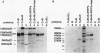Abstract
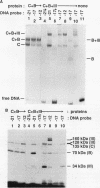
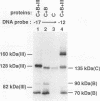
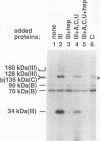
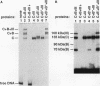
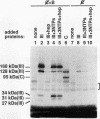
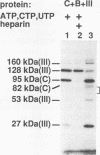
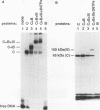
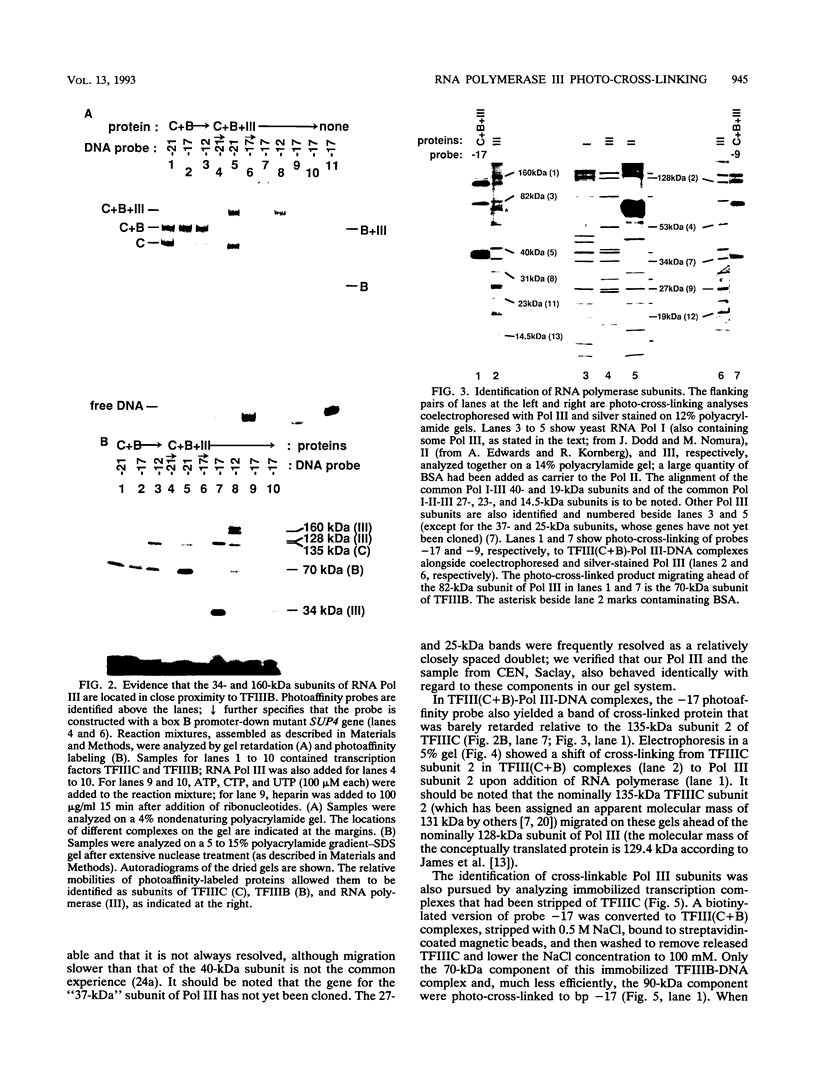
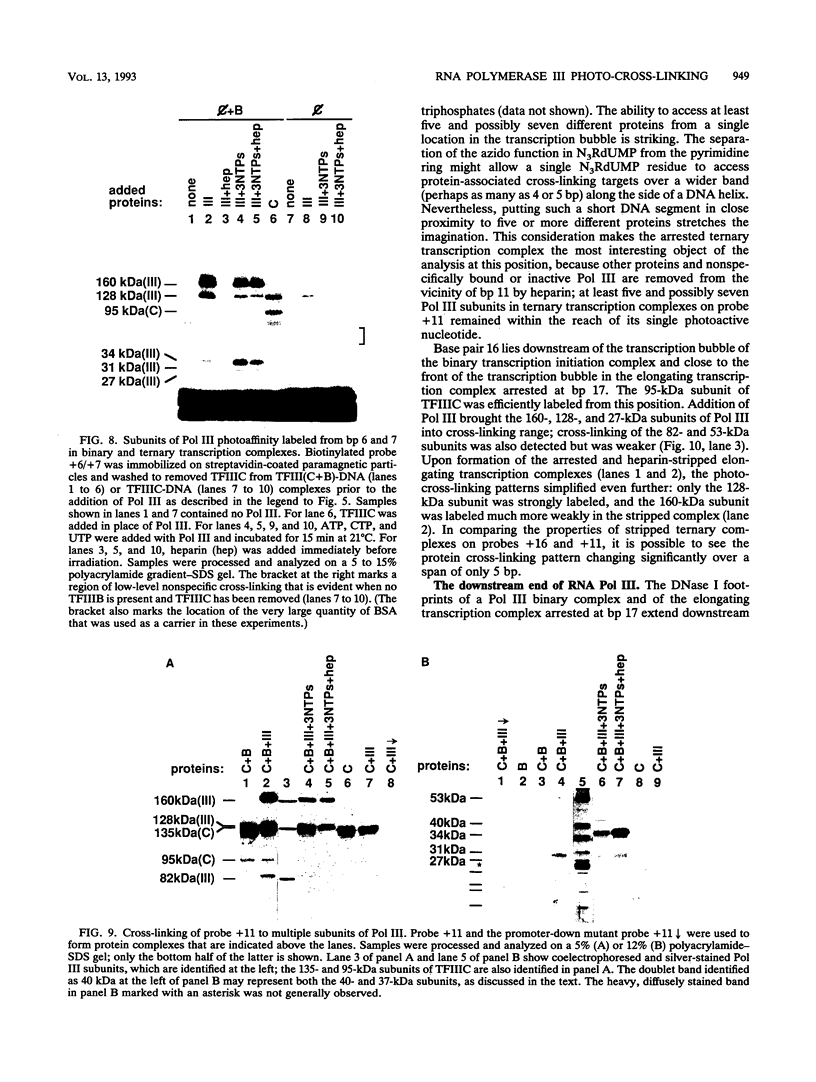
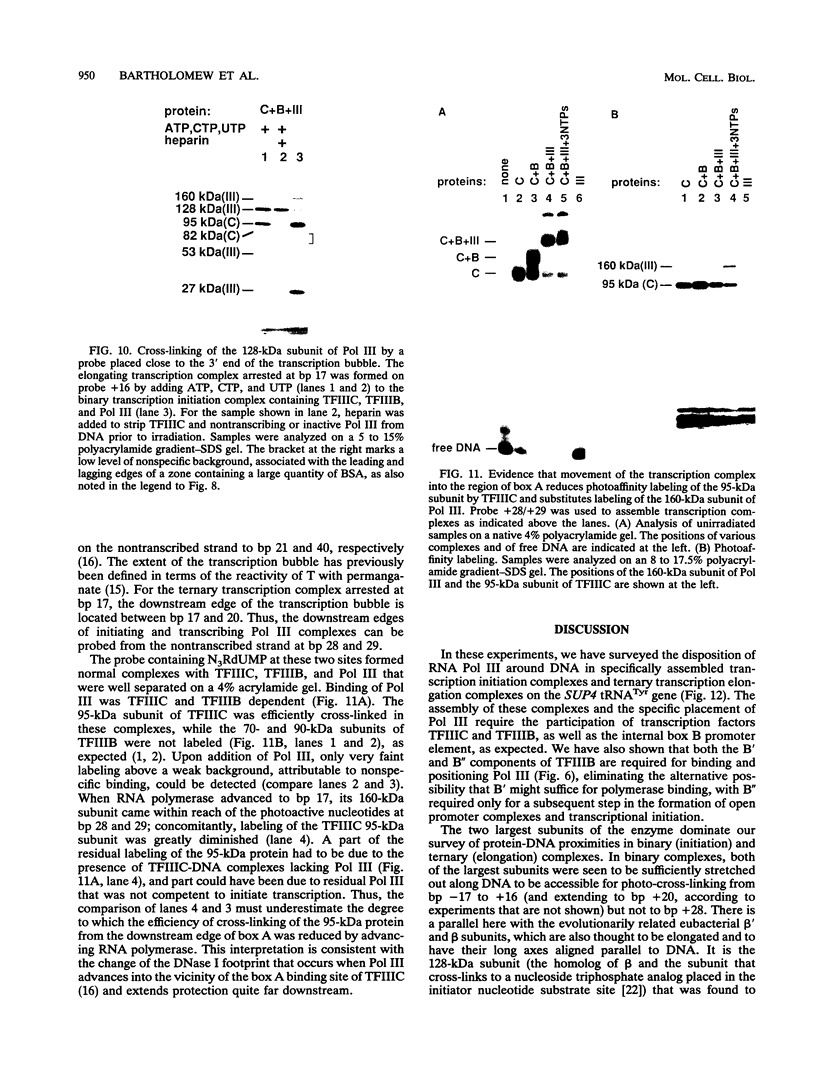
A photo-cross-linking method has been used to map the subunits of Saccharomyces cerevisiae RNA polymerase (Pol) III with respect to DNA in binary (preinitiation) and ternary (RNA-elongating) transcription complexes. Transcription factor- and Pol III-containing complexes have been assembled on S. cerevisiae SUP4 tRNA(Tyr) gene probes containing the photoactive nucleotide 5-[N-(p-azidobenzoyl)-3-aminoallyl]-dUMP in different specified positions. Covalent DNA-protein linkages form upon irradiation of these complexes, and the Pol III subunits that are cross-linked to individual positions in the SUP4 tRNA gene have been identified. RNA Pol III cross-linking has been shown to require the box B downstream promoter element of the tRNA gene and the presence of transcription factor TFIIIB. Further proof of specificity has been provided by demonstrating that particular Pol III subunits move out of the range of upstream-placed photoactive nucleotides, and that others move into the range of downstream-placed photoactive nucleotides, as a consequence of initiating and elongating RNA chains. Binding and specific placement of Pol III have also been shown to require both the B' and the B" components of TFIIIB. Nine Pol III subunits are cross-linked from different positions of the SUP4 tRNA gene's nontranscribed strand. In binary transcription complexes, the two largest Pol III subunits are accessible to photo-cross-linking over the entire stretch of the DNase I footprint. The 27- and 34-kDa Pol III subunits are also relatively extended along DNA; its upstream projection makes the 34-kDa subunit a candidate for interaction with TFIIIB, while the 27-kDa subunit