Abstract
Listeria monocytogenes strains belonging to serotypes 1/2a and 4b are frequently linked to listeriosis. While inlA mutations leading to premature stop codons (PMSCs) and attenuated virulence are common in 1/2a, they are rare in serotype 4b. We observed PMSCs in 35% of L. monocytogenes isolates (n = 54) recovered from the British Columbia food supply, including serotypes 1/2a (30%), 1/2c (100%), and 3a (100%), and a 3-codon deletion (amino acid positions 738 to 740) seen in 57% of 4b isolates from fish-processing facilities. Caco-2 invasion assays showed that two isolates with the deletion were significantly more invasive than EGD-SmR (P < 0.0001) and were either as (FF19-1) or more (FE13-1) invasive than a clinical control strain (08-5578) (P = 0.006). To examine whether serotype 1/2a was more likely to acquire mutations than other serotypes, strains were plated on agar with rifampin, revealing 4b isolates to be significantly more mutable than 1/2a, 1/2c, and 3a serotypes (P = 0.0002). We also examined the ability of 33 strains to adapt to cold temperature following a downshift from 37°C to 4°C. Overall, three distinct cold-adapting groups (CAG) were observed: 46% were fast (<70 h), 39% were intermediate (70 to 200 h), and 15% were slow (>200 h) adaptors. Intermediate CAG strains (70%) more frequently possessed inlA PMSCs than did fast (20%) and slow (10%) CAGs; in contrast, 87% of fast adaptors lacked inlA PMSCs. In conclusion, we report food chain-derived 1/2a and 4b serotypes with a 3-codon deletion possessing invasive behavior and the novel association of inlA genotypes encoding a full-length InlA with fast cold-adaptation phenotypes.
INTRODUCTION
Listeria monocytogenes is an environmentally ubiquitous organism that frequently contaminates food-processing environments. It is estimated that 99% of listeriosis cases are transmitted through the consumption of contaminated food (1–3). In healthy individuals, L. monocytogenes infections are rare, are restricted to the gastrointestinal environment, are self-limiting, and manifest as gastroenteritis and/or mild flu-like symptoms. In contrast, in susceptible populations (e.g., neonates, the elderly, and the immunocompromised), infections become invasive, leading to encephalitis, meningitis, septicemia, and/or spontaneous abortions during the last trimester of pregnancy (4); mortality rates for invasive listeriosis typically range from 20 to 40% (5–7).
Although there are 13 L. monocytogenes serotypes, 1/2a, 1/2b, and 4b account for the majority of human disease (3, 8). Historically, lineage I 4b strains have been overrepresented in clinical listeriosis cases and are less frequently recovered from foods (8–10). However, over the past decade, lineage II 1/2a strains typically overrepresented in food and environmental samples (3, 9, 11) have been frequently linked to human disease, causing notable outbreaks in Switzerland (12) and the United Kingdom and two 2008 outbreaks in Canada (7, 13). With regard to the Canadian outbreaks, 1/2a strains comprise the majority of Canadian clinical isolates, followed by 4b (14). Reasons for the prevalence of 1/2a strains in human disease in Canada may be linked to a recently identified clonal complex/epidemic 1/2a clone that was identified as a recurring cause of sporadic listeriosis since 1988 (15). Within this complex, the majority of 1/2a strains were found to possess the LGI1 genomic island, which was first identified in an outbreak strain linked to 23 deaths (16) and more recently recovered from smoked salmon in British Columbia (BC) (17).
Over the past decade, sequence analysis of inlA, which encodes a membrane-bound protein facilitating invasion into nonprofessional phagocytes, reveals that a significant proportion (45%) of strains recovered from ready-to-eat (RTE) food possess mutations resulting in a premature stop codon (PMSC) (18–20). Strains with inlA PMSCs produce either a truncated or nonsecreted InlA, resulting in virulence-attenuated phenotypes as measured by in vitro cell assays (21–25) and mammalian models (18, 26, 27).
