Abstract
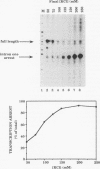
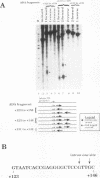
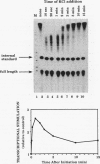
Transcription arrest plays a key role in the regulation of the murine adenosine deaminase (ADA) gene, as well as a number of other cellular and viral genes. We have previously characterized the ADA intron 1 arrest site, located 145 nucleotides downstream of the transcription start site, with respect to sequence and elongation factor requirements. Here, we show that the optimal conditions for both intron 1 arrest and overall ADA transcription involve the addition of high concentrations of KCl soon after initiation. As we have further delineated the sequence requirements for intron 1 arrest, we have found that sequences downstream of the arrest site are unnecessary for arrest. Also, a 24-bp fragment containing sequences upstream of the arrest site is sufficient to generate arrest downstream of the adenovirus major late promoter only in the native orientation. Surprisingly, we found that deletion of sequences encompassing the ADA transcription start site substantially reduced intron 1 arrest, with no effect on overall levels of transcription. At the same time, deletion of sequences upstream of the TATA box had no significant effect on either process. We believe the start site mutations have disrupted either the assembly or the composition of the transcription complex such that intron 1 site read-through is now favored. This finding, coupled with the increase in overall transcription after high-concentration KCl treatment, allows us to further refine our model of ADA gene regulation.
Full text
PDF


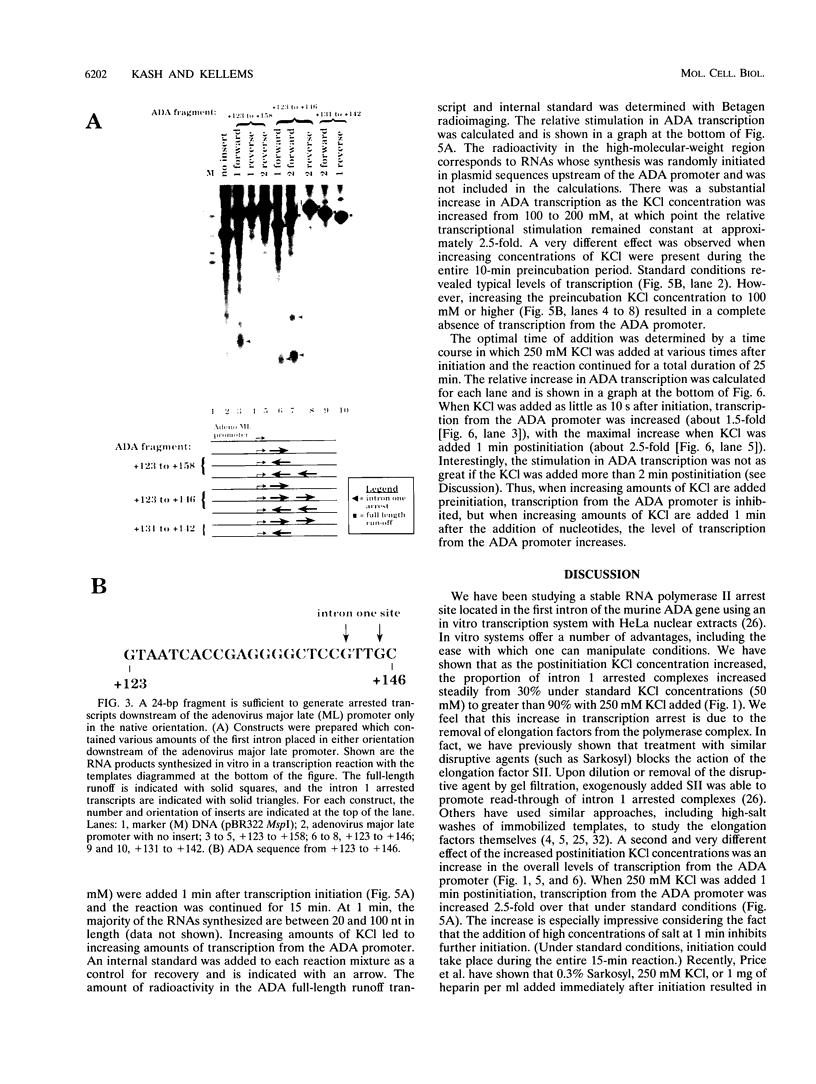
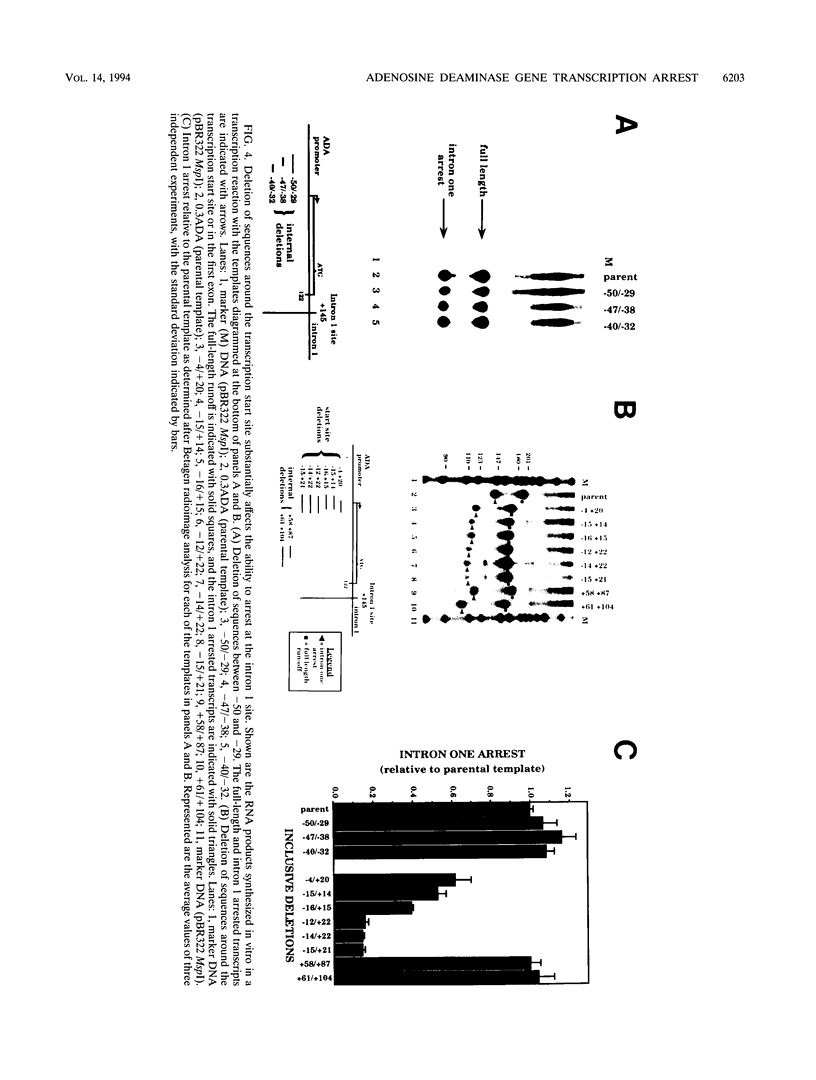
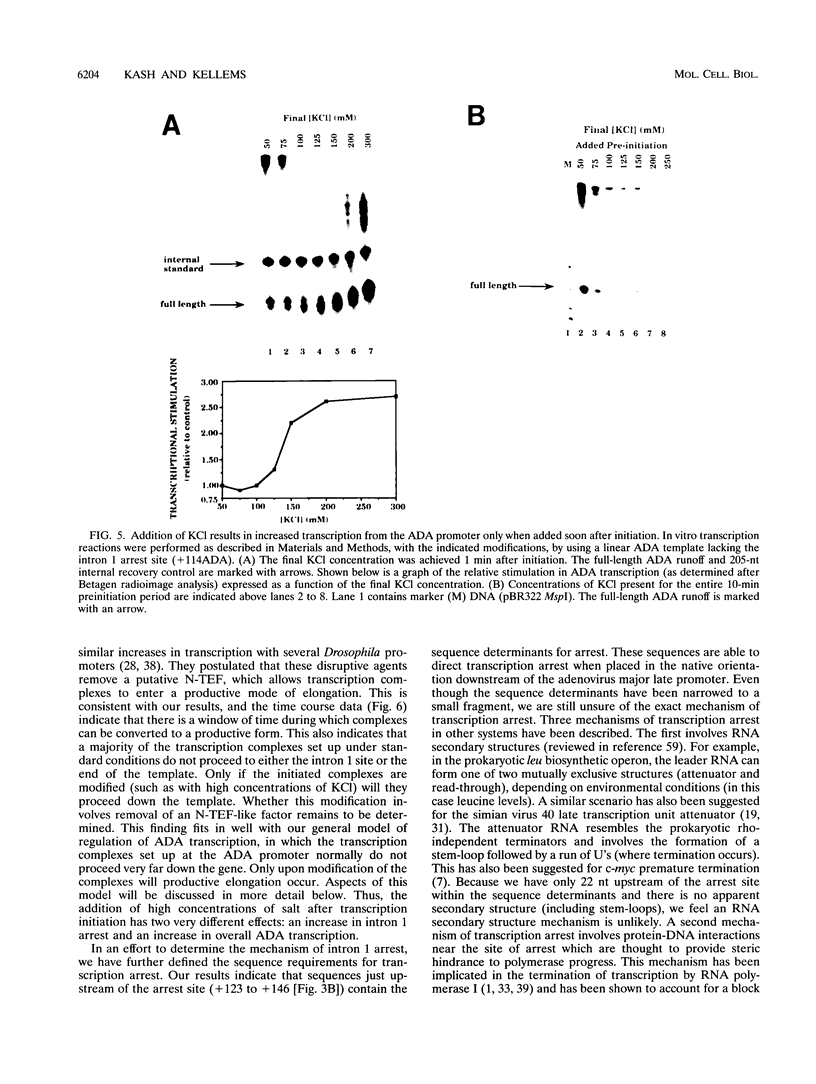
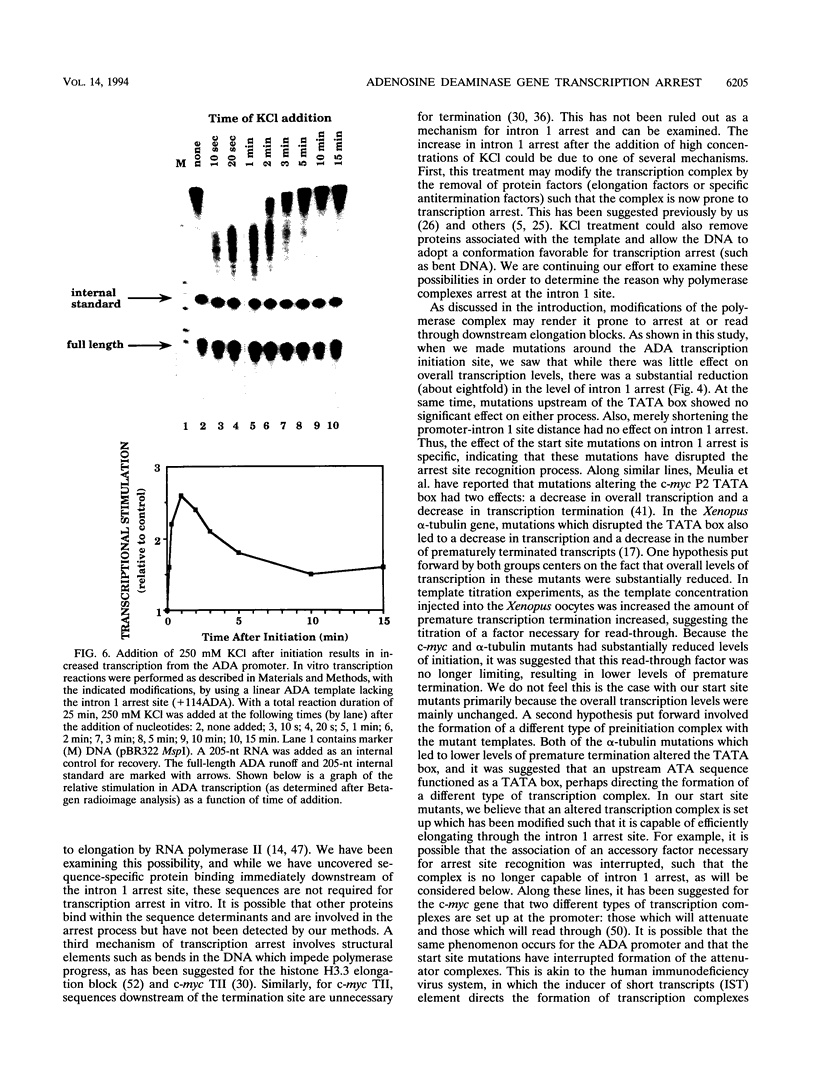


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bartsch I., Schoneberg C., Grummt I. Purification and characterization of TTFI, a factor that mediates termination of mouse ribosomal DNA transcription. Mol Cell Biol. 1988 Sep;8(9):3891–3897. doi: 10.1128/mcb.8.9.3891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ben-Asher E., Aloni Y. Transcription of minute virus of mice, an autonomous parvovirus, may be regulated by attenuation. J Virol. 1984 Oct;52(1):266–276. doi: 10.1128/jvi.52.1.266-276.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bender T. P., Thompson C. B., Kuehl W. M. Differential expression of c-myb mRNA in murine B lymphomas by a block to transcription elongation. Science. 1987 Sep 18;237(4821):1473–1476. doi: 10.1126/science.3498214. [DOI] [PubMed] [Google Scholar]
