Abstract
Biogas production technologies commonly involve the use of natural anaerobic consortia of microbes. The objective of this study was to elucidate the importance of hydrogen in this complex microbial food chain. Novel laboratory biogas reactor prototypes were designed and constructed. The fates of pure hydrogen-producing cultures of Caldicellulosiruptor saccharolyticus and Enterobacter cloacae were followed in time in thermophilic and mesophilic natural biogas-producing communities, respectively. Molecular biological techniques were applied to study the altered ecosystems. A systematic study in 5-litre CSTR digesters revealed that a key fermentation parameter in the maintenance of an altered population balance is the loading rate of total organic solids. Intensification of the biogas production was observed and the results corroborate that the enhanced biogas productivity is associated with the increased abundance of the hydrogen producers. Fermentation parameters did not indicate signs of failure in the biogas production process. Rational construction of more efficient and sustainable biogas-producing microbial consortia is proposed.
1. Introduction
Various approaches have been developed for the treatment and elimination of organic waste, often involving biological systems [1, 2]. Technologies that convert organic material into biogas or hydrogen (H2) in fermentation processes are the only ones that simultaneously allow the combined advantages of waste disposal and the generation of useful energy [3–6]. Biogas, a renewable energy carrier consisting mainly of methane (CH4) and carbon dioxide (CO2), is the end-product of the anaerobic digestion of organic material [7] and can be exploited in various ways. After the removal of trace contaminations consisting of hydrogen sulfide, xyloxanes, and water, it can be burnt to generate heat or can be used as fuel in gas engines, coupled to a generator to produce electricity and heat. If the CO2 is also eliminated from the biogas, the remaining gas, often called biomethane, has the properties of purified natural gas and can be utilized in every applications to replace fossil natural gas as transportation fuel, raw material for the chemical industry, or in fuel cells, which convert it to electricity with high efficiency [1].
Biogas technologies commonly apply natural anaerobic consortia of microbes. These communities form an intricate microbiological food chain. The population dynamics of the natural ecosystems were not adequately studied before the introduction of molecular biological techniques [8–12]. Research on the diversity of these microbial communities is needed for the optimization of biogas production technologies as the economic viability is closely related to the efficacy of the concerted microbiological action [13–18]. One of the rate-limiting factors in biogas-producing consortia is the actual level of H2 in the system [10, 18–20]. The presence of too much H2 inhibits the acetogenic bacteria that generate H2 in the system, whereas too little H2 has an adverse effect on an important group of methanogens, the hydrogenotrophic methanogens. In natural ecosystems, a very low partial pressure of H2 is maintained, which may be a limiting factor for the methanogens [10, 18, 21].
We demonstrated earlier that reductant accessibility is a regulating factor in biogas production and presented data supporting the hypothesis that the introduction of H2-producing bacteria into a natural biogas-generating consortium appreciably increases the efficacy of biogas production both in batch fermentations and in scaled-up anaerobic digestion [6, 10].
In the present study, systematic experiments were conducted in 5-liter continuously stirred tank reactor (CSTR) reactors specifically designed for biogas research on a laboratory scale. These devices model the real-life, large-scale biogas production plants much better than the routinely used batch systems and some of the first results are reported here. Thermophilic and mesophilic conditions were selected in order to study both of the temperature ranges applied in anaerobic digesters. The microbial diversity in the thermophilic natural consortia is lower, which necessitates a thorough inspection of the microbiological profiles. Introduction of a H2-producing new member into such consortia is therefore somewhat more challenging than altering the composition of a microbial consortium under mesophilic conditions. Caldicellulosiruptor saccharolyticus is an excellent thermophilic H2 producer and its addition to biogas- and biohydrogen-generating systems has a demonstrated beneficial effect [6, 10, 22, 23]. Enterobacter cloacae was selected as a promising candidate to play a similar role in the mesophilic environment. An important question that remained before the implementation of large-scale application related to the conditions under which C. saccharolyticus and E. cloacae become stable members of the respective biogas-producing microbial consortia. This can be determined only in systematic experiments with digesters functioning in continuous operation mode. The survival of C. saccharolyticus and of E. cloacae in a thermophilic and mesophilic biogas-producing community, respectively, was therefore tested by molecular biological methods.