is accessible to photo-cross-linking from the leading edge of the Pol III binding site. Several subunits, including the 82- and 53-kDa subunits in binary transcription complexes, are relatively localized in their accessibility to cross-linking. Multiple Pol III subunits are accessible to specific cross-linking from a single photoactive nucleotide in the middle of the transcription bubble of an arrested ternary transcription complex. It is suggested that this precisely placed transcription complex comprises a dynamic ensemble of structural states rather than a single perfectly constrained entity.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bartholomew B., Kassavetis G. A., Braun B. R., Geiduschek E. P. The subunit structure of Saccharomyces cerevisiae transcription factor IIIC probed with a novel photocrosslinking reagent. EMBO J. 1990 Jul;9(7):2197–2205. doi: 10.1002/j.1460-2075.1990.tb07389.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartholomew B., Kassavetis G. A., Geiduschek E. P. Two components of Saccharomyces cerevisiae transcription factor IIIB (TFIIIB) are stereospecifically located upstream of a tRNA gene and interact with the second-largest subunit of TFIIIC. Mol Cell Biol. 1991 Oct;11(10):5181–5189. doi: 10.1128/mcb.11.10.5181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borukhov S., Lee J., Goldfarb A. Mapping of a contact for the RNA 3' terminus in the largest subunit of RNA polymerase. J Biol Chem. 1991 Dec 15;266(35):23932–23935. [PubMed] [Google Scholar]
- Cormack B. P., Struhl K. The TATA-binding protein is required for transcription by all three nuclear RNA polymerases in yeast cells. Cell. 1992 May 15;69(4):685–696. doi: 10.1016/0092-8674(92)90232-2. [DOI] [PubMed] [Google Scholar]
- Dequard-Chablat M., Riva M., Carles C., Sentenac A. RPC19, the gene for a subunit common to yeast RNA polymerases A (I) and C (III). J Biol Chem. 1991 Aug 15;266(23):15300–15307. [PubMed] [Google Scholar]
- Hayward R. S., Igarashi K., Ishihama A. Functional specialization within the alpha-subunit of Escherichia coli RNA polymerase. J Mol Biol. 1991 Sep 5;221(1):23–29. doi: 10.1016/0022-2836(91)80197-3. [DOI] [PubMed] [Google Scholar]