- Bengal E., Flores O., Krauskopf A., Reinberg D., Aloni Y. Role of the mammalian transcription factors IIF, IIS, and IIX during elongation by RNA polymerase II. Mol Cell Biol. 1991 Mar;11(3):1195–1206. doi: 10.1128/mcb.11.3.1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bengal E., Goldring A., Aloni Y. Transcription complexes synthesizing attenuated RNA can serve as a model system for analyzing elongation factors. J Biol Chem. 1989 Nov 15;264(32):18926–18932. [PubMed] [Google Scholar]
- Bentley D. L., Groudine M. A block to elongation is largely responsible for decreased transcription of c-myc in differentiated HL60 cells. Nature. 1986 Jun 12;321(6071):702–706. doi: 10.1038/321702a0. [DOI] [PubMed] [Google Scholar]
- Bentley D. L., Groudine M. Sequence requirements for premature termination of transcription in the human c-myc gene. Cell. 1988 Apr 22;53(2):245–256. doi: 10.1016/0092-8674(88)90386-8. [DOI] [PubMed] [Google Scholar]
- Björklund S., Skogman E., Thelander L. An S-phase specific release from a transcriptional block regulates the expression of mouse ribonucleotide reductase R2 subunit. EMBO J. 1992 Dec;11(13):4953–4959. doi: 10.1002/j.1460-2075.1992.tb05602.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z., Harless M. L., Wright D. A., Kellems R. E. Identification and characterization of transcriptional arrest sites in exon 1 of the human adenosine deaminase gene. Mol Cell Biol. 1990 Sep;10(9):4555–4564. doi: 10.1128/mcb.10.9.4555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Z., Innis J. W., Sun M. H., Wright D. A., Kellems R. E. Sequence requirements for transcriptional arrest in exon 1 of the human adenosine deaminase gene. Mol Cell Biol. 1991 Dec;11(12):6248–6256. doi: 10.1128/mcb.11.12.6248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chinsky J. M., Maa M. C., Ramamurthy V., Kellems R. E. Adenosine deaminase gene expression. Tissue-dependent regulation of transcriptional elongation. J Biol Chem. 1989 Aug 25;264(24):14561–14565. [PubMed] [Google Scholar]