2. Materials and Methods
2.1. Laboratory Biogas Reactor
Explicitly designed reactors with a working volume of 5 L and a headspace of 1 L were custom-made from stainless steel by Biospin Ltd., Szeged, Hungary, and the design is presented here for the first time. The substrate is stored at ambient temperature in a reservoir and is mixed and mechanically pretreated by a shredder pump. It can be fed into the apparatus either continuously or intermittently, through a piston-type delivery system, which controls the substrate volume introduced into the reactor. Simultaneously with the feeding, the same volume of fermented material is removed through an overflow via U-shaped tubing in order to maintain a gas-tight closure and a constant working volume in the reactor. The biogas reactors are equipped with a spiral strip mixing device driven by an electronic engine. Three devices are operated by the same electronic engine through a belt transmission in order to maintain identical mixing conditions and to save on construction and operational costs. An electronically heated jacket surrounds the cylindrical reactor body. Temperature is measured with a thermistor sensor, and constant temperature is maintained with an accuracy of ±0.5°C. Electrodes for the measurement of pH and redox potential are inserted through the wall of the reaction vessel, in sealed sockets. Formation of the anaerobic environment can be facilitated via a gas delivery ring situated near the bottom of the apparatus. Ultrapure nitrogen gas is used to spurge the system at the beginning of the experiment. The device can be drained at the bottom, where samples for biomass analysis can likewise be removed. The top plate can be opened for inspection and cleaning. The plate is fitted with a neoprene O-ring and is secured to the main body by flexible clamps. The evolved gas leaves the reactor through flexible tubing connected to the top plate, where ports for gas sampling and the delivery of liquids by means of syringes through silicone rubber septa are also installed (Figure 1).
Figure 1.
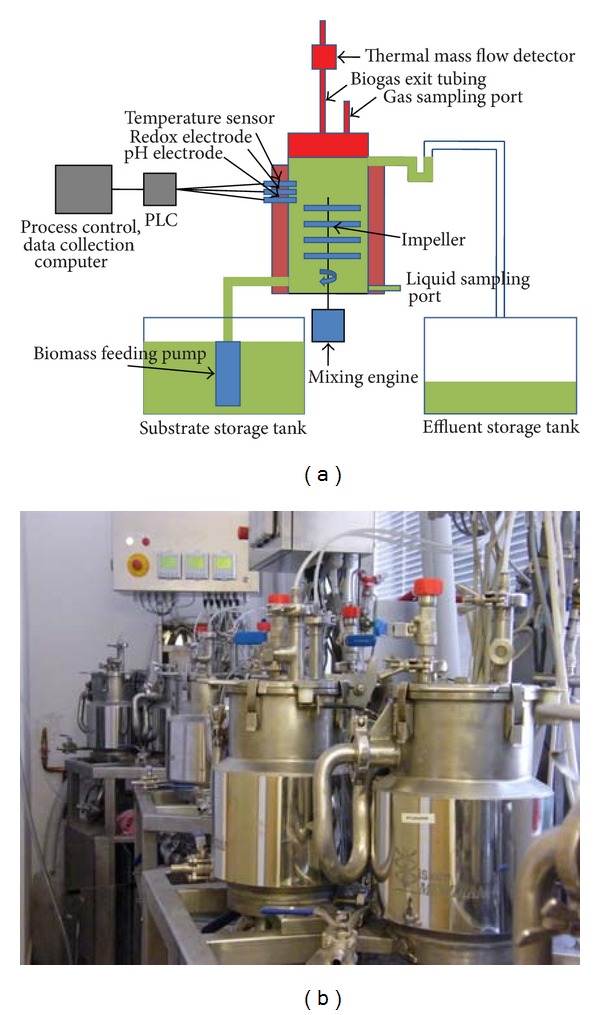
Scheme of operation of the reactors used in this study, some of which are illustrated below. An explanation is given in the text.
Temperature, pH, and redox potential are monitored continuously. The data are displayed on a panel above each apparatus and are fed into a process-control computer via a digital converter. Gas volume is measured by means of direct mass flow controllers (DMFC, Brooks Instruments) attached to each gas exit port. Data are collected, stored, and analyzed by special software developed by Merat Ltd., Budapest, Hungary. The key parameters (temperature, mixing speed, and pH) are continuously controlled by the software.
2.2. Methods
2.2.1. Microorganism, Medium, and Culture Conditions
Caldicellulosiruptor saccharolyticus (DSM 8903) [22] was purchased from Deutsche Sammlung von Mikroorganismen und Zellkulturen GmbH and propagated at 70°C on DSMZ medium 640 in anaerobic 50 mL hypovials (Supelco) until OD600 = 0.5.
Enterobacter cloacae (DSM30054) [24] was cultivated at 30°C on DSMZ medium 1 in sterile Erlenmeyer flasks until OD600 = 1.5.
Freshly grown cultures were added to the reactors directly at a concentration of 5% (v/v) by means of a syringe.
2.2.2. Cell Growth and Viable Biomass Determination
Viable cell counts of C. saccharolyticus were determined by plating serially diluted cell suspensions in the stationary phase on DSMZ medium 640 solidified with 2.5% (w/v) Gelrite Gellan Gum (Sigma-Aldrich) [25]. Plating was performed anaerobically in an anaerobic chamber (Bactron IV, Sheldon Manufacturing) [4, 5], and the plates were incubated at 70°C for 3 to 4 days for the determination of colony forming units.