While infrequent reporting of PMSCs in 4b strains may reflect their overrepresentation in clinical listeriosis, the contrary is postulated for 1/2a strains. Being frequently recovered in food and food production environments, positive selection for strains with PMSCs may serve as a phase switch that is important for environmental survival (28). In line with this, it has been suggested that lineage II 1/2a strains are better able to survive conditions associated with the food chain. Notably, 1/2a and 1/2c (lineage II) more frequently possess PMSCs than 1/2b or 3b serotypes, with 4b strains rarely having PMSCs (28). In general, serotype 4b appears more recalcitrant to genetic flux, being less likely to acquire or possess plasmids and to experience homologous recombination events that may afford rapid adaptation to niche-specific stresses.
In this study, we were interested in observing how L. monocytogenes strains with differing inlA genotypes respond to food chain-relevant conditions. In particular, the reliance on refrigeration to maintain the quality of fresh and RTE foods makes cold temperature a suitable and relevant parameter to examine. To this end, we ascertained the nature and prevalence of inlA genotypes in L. monocytogenes serotypes recovered from food production environments and food in BC, Canada, and assessed the capacity of strains to acquire adaptive mutations. Further, we explored how L. monocytogenes strains encoding a full-length InlA or carrying an inlA gene with a PMSC were able to adapt and subsequently resume growth at 4°C following a downshift from 37°C.
MATERIALS AND METHODS
Bacterial isolates.
Listeria monocytogenes strains used in this study were recovered from food-processing environment (FPE) swabs (n = 29), raw unprocessed food (RUF; n = 6), and ready-to-eat (RTE) foods (n = 19) that were collected from three dairy-processing (DPF), five fish-processing (FPF), and five meat-processing (MPF) facilities across BC as part of a previous study (29). Using origin of isolation, serotyping, and pulsed-field gel electrophoresis (PFGE) data, we selected 54 different isolates for inclusion in this study. Isolate origins and serotyping data are described in Table 1. Isolates were serotyped by slide agglutination, antisera were prepared according to Seeliger and Höhne (30) at the Canadian National Microbiology Laboratory, and PFGE was performed by the BCCDC Public Health Microbiology and Reference Laboratory. Bacterial cultures were maintained in peptone with 20% glycerol at −70°C. Prior to experiments, isolates were grown overnight on tryptic soy agar (TSA; Difco, Becton, Dickinson Diagnostics, Mississauga, ON, Canada) at 35°C.
Table 1.
Serotypes of Listeria monocytogenes strains recovered from food-processing environments, raw unprocessed foods, or ready-to-eat foods
| Source | No. of L. monocytogenes strains with each serotype |
|||||
|---|---|---|---|---|---|---|
| Total | 1/2a | 1/2b | 1/2c | 3a | 4b | |
| Processing environment | 29 | 11 | 1 | 5 | 4 | 8 |
| Fooda | ||||||
| RUF | 6 | 4 | 0 | 2 | 0 | 0 |
| RTE | 19 | 5 | 0 | 1 | 0 | 13 |
| Total | 54 | 20 | 1 | 8 | 4 | 21 |
RUF, raw unprocessed food; RTE, ready-to-eat food products.
Internalin A DNA sequencing.
Conventional PCRs were used to amplify the 2.4-kb inlA gene as described by Kovacevic et al. (17). Briefly, 5 U of AmpliTaq Gold 360 DNA polymerase (Invitrogen, Burlington, ON) was used with one set of custom primers (inlA-JK-F, 5′-TAC AAC GAA ACC TGA TAT TG-3′, and inlA-JK-R, 5′-GCT AGA TAT AGT CCG AAA AC-3′), each at 0.5 mM, 200 mM deoxynucleoside triphosphates (dNTPs) (Invitrogen), and 50 to 100 ng DNA template obtained using a DNeasy blood and tissue kit (Qiagen, Toronto, ON). Thermocycling was performed as follows: initial denaturation at 94°C for 2 min; 20 cycles of 94°C for 1 min, 60 to 50°C for 1 min with a touchdown decrease of 0.5°C per cycle, and 72°C for 2.5 min; 20 cycles of 94°C for 1 min, 50°C for 1 min, and 72°C for 2.5 min; and a final extension step at 72°C for 7 min (19). The PCR product was purified using a QIAquick PCR purification kit (Qiagen) and sequenced at Canada's Michael Smith Genome Science Centre using the inlA-JK primer set and previously published primers (19). Nucleotide sequences were assembled and analyzed with Geneious 5.4 software (Biomatters Ltd., Auckland, New Zealand). The presence of PMSCs was determined by comparing inlA sequence data to that of L. monocytogenes EGD-e (31).