- Collart M. A., Tourkine N., Belin D., Vassalli P., Jeanteur P., Blanchard J. M. c-fos gene transcription in murine macrophages is modulated by a calcium-dependent block to elongation in intron 1. Mol Cell Biol. 1991 May;11(5):2826–2831. doi: 10.1128/mcb.11.5.2826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cullen B. R. The HIV-1 Tat protein: an RNA sequence-specific processivity factor? Cell. 1990 Nov 16;63(4):655–657. doi: 10.1016/0092-8674(90)90129-3. [DOI] [PubMed] [Google Scholar]
- Deuschle U., Hipskind R. A., Bujard H. RNA polymerase II transcription blocked by Escherichia coli lac repressor. Science. 1990 Apr 27;248(4954):480–483. doi: 10.1126/science.2158670. [DOI] [PubMed] [Google Scholar]
- Dignam J. D., Lebovitz R. M., Roeder R. G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 1983 Mar 11;11(5):1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grayhack E. J., Yang X. J., Lau L. F., Roberts J. W. Phage lambda gene Q antiterminator recognizes RNA polymerase near the promoter and accelerates it through a pause site. Cell. 1985 Aug;42(1):259–269. doi: 10.1016/s0092-8674(85)80121-5. [DOI] [PubMed] [Google Scholar]
- Hair A., Morgan G. T. Premature termination of tubulin gene transcription in Xenopus oocytes is due to promoter-dependent disruption of elongation. Mol Cell Biol. 1993 Dec;13(12):7925–7934. doi: 10.1128/mcb.13.12.7925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haley J. D., Waterfield M. D. Contributory effects of de novo transcription and premature transcript termination in the regulation of human epidermal growth factor receptor proto-oncogene RNA synthesis. J Biol Chem. 1991 Jan 25;266(3):1746–1753. [PubMed] [Google Scholar]
- Hay N., Aloni Y. Attenuation in SV40 as a mechanism of transcription-termination by RNA polymerase B. Nucleic Acids Res. 1984 Feb 10;12(3):1401–1414. doi: 10.1093/nar/12.3.1401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hernandez N., Lucito R. Elements required for transcription initiation of the human U2 snRNA gene coincide with elements required for snRNA 3' end formation. EMBO J. 1988 Oct;7(10):3125–3134. doi: 10.1002/j.1460-2075.1988.tb03179.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hipskind R. A., Nordheim A. In vitro transcriptional analysis of the human c-fos proto-oncogene. J Biol Chem. 1991 Oct 15;266(29):19572–19582. [PubMed] [Google Scholar]
- Horwitz R. J., Li J., Greenblatt J. An elongation control particle containing the N gene transcriptional antitermination protein of bacteriophage lambda. Cell. 1987 Nov 20;51(4):631–641. doi: 10.1016/0092-8674(87)90132-2. [DOI] [PubMed] [Google Scholar]