E. cloacae cells were counted on agar plates of DSMZ medium 1 solidified with 1.5% (w/v) agar (Biolab). The plates were incubated overnight at 37°C.
2.2.3. Biogas Substrate
The substrate for anaerobic digestion consisted of a mixture of pig slurry (25% w/v) and chopped sweet sorghum (75% w/v). The sweet sorghum plant material was collected in fresh green form, chopped to pieces measuring less than 5 mm and stored frozen at −20°C before use. An inoculum from a thermophilic sewage sludge digester was used to start the experiments involving C. saccharolyticus. An inoculum from a mesophilic agricultural biogas plant was used to initiate the fermentation in the case of E. cloacae. The inocula were preincubated for at least 2 weeks at 55 or 37°C, respectively, before use.
2.2.4. Gas Analysis
The composition of the evolved biogas was measured by taking 250 μL aliquots from the headspace and injecting them into a gas chromatograph (6890N Network GC System, Agilent Technologies) equipped with a 5 Å molecular sieve column (length 30 m, I.D. 0.53 megabore, film 25 μm) and a thermal conductivity detector. Nitrogen was used as carrier gas.
2.2.5. Volatile Acids
Volatile acids were determined by HPLC (Hitachi Elite, equipped with an ICSep ICE-COREGEL 64H column and a refractive index detector L2490), under the following conditions: solvent 0.1 N H2SO4, flow rate 0.8 mL min−1, column temperature 50°C, detector temperature 41°C.
2.2.6. Organic Carbon and Dry Matter Content Measurement
Total organic carbon (TOC) was determined with a Teledyne Tekmar Apollo 9000 automatic TOC instrument. This apparatus burns the biomass at 730°C and measures the released COx and NOx by infrared absorption. The dry matter content was quantified by drying the biomass at 105°C overnight and weighing the residue. Further heating of this residue at 550°C until its weight did not change yielded the organic dry matter content [10].
2.2.7. Molecular Biology Techniques
For the purification of genomic DNA, the QIAmp DNA Stool Mini Kit (Qiagen) was used in accordance with the manufacturer's recommendations, except that DNA was eluted from the column in 50 μL of elution buffer. The total bacterial DNA was extracted from the pure cultures with the GenElute Bacterial Genomic DNA kit (Sigma-Aldrich).
PCR primer pairs were designed, using Primer Express 2.0 software (Life Technologies). Two targeted genes were selected from the C. saccharolyticus strain to create specific PCR primers. The first set was EchA-F1 (5′-TCAGCACAGTTTCCGTTCCA-3′) and EchA-R1 (5′-TCCTGCTTTTACCATTGTACTTGAA-3′). These primers were designed to amplify a 100-bp segment of the echA gene, which codes a putative, membrane-associated [NiFe]-hydrogenase subunit. The other primer pair was celA-F1 (5′-GGGTCCTGCTGAGGTTATGC-3′) and celA-R1 (5′-GCTAAGGAAGCTGCCGTCTCT-3′). The PRC product was a 100-bp fragment of the celA gene, which codes for a subunit of cellulase. In the case of E. cloacae, the PCR reaction targeted a fragment of the gene coding for the large subunit of [NiFe]-hydrogenase3. The 100-bp product was amplified with the primer pair HycE_F2 (5′-TGTTGCCGCGCAGCATGTAG-3′) and HycE_R2 (5′-TGACCGGCGACAACCAGAAG-3′).
Sensitivity studies were performed to determine the lowest amount of DNA that can be used as positive control. The sensitivity measurements were carried out with pure genomic DNA.
All PCR reactions were performed in a 7500 Fast Real-Time system, using Power SYBR Green PCR Master Mix (Life Technologies). The original concentration of the genomic DNA was 150 ng μL−1. Five dilutions were made from both DNA samples; each of them was attenuated ten times relative to the previous one. Real-time PCR reaction experiments included a non-template control, five positive controls, and various samples in three parallel measurements. The PCR reactions contained the primers (1.5 pg μL−1), the template in various amounts (1.5 ng μL−1–200 ng μL−1), the manufacturer's SYBR Green PCR Master Mix, and water to a final volume of 25 μL. The temperature profile was as follows: a 10 min initial enzyme activation at 95°C, followed by 40 consecutive cycles at 95°C for 15 s, and termination at 60°C for 1 min.
