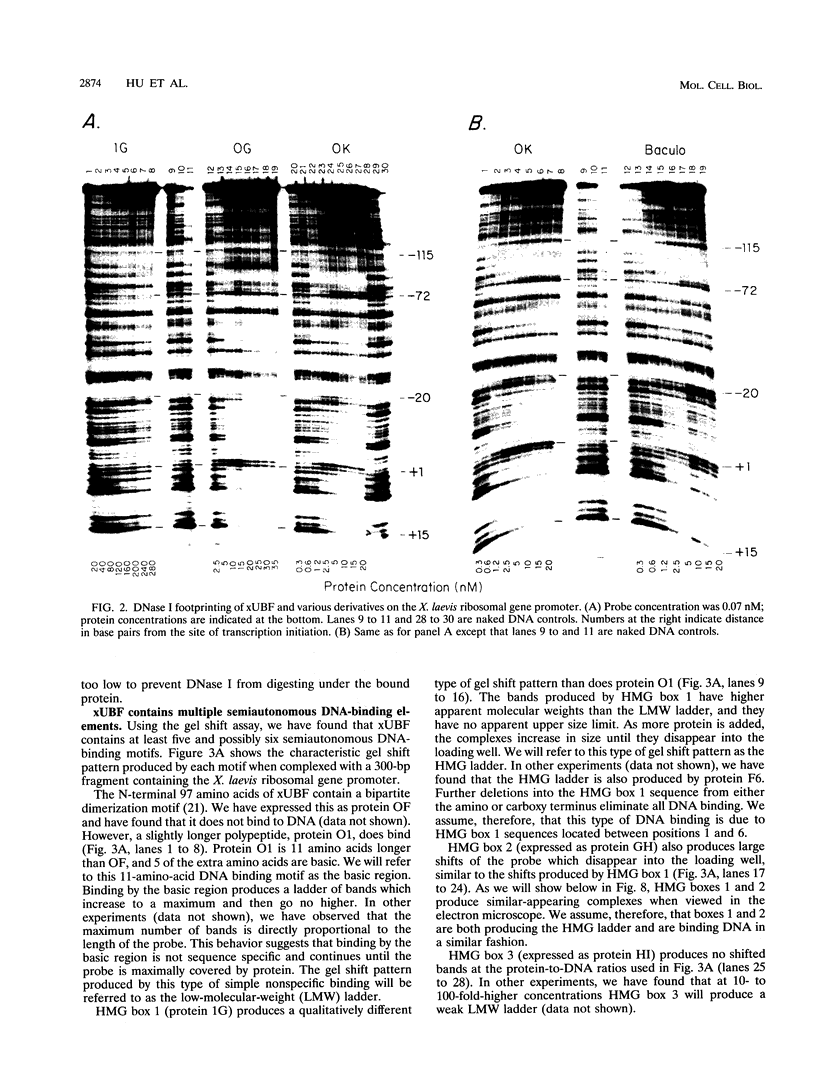








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bachvarov D., Moss T. The RNA polymerase I transcription factor xUBF contains 5 tandemly repeated HMG homology boxes. Nucleic Acids Res. 1991 May 11;19(9):2331–2335. doi: 10.1093/nar/19.9.2331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bell S. P., Jantzen H. M., Tjian R. Assembly of alternative multiprotein complexes directs rRNA promoter selectivity. Genes Dev. 1990 Jun;4(6):943–954. doi: 10.1101/gad.4.6.943. [DOI] [PubMed] [Google Scholar]
- Bell S. P., Learned R. M., Jantzen H. M., Tjian R. Functional cooperativity between transcription factors UBF1 and SL1 mediates human ribosomal RNA synthesis. Science. 1988 Sep 2;241(4870):1192–1197. doi: 10.1126/science.3413483. [DOI] [PubMed] [Google Scholar]
- Choe S. Y., Schultz M. C., Reeder R. H. In vitro definition of the yeast RNA polymerase I promoter. Nucleic Acids Res. 1992 Jan 25;20(2):279–285. doi: 10.1093/nar/20.2.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Comai L., Tanese N., Tjian R. The TATA-binding protein and associated factors are integral components of the RNA polymerase I transcription factor, SL1. Cell. 1992 Mar 6;68(5):965–976. doi: 10.1016/0092-8674(92)90039-f. [DOI] [PubMed] [Google Scholar]
- Cormack B. P., Struhl K. The TATA-binding protein is required for transcription by all three nuclear RNA polymerases in yeast cells. Cell. 1992 May 15;69(4):685–696. doi: 10.1016/0092-8674(92)90232-2. [DOI] [PubMed] [Google Scholar]
- Ferrari S., Harley V. R., Pontiggia A., Goodfellow P. N., Lovell-Badge R., Bianchi M. E. SRY, like HMG1, recognizes sharp angles in DNA. EMBO J. 1992 Dec;11(12):4497–4506. doi: 10.1002/j.1460-2075.1992.tb05551.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher R. P., Lisowsky T., Parisi M. A., Clayton D. A. DNA wrapping and bending by a mitochondrial high mobility group-like transcriptional activator protein. J Biol Chem. 1992 Feb 15;267(5):3358–3367. [PubMed] [Google Scholar]
- Guimond A., Moss T. Variants of the Xenopus laevis ribosomal transcription factor xUBF are developmentally regulated by differential splicing. Nucleic Acids Res. 1992 Jul 11;20(13):3361–3366. doi: 10.1093/nar/20.13.3361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamada H., Bustin M. Hierarchy of binding sites for chromosomal proteins HMG 1 and 2 in supercoiled deoxyribonucleic acid. Biochemistry. 1985 Mar 12;24(6):1428–1433. doi: 10.1021/bi00327a022. [DOI] [PubMed] [Google Scholar]
- Hisatake K., Nishimura T., Maeda Y., Hanada K., Song C. Z., Muramatsu M. Cloning and structural analysis of cDNA and the gene for mouse transcription factor UBF. Nucleic Acids Res. 1991 Sep 11;19(17):4631–4637. doi: 10.1093/nar/19.17.4631. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jantzen H. M., Admon A., Bell S. P., Tjian R. Nucleolar transcription factor hUBF contains a DNA-binding motif with homology to HMG proteins. Nature. 1990 Apr 26;344(6269):830–836. doi: 10.1038/344830a0. [DOI] [PubMed] [Google Scholar]