- Innis J. W., Kellems R. E. A heat-labile factor promotes premature 3' end formation in exon 1 of the murine adenosine deaminase gene in a cell-free transcription system. Mol Cell Biol. 1991 Nov;11(11):5398–5409. doi: 10.1128/mcb.11.11.5398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Innis J. W., Moore D. J., Kash S. F., Ramamurthy V., Sawadogo M., Kellems R. E. The murine adenosine deaminase promoter requires an atypical TATA box which binds transcription factor IID and transcriptional activity is stimulated by multiple upstream Sp1 binding sites. J Biol Chem. 1991 Nov 15;266(32):21765–21772. [PubMed] [Google Scholar]
- Izban M. G., Luse D. S. Factor-stimulated RNA polymerase II transcribes at physiological elongation rates on naked DNA but very poorly on chromatin templates. J Biol Chem. 1992 Jul 5;267(19):13647–13655. [PubMed] [Google Scholar]
- Kash S. F., Innis J. W., Jackson A. U., Kellems R. E. Functional analysis of a stable transcription arrest site in the first intron of the murine adenosine deaminase gene. Mol Cell Biol. 1993 May;13(5):2718–2729. doi: 10.1128/mcb.13.5.2718. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato H., Sumimoto H., Pognonec P., Chen C. H., Rosen C. A., Roeder R. G. HIV-1 Tat acts as a processivity factor in vitro in conjunction with cellular elongation factors. Genes Dev. 1992 Apr;6(4):655–666. doi: 10.1101/gad.6.4.655. [DOI] [PubMed] [Google Scholar]
- Kephart D. D., Marshall N. F., Price D. H. Stability of Drosophila RNA polymerase II elongation complexes in vitro. Mol Cell Biol. 1992 May;12(5):2067–2077. doi: 10.1128/mcb.12.5.2067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerppola T. K., Kane C. M. Analysis of the signals for transcription termination by purified RNA polymerase II. Biochemistry. 1990 Jan 9;29(1):269–278. doi: 10.1021/bi00453a037. [DOI] [PubMed] [Google Scholar]
- Kerppola T. K., Kane C. M. Intrinsic sites of transcription termination and pausing in the c-myc gene. Mol Cell Biol. 1988 Oct;8(10):4389–4394. doi: 10.1128/mcb.8.10.4389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kessler M., Aloni Y. The block to transcription elongation at the SV40 attenuation site is decreased in vitro by oligomers complementary to segments of the attenuator RNA. Gene. 1989 Dec 7;84(1):65–72. doi: 10.1016/0378-1119(89)90140-6. [DOI] [PubMed] [Google Scholar]
