Abstract
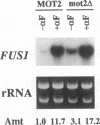
The STE4 gene encodes the beta subunit of a heterotrimeric G protein that is an essential component of the pheromone signal transduction pathway. To identify downstream component(s) of Ste4, we sought pseudo-revertants that restored mating competence to ste4 mutants. The suppressor mot2 was isolated as a recessive mutation that restored conjugational competence to a temperature-sensitive ste4 mutant and simultaneously conferred a temperature-sensitive growth phenotype. The MOT2 gene encodes a putative zinc finger protein, the deletion of which resulted in temperature-sensitive growth, increased expression of FUS1 in the absence of pheromones, and suppression of a deletion of the alpha-factor receptor. On the other hand, sterility resulting from deletion of STE4 was not suppressed by the mot2 deletion. These phenotypes are similar to those associated with temperature-sensitive mutations in CDC36 and CDC39, which are proposed to encode general negative regulators of transcription rather than factors involved in the pheromone response pathway. Deletion of MOT2 also caused increased transcription of unrelated genes such as GAL7 and PHO84. Overexpression of MOT2 suppresses the growth defect of temperature-sensitive mutations in CDC36 and CDC39. These observations suggest that Mot2 functions as a general negative regulator of transcription in the same processes as Cdc36 and Cdc39.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aebi M., Clark M. W., Vijayraghavan U., Abelson J. A yeast mutant, PRP20, altered in mRNA metabolism and maintenance of the nuclear structure, is defective in a gene homologous to the human gene RCC1 which is involved in the control of chromosome condensation. Mol Gen Genet. 1990 Oct;224(1):72–80. doi: 10.1007/BF00259453. [DOI] [PubMed] [Google Scholar]
- Belhumeur P., Lee A., Tam R., DiPaolo T., Fortin N., Clark M. W. GSP1 and GSP2, genetic suppressors of the prp20-1 mutant in Saccharomyces cerevisiae: GTP-binding proteins involved in the maintenance of nuclear organization. Mol Cell Biol. 1993 Apr;13(4):2152–2161. doi: 10.1128/mcb.13.4.2152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg J. M. Zinc fingers and other metal-binding domains. Elements for interactions between macromolecules. J Biol Chem. 1990 Apr 25;265(12):6513–6516. [PubMed] [Google Scholar]
- Blinder D., Bouvier S., Jenness D. D. Constitutive mutants in the yeast pheromone response: ordered function of the gene products. Cell. 1989 Feb 10;56(3):479–486. doi: 10.1016/0092-8674(89)90250-x. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Bun-Ya M., Nishimura M., Harashima S., Oshima Y. The PHO84 gene of Saccharomyces cerevisiae encodes an inorganic phosphate transporter. Mol Cell Biol. 1991 Jun;11(6):3229–3238. doi: 10.1128/mcb.11.6.3229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burkholder A. C., Hartwell L. H. The yeast alpha-factor receptor: structural properties deduced from the sequence of the STE2 gene. Nucleic Acids Res. 1985 Dec 9;13(23):8463–8475. doi: 10.1093/nar/13.23.8463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark K. L., Ohtsubo M., Nishimoto T., Goebl M., Sprague G. F., Jr The yeast SRM1 protein and human RCC1 protein share analogous functions. Cell Regul. 1991 Oct;2(10):781–792. doi: 10.1091/mbc.2.10.781. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark K. L., Sprague G. F., Jr Yeast pheromone response pathway: characterization of a suppressor that restores mating to receptorless mutants. Mol Cell Biol. 1989 Jun;9(6):2682–2694. doi: 10.1128/mcb.9.6.2682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collart M. A., Struhl K. CDC39, an essential nuclear protein that negatively regulates transcription and differentially affects the constitutive and inducible HIS3 promoters. EMBO J. 1993 Jan;12(1):177–186. doi: 10.1002/j.1460-2075.1993.tb05643.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dietzel C., Kurjan J. The yeast SCG1 gene: a G alpha-like protein implicated in the a- and alpha-factor response pathway. Cell. 1987 Sep 25;50(7):1001–1010. doi: 10.1016/0092-8674(87)90166-8. [DOI] [PubMed] [Google Scholar]
- Duronio R. J., Towler D. A., Heuckeroth R. O., Gordon J. I. Disruption of the yeast N-myristoyl transferase gene causes recessive lethality. Science. 1989 Feb 10;243(4892):796–800. doi: 10.1126/science.2644694. [DOI] [PubMed] [Google Scholar]
- Fleischmann M., Clark M. W., Forrester W., Wickens M., Nishimoto T., Aebi M. Analysis of yeast prp20 mutations and functional complementation by the human homologue RCC1, a protein involved in the control of chromosome condensation. Mol Gen Genet. 1991 Jul;227(3):417–423. doi: 10.1007/BF00273932. [DOI] [PubMed] [Google Scholar]