- Jantzen H. M., Chow A. M., King D. S., Tjian R. Multiple domains of the RNA polymerase I activator hUBF interact with the TATA-binding protein complex hSL1 to mediate transcription. Genes Dev. 1992 Oct;6(10):1950–1963. doi: 10.1101/gad.6.10.1950. [DOI] [PubMed] [Google Scholar]
- Kerrigan L. A., Kadonaga J. T. Periodic binding of individual core histones to DNA: inadvertent purification of the core histone H2B as a putative enhancer-binding factor. Nucleic Acids Res. 1992 Dec 25;20(24):6673–6680. doi: 10.1093/nar/20.24.6673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuhn A., Grummt I. Dual role of the nucleolar transcription factor UBF: trans-activator and antirepressor. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7340–7344. doi: 10.1073/pnas.89.16.7340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Leblanc B., Read C., Moss T. Recognition of the Xenopus ribosomal core promoter by the transcription factor xUBF involves multiple HMG box domains and leads to an xUBF interdomain interaction. EMBO J. 1993 Feb;12(2):513–525. doi: 10.1002/j.1460-2075.1993.tb05683.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lilley D. M. DNA--protein interactions. HMG has DNA wrapped up. Nature. 1992 May 28;357(6376):282–283. doi: 10.1038/357282a0. [DOI] [PubMed] [Google Scholar]
- Maeda Y., Hisatake K., Kondo T., Hanada K., Song C. Z., Nishimura T., Muramatsu M. Mouse rRNA gene transcription factor mUBF requires both HMG-box1 and an acidic tail for nucleolar accumulation: molecular analysis of the nucleolar targeting mechanism. EMBO J. 1992 Oct;11(10):3695–3704. doi: 10.1002/j.1460-2075.1992.tb05454.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McStay B., Frazier M. W., Reeder R. H. xUBF contains a novel dimerization domain essential for RNA polymerase I transcription. Genes Dev. 1991 Nov;5(11):1957–1968. doi: 10.1101/gad.5.11.1957. [DOI] [PubMed] [Google Scholar]
- McStay B., Hu C. H., Pikaard C. S., Reeder R. H. xUBF and Rib 1 are both required for formation of a stable polymerase I promoter complex in X. laevis. EMBO J. 1991 Aug;10(8):2297–2303. doi: 10.1002/j.1460-2075.1991.tb07766.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McStay B., Reeder R. H. A termination site for Xenopus RNA polymerase I also acts as an element of an adjacent promoter. Cell. 1986 Dec 26;47(6):913–920. doi: 10.1016/0092-8674(86)90806-8. [DOI] [PubMed] [Google Scholar]
- O'Mahony D. J., Rothblum L. I. Identification of two forms of the RNA polymerase I transcription factor UBF. Proc Natl Acad Sci U S A. 1991 Apr 15;88(8):3180–3184. doi: 10.1073/pnas.88.8.3180. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Mahony D. J., Smith S. D., Xie W., Rothblum L. I. Analysis of the phosphorylation, DNA-binding and dimerization properties of the RNA polymerase I transcription factors UBF1 and UBF2. Nucleic Acids Res. 1992 Mar 25;20(6):1301–1308. doi: 10.1093/nar/20.6.1301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Mahony D. J., Xie W. Q., Smith S. D., Singer H. A., Rothblum L. I. Differential phosphorylation and localization of the transcription factor UBF in vivo in response to serum deprivation. In vitro dephosphorylation of UBF reduces its transactivation properties. J Biol Chem. 1992 Jan 5;267(1):35–38. [PubMed] [Google Scholar]