3. Results and Discussion
Anaerobic digestion is one of the most promising of the various biomass conversion processes. H2 is an important ingredient in the anaerobic fermentation of organic materials. The regulatory roles of the H2 levels and interspecies H2 transfer optimize the concerted action of the complex microbial population [6, 10, 20, 21]. The H2 concentration has been shown to determine the structure of the methanogenic community [9]. In anaerobic habitats, methanogens keep fermentative pathways energetically favorable by maintaining an extremely low partial pressure of H2. The methanogenic archaea are a highly specialized group of microbes as they produce CH4, which is both a useful energy source and a powerful greenhouse gas [1, 8]. These organisms are ubiquitous in aquatic and marine sediments, sewage sludge, and the intestines of ruminants and some other animals. They are also responsible for the final steps of biogas formation in anaerobic digesters [14, 15, 17, 26]. The hydrogenotrophic methanogens use H2 to reduce CO2 to CH4, while the acetotrophic methanogens split acetate to CH4 and CO2 [21]. The expressions of up to 10% of the total proteins in a hydrogenotrophic methanogen were reported to change in response to a H2 limitation [12], indicating that the H2 availability is sensed by the methanogens and that this gas has a major effect on their physiology. It has been found that the addition of a pure culture of H2-producing bacterium intensifies biogas production in laboratory batch experiments, and the effect was also observed in a 5 m3 semi-continuous biogas digester [6, 10]. To test the phenomenon at thermophilic temperature, C. saccharolyticus was selected, while E. cloacae was chosen for similar experiments under mesophilic conditions. C. saccharolyticus is detectable in anaerobic digesters only through the use of sophisticated metagenomics tools [18]. To follow up these observations, a systematic study was carried out under standardized anaerobic digestion conditions. All experiments were performed in three separate, parallel fermentations. Under the commonly employed feeding conditions (4 g total organic solids L−1 day−1), addition of C. saccharolyticus or E. cloacae culture at the time when stable and reproducible daily biogas production had been attained led to an intensification effect similar to that observed in the batch experiments (Figures 3 and 7) [10]. However, the bacteria in the semicontinuously fed reactors gradually diluted out and disappeared within 2-3 weeks, and no C. saccharolyticus or E. cloacae was then detectable with the DNA molecular marker method (Figures 2 and 6).
Figure 3.
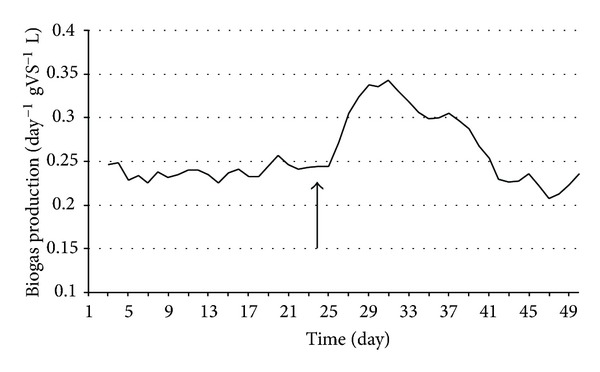
Biogas production in CSTR reactors. C. saccharolyticus was inoculated at the time point indicated by the arrow. Feeding rate: 4 g total organic solids L−1 day−1.
Figure 7.
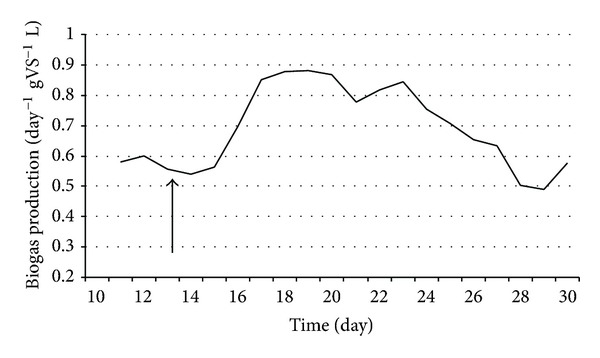
Biogas production in CSTR reactors. E. cloacae was inoculated at the time point indicated by the arrow. Feeding rate: 4 g total organic solids L−1 day−1.
Figure 2.
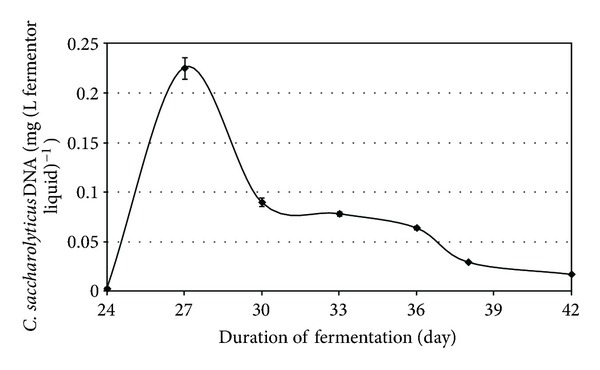
Detection of C. saccharolyticus DNA in the reactors at various times on use of a low organic loading rate, that is, 4 g total organic solids L−1 day−1. PCR was carried out with the EchA-F1 and EchA-R1 probes. The cellulose-specific celA-F1 and celA-R1 probes corroborated these findings and are therefore not shown.