- Igarashi K., Hanamura A., Makino K., Aiba H., Aiba H., Mizuno T., Nakata A., Ishihama A. Functional map of the alpha subunit of Escherichia coli RNA polymerase: two modes of transcription activation by positive factors. Proc Natl Acad Sci U S A. 1991 Oct 15;88(20):8958–8962. doi: 10.1073/pnas.88.20.8958. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Igarashi K., Ishihama A. Bipartite functional map of the E. coli RNA polymerase alpha subunit: involvement of the C-terminal region in transcription activation by cAMP-CRP. Cell. 1991 Jun 14;65(6):1015–1022. doi: 10.1016/0092-8674(91)90553-b. [DOI] [PubMed] [Google Scholar]
- Izban M. G., Luse D. S. The RNA polymerase II ternary complex cleaves the nascent transcript in a 3'----5' direction in the presence of elongation factor SII. Genes Dev. 1992 Jul;6(7):1342–1356. doi: 10.1101/gad.6.7.1342. [DOI] [PubMed] [Google Scholar]
- James P., Whelen S., Hall B. D. The RET1 gene of yeast encodes the second-largest subunit of RNA polymerase III. Structural analysis of the wild-type and ret1-1 mutant alleles. J Biol Chem. 1991 Mar 25;266(9):5616–5624. [PubMed] [Google Scholar]
- Kassavetis G. A., Bartholomew B., Blanco J. A., Johnson T. E., Geiduschek E. P. Two essential components of the Saccharomyces cerevisiae transcription factor TFIIIB: transcription and DNA-binding properties. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7308–7312. doi: 10.1073/pnas.88.16.7308. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kassavetis G. A., Braun B. R., Nguyen L. H., Geiduschek E. P. S. cerevisiae TFIIIB is the transcription initiation factor proper of RNA polymerase III, while TFIIIA and TFIIIC are assembly factors. Cell. 1990 Jan 26;60(2):235–245. doi: 10.1016/0092-8674(90)90739-2. [DOI] [PubMed] [Google Scholar]