- Parisi M. A., Clayton D. A. Similarity of human mitochondrial transcription factor 1 to high mobility group proteins. Science. 1991 May 17;252(5008):965–969. doi: 10.1126/science.2035027. [DOI] [PubMed] [Google Scholar]
- Pikaard C. S., McStay B., Schultz M. C., Bell S. P., Reeder R. H. The Xenopus ribosomal gene enhancers bind an essential polymerase I transcription factor, xUBF. Genes Dev. 1989 Nov;3(11):1779–1788. doi: 10.1101/gad.3.11.1779. [DOI] [PubMed] [Google Scholar]
- Putnam C. D., Pikaard C. S. Cooperative binding of the Xenopus RNA polymerase I transcription factor xUBF to repetitive ribosomal gene enhancers. Mol Cell Biol. 1992 Nov;12(11):4970–4980. doi: 10.1128/mcb.12.11.4970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roussel P., André C., Masson C., Géraud G., Hernandez-Verdun D. Localization of the RNA polymerase I transcription factor hUBF during the cell cycle. J Cell Sci. 1993 Feb;104(Pt 2):327–337. doi: 10.1242/jcs.104.2.327. [DOI] [PubMed] [Google Scholar]
- Schultz M. C., Reeder R. H., Hahn S. Variants of the TATA-binding protein can distinguish subsets of RNA polymerase I, II, and III promoters. Cell. 1992 May 15;69(4):697–702. doi: 10.1016/0092-8674(92)90233-3. [DOI] [PubMed] [Google Scholar]
- Simpson R. T. Nucleosome positioning: occurrence, mechanisms, and functional consequences. Prog Nucleic Acid Res Mol Biol. 1991;40:143–184. doi: 10.1016/s0079-6603(08)60841-7. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Travis A., Amsterdam A., Belanger C., Grosschedl R. LEF-1, a gene encoding a lymphoid-specific protein with an HMG domain, regulates T-cell receptor alpha enhancer function [corrected]. Genes Dev. 1991 May;5(5):880–894. doi: 10.1101/gad.5.5.880. [DOI] [PubMed] [Google Scholar]
- Voit R., Schnapp A., Kuhn A., Rosenbauer H., Hirschmann P., Stunnenberg H. G., Grummt I. The nucleolar transcription factor mUBF is phosphorylated by casein kinase II in the C-terminal hyperacidic tail which is essential for transactivation. EMBO J. 1992 Jun;11(6):2211–2218. doi: 10.1002/j.1460-2075.1992.tb05280.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weir H. M., Kraulis P. J., Hill C. S., Raine A. R., Laue E. D., Thomas J. O. Structure of the HMG box motif in the B-domain of HMG1. EMBO J. 1993 Apr;12(4):1311–1319. doi: 10.1002/j.1460-2075.1993.tb05776.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zechiedrich E. L., Osheroff N. Eukaryotic topoisomerases recognize nucleic acid topology by preferentially interacting with DNA crossovers. EMBO J. 1990 Dec;9(13):4555–4562. doi: 10.1002/j.1460-2075.1990.tb07908.x. [DOI] [PMC free article] [PubMed] [Google Scholar]