Figure 6.
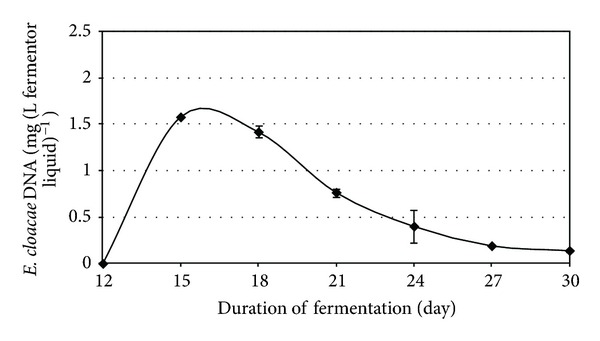
Detection of E. cloacae DNA in the reactors at various times on use of a low organic loading rate, that is, 4 g total organic solids L−1 day−1.
A systematic study was launched to elucidate the reason for the discrepancy between these results and those of the earlier scale-up experiment.
The biogas productivity correlated strongly with the presence of C. saccharolyticus in the system (Figure 5).
Figure 5.
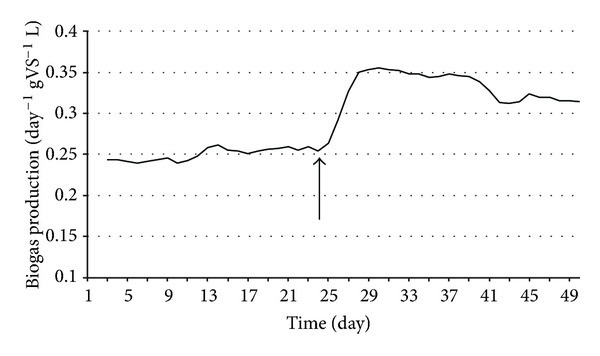
Biogas production in CSTR reactors. C. saccharolyticus was added at the time point indicated by the arrow. Feeding rate: 8 g total organic solids L−1 day−1.
The loading rate of the organic solids was identified as one of the parameters potentially responsible for the effect. Indeed, when the loading rate was increased, the obvious beneficial effect of the added H2-producer C. saccharolyticus lasted substantially longer (Figures 4 and 5). Figures 6 and 7 demonstrate a very similar effect, observed under mesophilic conditions after inoculation with E. cloacae. The extent of the intensification decreases in time, as the microbe is gradually diluted out of the system (Figure 7). This observation is substantiated by the measurement of the E. cloacae-specific DNA (Figure 6). Roughly 2 weeks after inoculation, the added E. cloacae has disappeared from the reactor because it cannot not keep up with the rest of the microbial consortium and is washed out.
Figure 4.
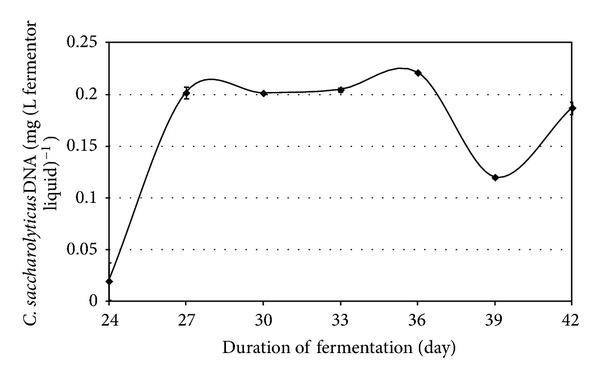
Detection of C. saccharolyticus DNA in the reactors after various times when a high organic loading rate was applied, that is, 8 g total organic solids L−1 day−1. PCR was carried out with the the EchA-F1 and EchA-R1 probes. The cellulase-specific celA-F1 and celA-R1 probes corroborated these findings and are therefore not shown.
Similarly to the thermophilic anaerobic digestion, in the case of the mesophilic fermentation the bioaugmentation persisted when the loading rate was elevated to 8 g total organic solids L−1 day−1 (Figures 8 and 9). Although the enhancement of biogas production was not so spectacular as in the thermophilic case, the approximately 30% increase persisted for an extended period of time, well beyond the wash-out period (Figure 9).
Figure 8.