- Gietz R. D., Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene. 1988 Dec 30;74(2):527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- Hagen D. C., McCaffrey G., Sprague G. F., Jr Evidence the yeast STE3 gene encodes a receptor for the peptide pheromone a factor: gene sequence and implications for the structure of the presumed receptor. Proc Natl Acad Sci U S A. 1986 Mar;83(5):1418–1422. doi: 10.1073/pnas.83.5.1418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagen D. C., McCaffrey G., Sprague G. F., Jr Pheromone response elements are necessary and sufficient for basal and pheromone-induced transcription of the FUS1 gene of Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jun;11(6):2952–2961. doi: 10.1128/mcb.11.6.2952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jahng K. Y., Ferguson J., Reed S. I. Mutations in a gene encoding the alpha subunit of a Saccharomyces cerevisiae G protein indicate a role in mating pheromone signaling. Mol Cell Biol. 1988 Jun;8(6):2484–2493. doi: 10.1128/mcb.8.6.2484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kadowaki T., Zhao Y., Tartakoff A. M. A conditional yeast mutant deficient in mRNA transport from nucleus to cytoplasm. Proc Natl Acad Sci U S A. 1992 Mar 15;89(6):2312–2316. doi: 10.1073/pnas.89.6.2312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kronstad J. W., Holly J. A., MacKay V. L. A yeast operator overlaps an upstream activation site. Cell. 1987 Jul 31;50(3):369–377. doi: 10.1016/0092-8674(87)90491-0. [DOI] [PubMed] [Google Scholar]
- Marsh L., Neiman A. M., Herskowitz I. Signal transduction during pheromone response in yeast. Annu Rev Cell Biol. 1991;7:699–728. doi: 10.1146/annurev.cb.07.110191.003411. [DOI] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Miyajima I., Nakafuku M., Nakayama N., Brenner C., Miyajima A., Kaibuchi K., Arai K., Kaziro Y., Matsumoto K. GPA1, a haploid-specific essential gene, encodes a yeast homolog of mammalian G protein which may be involved in mating factor signal transduction. Cell. 1987 Sep 25;50(7):1011–1019. doi: 10.1016/0092-8674(87)90167-x. [DOI] [PubMed] [Google Scholar]
- Morrison D. A. Transformation in Escherichia coli: cryogenic preservation of competent cells. J Bacteriol. 1977 Oct;132(1):349–351. doi: 10.1128/jb.132.1.349-351.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakayama N., Miyajima A., Arai K. Nucleotide sequences of STE2 and STE3, cell type-specific sterile genes from Saccharomyces cerevisiae. EMBO J. 1985 Oct;4(10):2643–2648. doi: 10.1002/j.1460-2075.1985.tb03982.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neiman A. M., Chang F., Komachi K., Herskowitz I. CDC36 and CDC39 are negative elements in the signal transduction pathway of yeast. Cell Regul. 1990 Apr;1(5):391–401. doi: 10.1091/mbc.1.5.391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ninomiya-Tsuji J., Nomoto S., Yasuda H., Reed S. I., Matsumoto K. Cloning of a human cDNA encoding a CDC2-related kinase by complementation of a budding yeast cdc28 mutation. Proc Natl Acad Sci U S A. 1991 Oct 15;88(20):9006–9010. doi: 10.1073/pnas.88.20.9006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nogi Y., Fukasawa T. Nucleotide sequence of the transcriptional initiation region of the yeast GAL7 gene. Nucleic Acids Res. 1983 Dec 20;11(24):8555–8568. doi: 10.1093/nar/11.24.8555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohtsubo M., Yoshida T., Seino H., Nishitani H., Clark K. L., Sprague G. F., Jr, Frasch M., Nishimoto T. Mutation of the hamster cell cycle gene RCC1 is complemented by the homologous genes of Drosophila and S.cerevisiae. EMBO J. 1991 May;10(5):1265–1273. doi: 10.1002/j.1460-2075.1991.tb08068.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parent S. A., Fenimore C. M., Bostian K. A. Vector systems for the expression, analysis and cloning of DNA sequences in S. cerevisiae. Yeast. 1985 Dec;1(2):83–138. doi: 10.1002/yea.320010202. [DOI] [PubMed] [Google Scholar]
- Reed S. I., Ferguson J., Jahng K. Y. Isolation and characterization of two genes encoding yeast mating pheromone signaling elements: CDC72 and CDC73. Cold Spring Harb Symp Quant Biol. 1988;53(Pt 2):621–627. doi: 10.1101/sqb.1988.053.01.071. [DOI] [PubMed] [Google Scholar]
- Reed S. I. The selection of S. cerevisiae mutants defective in the start event of cell division. Genetics. 1980 Jul;95(3):561–577. doi: 10.1093/genetics/95.3.561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose M. D., Novick P., Thomas J. H., Botstein D., Fink G. R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60(2-3):237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stone D. E., Cole G. M., de Barros Lopes M., Goebl M., Reed S. I. N-myristoylation is required for function of the pheromone-responsive G alpha protein of yeast: conditional activation of the pheromone response by a temperature-sensitive N-myristoyl transferase. Genes Dev. 1991 Nov;5(11):1969–1981. doi: 10.1101/gad.5.11.1969. [DOI] [PubMed] [Google Scholar]
- Trueheart J., Boeke J. D., Fink G. R. Two genes required for cell fusion during yeast conjugation: evidence for a pheromone-induced surface protein. Mol Cell Biol. 1987 Jul;7(7):2316–2328. doi: 10.1128/mcb.7.7.2316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whiteway M., Hougan L., Dignard D., Thomas D. Y., Bell L., Saari G. C., Grant F. J., O'Hara P., MacKay V. L. The STE4 and STE18 genes of yeast encode potential beta and gamma subunits of the mating factor receptor-coupled G protein. Cell. 1989 Feb 10;56(3):467–477. doi: 10.1016/0092-8674(89)90249-3. [DOI] [PubMed] [Google Scholar]
- de Barros Lopes M., Ho J. Y., Reed S. I. Mutations in cell division cycle genes CDC36 and CDC39 activate the Saccharomyces cerevisiae mating pheromone response pathway. Mol Cell Biol. 1990 Jun;10(6):2966–2972. doi: 10.1128/mcb.10.6.2966. [DOI] [PMC free article] [PubMed] [Google Scholar]