- Krauskopf A., Bengal E., Aloni Y. The block to transcription elongation at the minute virus of mice attenuator is regulated by cellular elongation factors. Mol Cell Biol. 1991 Jul;11(7):3515–3521. doi: 10.1128/mcb.11.7.3515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuhn A., Bartsch I., Grummt I. Specific interaction of the murine transcription termination factor TTF I with class-I RNA polymerases. Nature. 1990 Apr 5;344(6266):559–562. doi: 10.1038/344559a0. [DOI] [PubMed] [Google Scholar]
- Lattier D. L., States J. C., Hutton J. J., Wiginton D. A. Cell type-specific transcriptional regulation of the human adenosine deaminase gene. Nucleic Acids Res. 1989 Feb 11;17(3):1061–1076. doi: 10.1093/nar/17.3.1061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee H., Kraus K. W., Wolfner M. F., Lis J. T. DNA sequence requirements for generating paused polymerase at the start of hsp70. Genes Dev. 1992 Feb;6(2):284–295. doi: 10.1101/gad.6.2.284. [DOI] [PubMed] [Google Scholar]
- London L., Keene R. G., Landick R. Analysis of premature termination in c-myc during transcription by RNA polymerase II in a HeLa nuclear extract. Mol Cell Biol. 1991 Sep;11(9):4599–4615. doi: 10.1128/mcb.11.9.4599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maa M. C., Chinsky J. M., Ramamurthy V., Martin B. D., Kellems R. E. Identification of transcription stop sites at the 5' and 3' ends of the murine adenosine deaminase gene. J Biol Chem. 1990 Jul 25;265(21):12513–12519. [PubMed] [Google Scholar]
- Marshall N. F., Price D. H. Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol Cell Biol. 1992 May;12(5):2078–2090. doi: 10.1128/mcb.12.5.2078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McStay B., Reeder R. H. A DNA-binding protein is required for termination of transcription by RNA polymerase I in Xenopus laevis. Mol Cell Biol. 1990 Jun;10(6):2793–2800. doi: 10.1128/mcb.10.6.2793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mechti N., Piechaczyk M., Blanchard J. M., Jeanteur P., Lebleu B. Sequence requirements for premature transcription arrest within the first intron of the mouse c-fos gene. Mol Cell Biol. 1991 May;11(5):2832–2841. doi: 10.1128/mcb.11.5.2832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meulia T., Krumm A., Groudine M. Distinct properties of c-myc transcriptional elongation are revealed in Xenopus oocytes and mammalian cells