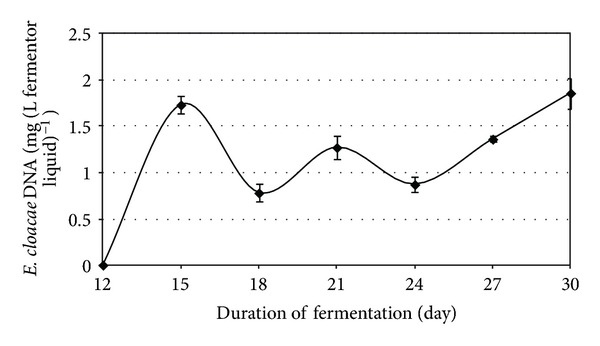
Detection of E. cloacae DNA in the reactors at various times when a high organic loading rate was applied, that is, 8 g total organic solids L−1 day−1.
Figure 9.
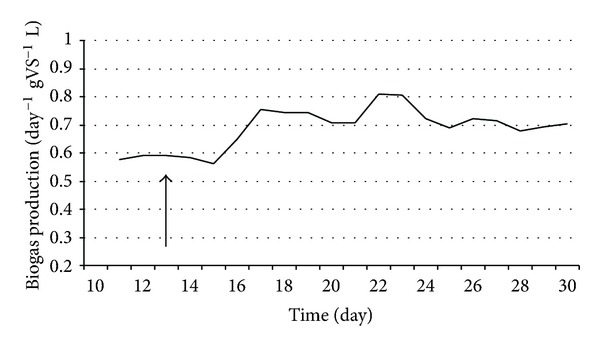
Biogas production in CSTR reactors. E. cloacae was added at the time point indicated by the arrow. Feeding rate: 8 g total organic solids L−1 day−1.
During this period, E. cloacae DNA appeared stable immediately after inoculation The observed intensification of the biogas by E. cloacae was proportional to its cell number.
Other factors influencing the long-lasting intensification of biogas production in this system are currently being studied. It should be noted, for example, that the temperature of 55°C routinely used in thermophilic anaerobic digesters is not ideal for growth of the extremely thermophilic C. saccharolyticus. The two sets of fermentations gave concordant results, as an indication that the organic loading rate is one of the most important parameters when the survival of a bioaugmentation strain is crucial. Intensification of the biogas production can be achieved under mesophilic or thermophilic conditions. The marked difference in overall biogas production rate in the two thermal milieu may be explained by two reasons. First, the inoculum of the thermophilic reactor originated from a biogas plant, where mainly sewage sludge was used, and the microbial community was therefore not perfectly suitable for the degradation of sweat sorghum and pig slurry. Second, the plant material used in the mesophilic and thermophilic experiments originated from different harvests. The inconsistent sugar content of the biomass could also have been responsible for the difference in gas production rate.
The volatile fatty acid (VFA) concentration in the reactors were somewhat elevated during the experiments involving high organic loading rate, but remained stable (Figure 10). The fact that VFA did not accumulate in time suggests a stable anaerobic degradation process following the addition of the H2 producers therefore the system operated in a balanced fashion upon intensification. It is not surprising that the VFA level was elevated when the organic loading rate increased (Figure 10) relative to the low organic loading rate condition (Figure 11) indicating that the microbiological community was overloaded with substrate. Essentially the same behavior was observed under mesophilic conditions (data not shown). The stability of the VFA levels suggests that the community could handle well this situation. Longer chain VFAs were not produced in measurable amounts.
Figure 10.
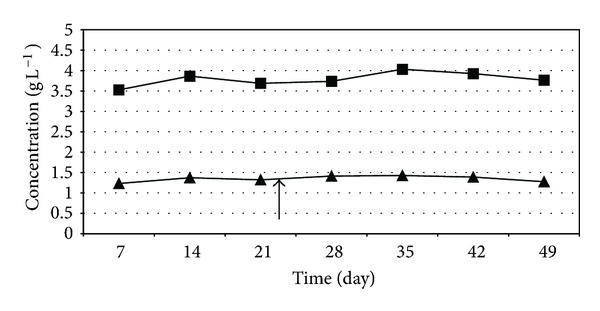
Volatile fatty acid levels during the thermophilic biogas intensification experiment. C. saccharolyticus was added at the time point indicated by the arrow. ■—acetate concentration, ▲—propionate concentration. Feeding rate: 8 g total organic solids L−1 day−1.
Figure 11.
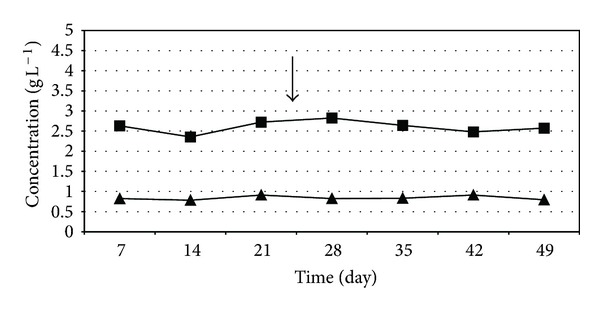
Volatile fatty acid levels during the thermophilic biogas intensification experiment. C. saccharolyticus was added at the time point indicated by the arrow. ■—acetate concentration, ▲—propionate concentration. Feeding rate: 4 g total organic solids L−1 day−1.