- Kassavetis G. A., Riggs D. L., Negri R., Nguyen L. H., Geiduschek E. P. Transcription factor IIIB generates extended DNA interactions in RNA polymerase III transcription complexes on tRNA genes. Mol Cell Biol. 1989 Jun;9(6):2551–2566. doi: 10.1128/mcb.9.6.2551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall N. F., Price D. H. Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol Cell Biol. 1992 May;12(5):2078–2090. doi: 10.1128/mcb.12.5.2078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmer J. M., Folk W. R. Unraveling the complexities of transcription by RNA polymerase III. Trends Biochem Sci. 1990 Aug;15(8):300–304. doi: 10.1016/0968-0004(90)90018-7. [DOI] [PubMed] [Google Scholar]
- Parsons M. C., Weil P. A. Purification and characterization of Saccharomyces cerevisiae transcription factor TFIIIC. Polypeptide composition defined with polyclonal antibodies. J Biol Chem. 1990 Mar 25;265(9):5095–5103. [PubMed] [Google Scholar]
- Reines D. Elongation factor-dependent transcript shortening by template-engaged RNA polymerase II. J Biol Chem. 1992 Feb 25;267(6):3795–3800. [PMC free article] [PubMed] [Google Scholar]
- Riva M., Schäffner A. R., Sentenac A., Hartmann G. R., Mustaev A. A., Zaychikov E. F., Grachev M. A. Active site labeling of the RNA polymerases A, B, and C from yeast. J Biol Chem. 1987 Oct 25;262(30):14377–14380. [PubMed] [Google Scholar]
- Schultz P., Marzouki N., Marck C., Ruet A., Oudet P., Sentenac A. The two DNA-binding domains of yeast transcription factor tau as observed by scanning transmission electron microscopy. EMBO J. 1989 Dec 1;8(12):3815–3824. doi: 10.1002/j.1460-2075.1989.tb08559.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sentenac A. Eukaryotic RNA polymerases. CRC Crit Rev Biochem. 1985;18(1):31–90. doi: 10.3109/10409238509082539. [DOI] [PubMed] [Google Scholar]
- Surratt C. K., Milan S. C., Chamberlin M. J. Spontaneous cleavage of RNA in ternary complexes of Escherichia coli RNA polymerase and its significance for the mechanism of transcription. Proc Natl Acad Sci U S A. 1991 Sep 15;88(18):7983–7987. doi: 10.1073/pnas.88.18.7983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas M. S., Glass R. E. Escherichia coli rpoA mutation which impairs transcription of positively regulated systems. Mol Microbiol. 1991 Nov;5(11):2719–2725. doi: 10.1111/j.1365-2958.1991.tb01980.x. [DOI] [PubMed] [Google Scholar]