and by template titration, 5,6-dichloro-1-beta-D-ribofuranosylbenzimidazole (DRB), and promoter mutagenesis. Mol Cell Biol. 1993 Sep;13(9):5647–5658. doi: 10.1128/mcb.13.9.5647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mok M., Maderious A., Chen-Kiang S. Premature termination by human RNA polymerase II occurs temporally in the adenovirus major late transcriptional unit. Mol Cell Biol. 1984 Oct;4(10):2031–2040. doi: 10.1128/mcb.4.10.2031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Brien T., Lis J. T. RNA polymerase II pauses at the 5' end of the transcriptionally induced Drosophila hsp70 gene. Mol Cell Biol. 1991 Oct;11(10):5285–5290. doi: 10.1128/mcb.11.10.5285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramamurthy V., Maa M. C., Harless M. L., Wright D. A., Kellems R. E. Sequence requirements for transcriptional arrest in exon 1 of the murine adenosine deaminase gene. Mol Cell Biol. 1990 Apr;10(4):1484–1491. doi: 10.1128/mcb.10.4.1484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ratnasabapathy R., Sheldon M., Johal L., Hernandez N. The HIV-1 long terminal repeat contains an unusual element that induces the synthesis of short RNAs from various mRNA and snRNA promoters. Genes Dev. 1990 Dec;4(12A):2061–2074. doi: 10.1101/gad.4.12a.2061. [DOI] [PubMed] [Google Scholar]
- Reines D., Mote J., Jr Elongation factor SII-dependent transcription by RNA polymerase II through a sequence-specific DNA-binding protein. Proc Natl Acad Sci U S A. 1993 Mar 1;90(5):1917–1921. doi: 10.1073/pnas.90.5.1917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnekov O., Aloni Y. RNA polymerase II is capable of pausing and prematurely terminating transcription at a precise location in vivo and in vitro. Proc Natl Acad Sci U S A. 1989 Jan;86(1):12–16. doi: 10.1073/pnas.86.1.12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnekov O., Pruzan R., Aloni Y. Elements involved in an in vitro block to transcription elongation at the end of the L1 mRNA family of adenovirus 2. Nucleic Acids Res. 1991 Apr 25;19(8):1783–1790. doi: 10.1093/nar/19.8.1783. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts S., Bentley D. L. Distinct modes of transcription read through or terminate at the c-myc attenuator. EMBO J. 1992 Mar;11(3):1085–1093. doi: 10.1002/j.1460-2075.1992.tb05147.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rougvie A. E., Lis J. T. The RNA polymerase II molecule at the 5' end of the uninduced hsp70 gene of D. melanogaster is transcriptionally engaged. Cell. 1988 Sep 9;54(6):795–804. doi: 10.1016/s0092-8674(88)91087-2. [DOI] [PubMed] [Google Scholar]