Process parameters such as pH and gas composition did not change during the experiments (data not shown). These indicate that the fermentation was not disturbed by the addition of the hydrogen producing bacteria.
4. Conclusions
A positive correlation was demonstrated between the intensification of biogas production and the presence of both added H2-producing microorganism strains in a natural biogas-generating ecosystem. The substrate composition did not markedly affect the elevated biogas production relative to the untreated controls. It is therefore envisaged that a rational design and engineering of the biogas-producing microbial community is possible.
C. saccharolyticus with its versatile H2-production activity augments biogas productivity from various substrates to a similar extent to E. cloacae. In a continuously operated industrial biogas facility, an important factor determining the value of this biotechnological improvement of performance is the persistence of the beneficial effect in time. C. saccharolyticus and E. cloacae are both capable of incorporating into the natural biogas-producing consortia under appropriate conditions. A significant parameter in this respect is the rate at which the reactor is loaded with organic substrates. Future studies will have the aim of the identification of other rate-limiting boundary conditions with a view to improving the yield and economic viability of biogas technology.
A laboratory device specifically designed for biogas studies has been developed and tested. It proved superior to the commonly used batch fermentation systems and is a suitable model of continuously operated industrial-scale biogas reactors.
Acknowledgments
This work was supported by EU Projects HUSRB/1002/214/041 IPA, HURO/1001/193/2.2.2. CBC, and IEE/10/235 SI2.591589 GreenGasGrids. Hungarian funds from TÁMOP-4.2.1/B-09/1/KONV-2010-0005 and TÁMOP-4.2.2/B-10/1-2010-0012 are gratefully acknowledged.
References
- 1.Ahring BK. Perspectives for anaerobic digestion. Advances in Biochemical Engineering/Biotechnology. 2003;81:1–30. doi: 10.1007/3-540-45839-5_1. [DOI] [PubMed] [Google Scholar]
- 2.Pind PF, Angelidaki I, Ahring BK. Dynamics of the anaerobic process: effects of volatile fatty acids. Biotechnology and Bioengineering. 2003;82(7):791–801. doi: 10.1002/bit.10628. [DOI] [PubMed] [Google Scholar]
- 3.Zeeman G, Lettinga G. The role of anaerobic digestion of domestic sewage in closing the water and nutrient cycle at community level. Water Science and Technology. 1999;39(5):187–194. [Google Scholar]
- 4.Ivanova G, Rákhely G, Kovács KL. Hydrogen production from biopolymers by Caldicellulosiruptor saccharolyticus and stabilization of the system by immobilization. International Journal of Hydrogen Energy. 2008;33(23):6953–6961. [Google Scholar]
- 5.Ivanova G, Rákhely G, Kovács KL. Thermophilic biohydrogen production from energy plants by Caldicellulosiruptor saccharolyticus and comparison with related studies. International Journal of Hydrogen Energy. 2009;34(9):3659–3670. [Google Scholar]
- 6.Herbel Z, Rákhely G, Bagi Z, et al. Exploitation of the extremely thermophilic Caldicellulosiruptor saccharolyticus in hydrogen and biogas production from biomasses. Environmental Technology. 2010;31(8-9):1017–1024. doi: 10.1080/09593330.2010.484075. [DOI] [PubMed] [Google Scholar]
- 7.Miron Y, Zeeman G, Van Lier JB, Lettinga G. The role of sludge retention time in the hydrolysis and acidification of lipids, carbohydrates and proteins during digestion of primary sludge in CSTR systems. Water Research. 2000;34(5):1705–1713. [Google Scholar]
- 8.Hendrickson EL, Kaul R, Zhou Y, et al. Complete genome sequence of the genetically tractable hydrogenotrophic methanogen Methanococcus maripaludis . Journal of Bacteriology. 2004;186(20):6956–6969. doi: 10.1128/JB.186.20.6956-6969.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Leybo AI, Netrusov AI, Conrad R. Effect of hydrogen concentration on the community structure of hydrogenotrophic methanogens studied by T-RELP analysis of 16S rRNA gene amplicons. Microbiology. 2006;75(6):683–688. [PubMed] [Google Scholar]
- 10.Bagi Z, Ács N, Bálint B, et al. Biotechnological intensification of biogas production. Applied Microbiology and Biotechnology. 2007;76(2):473–482. doi: 10.1007/s00253-007-1009-6. [DOI] [PubMed] [Google Scholar]
- 11.Van De Werken HJG, Verhaart MRA, VanFossen AL, et al. Hydrogenomics of the extremely thermophilic bacterium Caldicellulosiruptor saccharolyticus . Applied and Environmental Microbiology. 2008;74(21):6720–6729. doi: 10.1128/AEM.00968-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Xia Q, Wang T, Hendrickson EL, Lie TJ, Hackett M, Leigh JA. Quantitative proteomics of nutrient limitation in the hydrogenotrophic methanogen Methanococcus maripaludis . BMC Microbiology. 2009;9, article 149 doi: 10.1186/1471-2180-9-149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Hofman-Bang J, Zheng D, Westermann P, Ahring BK, Raskin L. Molecular ecology of anaerobic reactor systems. Advances in Biochemical Engineering/Biotechnology. 2003;81:151–203. doi: 10.1007/3-540-45839-5_5. [DOI] [PubMed] [Google Scholar]
- 14.Nettmann E, Bergmann I, Mundt K, Linke B, Klocke M. Archaea diversity within a commercial biogas plant utilizing herbal biomass determined by 16S rDNA and mcrA analysis. Journal of Applied Microbiology. 2008;105(6):1835–1850. doi: 10.1111/j.1365-2672.2008.03949.x. [DOI] [PubMed] [Google Scholar]
- 15.Klocke M, Nettmann E, Bergmann I, et al. Characterization of the methanogenic Archaea within two-phase biogas reactor systems operated with plant biomass. Systematic and Applied Microbiology. 2008;31(3):190–205. doi: 10.1016/j.syapm.2008.02.003. [DOI] [PubMed] [Google Scholar]
- 16.Franke-Whittle IH, Goberna M, Pfister V, Insam H. Design and development of the ANAEROCHIP microarray for investigation of methanogenic communities. Journal of Microbiological Methods. 2009;79(3):279–288. doi: 10.1016/j.mimet.2009.09.017. [DOI] [PubMed] [Google Scholar]
- 17.Goberna M, Insam H, Franke-Whittle IH. Effect of biowaste sludge maturation on the diversity of thermophilic bacteria and archaea in an anaerobic reactor. Applied and Environmental Microbiology. 2009;75(8):2566–2572. doi: 10.1128/AEM.02260-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wirth R, Kovács E, Maróti G, Bagi Z, Rákhely G, Kovács KL. Characterization of biogas-producing microbial community by short-read next generation DNA sequencing. Biotechnology for Biofuels. 2012;5, article 41 doi: 10.1186/1754-6834-5-41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Bernstead J, Archer DB, Lloyd D. Role of hydrogen in the growth of mutualistic methanogenic cocultures. In: Baich J-P, Bruschi M, Garcia J-L, editors. Microbiology and Biochemistry of Strict Anaerobes Involved in Interspecies Transfer. New York, NY, USA: Plenum Press; 1990. pp. 161–171. [Google Scholar]
- 20.Schink B, Stams AJM. Syntrophism among prokaryotes. In: Dworkin M, Schleifer KH, Stackebrandt E, editors. The Prokaryotes. New York, NY, USA: Springer; 2002. [Google Scholar]
- 21.Demirel B, Scherer P. The roles of acetotrophic and hydrogenotrophic methanogens during anaerobic conversion of biomass to methane: a review. Reviews in Environmental Science and Biotechnology. 2008;7(2):173–190. [Google Scholar]
- 22.Rainey FA, Donnison AM, Janssen PH, et al. Description of Caldicellulosiruptor saccharolyticus gen. nov., sp. nov: an obligately anaerobic, extremely thermophilic, cellulolytic bacterium. FEMS Microbiology Letters. 1994;120(3):263–266. doi: 10.1111/j.1574-6968.1994.tb07043.x. [DOI] [PubMed] [Google Scholar]
- 23.Ivanova G, Rákhely G, Kovács KL. Hydrogen production from biopolymers by Caldicellulosiruptor saccharolyticus and stabilization of the system by immobilization. International Journal of Hydrogen Energy. 2008;33(23):6953–6961. [Google Scholar]
- 24.Ren Y, Ren Y, Zhou Z, et al. Complete genome sequence of Enterobacter cloacae subsp. cloacae type strain ATCC 13047. Journal of Bacteriology. 2010;192(9):2463–2464. doi: 10.1128/JB.00067-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Rákhely G, Kovács KL. Plating hyperthermophilic archea on solid surface. Analytical Biochemistry. 1996;243(1):181–183. doi: 10.1006/abio.1996.0499. [DOI] [PubMed] [Google Scholar]
- 26.Goberna M, Gadermaier M, García C, Wett B, Insam H. Adaptation of methanogenic communities to the cofermentation of cattle excreta and olive mill wastes at 37°C and 55°C. Applied and Environmental Microbiology. 2010;76(19):6564–6571. doi: 10.1128/AEM.00961-10. [DOI] [PMC free article] [PubMed] [Google Scholar]