- SivaRaman L., Reines D., Kane C. M. Purified elongation factor SII is sufficient to promote read-through by purified RNA polymerase II at specific termination sites in the human histone H3.3 gene. J Biol Chem. 1990 Aug 25;265(24):14554–14560. [PubMed] [Google Scholar]
- Skarnes W. C., Tessier D. C., Acheson N. H. RNA polymerases stall and/or prematurely terminate nearby both early and late promoters on polyomavirus DNA. J Mol Biol. 1988 Sep 5;203(1):153–171. doi: 10.1016/0022-2836(88)90099-x. [DOI] [PubMed] [Google Scholar]
- Toohey M. G., Jones K. A. In vitro formation of short RNA polymerase II transcripts that terminate within the HIV-1 and HIV-2 promoter-proximal downstream regions. Genes Dev. 1989 Mar;3(3):265–282. doi: 10.1101/gad.3.3.265. [DOI] [PubMed] [Google Scholar]
- Watson R. J. Expression of the c-myb and c-myc genes is regulated independently in differentiating mouse erythroleukemia cells by common processes of premature transcription arrest and increased mRNA turnover. Mol Cell Biol. 1988 Sep;8(9):3938–3942. doi: 10.1128/mcb.8.9.3938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wiest D. K., Wang D., Hawley D. K. Mechanistic studies of transcription arrest at the adenovirus major late attenuation site. Comparison of purified RNA polymerase II and washed elongation complexes. J Biol Chem. 1992 Apr 15;267(11):7733–7744. [PubMed] [Google Scholar]
- Wright S., Mirels L. F., Calayag M. C., Bishop J. M. Premature termination of transcription from the P1 promoter of the mouse c-myc gene. Proc Natl Acad Sci U S A. 1991 Dec 15;88(24):11383–11387. doi: 10.1073/pnas.88.24.11383. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wright S. Regulation of eukaryotic gene expression by transcriptional attenuation. Mol Biol Cell. 1993 Jul;4(7):661–668. doi: 10.1091/mbc.4.7.661. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanofsky C. Transcription attenuation. J Biol Chem. 1988 Jan 15;263(2):609–612. [PubMed] [Google Scholar]
- de Vegvar H. E., Lund E., Dahlberg J. E. 3' end formation of U1 snRNA precursors is coupled to transcription from snRNA promoters. Cell. 1986 Oct 24;47(2):259–266. doi: 10.1016/0092-8674(86)90448-4. [DOI] [PubMed] [Google Scholar]