Invasion of Caco-2 cells.
The invasion efficiencies of seven representative L. monocytogenes isolates from different serotypes, food and environmental samples, and inlA genotypes were assessed in 24-well tissue culture plates according to the method of Gaillard et al. (32), with minor modifications. Briefly, Caco-2 cells (∼2 × 105 cells per well; passages 5 to 20) were cultured in Dulbecco's modified Eagle's minimum essential medium (DMEM) (HyClone; Thermo Scientific, Toronto, ON) supplemented with 10% inactivated fetal calf serum (Gibco, Life Technologies, Burlington, ON), 1% nonessential amino acids (Gibco), and 1% GlutaMAX (Gibco) for 2 days (5% CO2, 37°C) to reach confluence. Bacterial cultures grown statically overnight in brain heart infusion (BHI) at 30°C were pelleted by centrifugation (5,939 × g; Eppendorf 5415 R), washed once, rehydrated in 1× Dulbecco's phosphate-buffered saline (DPBS) (HyClone) with magnesium and calcium, and adjusted to an optical density at 600 nm (OD600) of 0.5 (Genesys 10UV; Thermo Spectronic, Rochester, NY). Prior to infection, cultures were diluted in DMEM to approximately 4 × 107 CFU per ml, as assessed by plating on TSA. Bacterial suspensions (0.5 ml) were added to Caco-2 cells and incubated at 37°C for 1 h to allow bacterial entry. Cells were washed three times, overlaid with fresh DMEM containing gentamicin (10 mg/liter), and incubated at 37°C for 2 h. Following incubation, the cell monolayer was washed three times with DPBS and treated with 1% Triton X-100 for 10 min at 37°C. The number of viable bacteria released was quantified by plating them onto TSA. L. monocytogenes EGD-SmR and BUG5 (Tn1545-induced inlA mutant from EGD-SmR) (33) and 08-5578 (16), kindly provided by Pascale Cossart (Institut Pasteur) and Matthew Gilmour (Public Health Agency of Canada), respectively, were used as controls. The gentamicin concentration used (10 mg/liter) was confirmed to kill all extracellular bacteria by plating of the inoculum onto TSA. Invasion assays for each isolate were carried out in triplicate and repeated two times.
Mutation frequency.
Since differences in the occurrence of PMSCs exist among different serotypes, we examined how frequently isolates (n = 54) acquired mutations following exposure to rifampin (RIF) by using previously published methodology described for Enterobacteriaceae, with some modification (34, 35). Briefly, isolates were grown overnight at 35°C in BHI broth and adjusted to an OD600 of 1.0. A 100-μl aliquot was spread-plated onto BHI agar plates with 100 μg/ml RIF. Following incubation for 48 h at 35°C, the number of CFU was counted. The assay for each isolate was carried out in triplicate and repeated two times. The mean number of colonies for all strains was determined, and comparisons were made between strains with and without PMSCs and across serotypes.
Cold growth evaluation.
A subset of isolates (n = 33) representing L. monocytogenes strains with full-length inlA, a 3-codon deletion (amino acids [aa] 738 to 740), and each type of PMSC observed in our collection was assessed for cold growth adaptation as described by Arguedas-Villa et al. (36). In short, a single colony was inoculated into 10 ml brain heart infusion broth (Oxoid, Ottawa, ON) and grown overnight at 37°C with shaking (220 rpm) (∼109 CFU/ml). Fresh BHI (10 ml) was inoculated with approximately 103 CFU/ml and incubated at 4°C until bacteria reached stationary phase. Growth was monitored by plating 10-fold serial dilutions prepared in peptone-buffered saline onto plate count agar (Oxoid), incubating them at 37°C for 24 h, and counting CFU. The lag phase duration (LPD) and exponential growth rate (EGR) of each strain were calculated from log-converted growth (CFU/ml) data using the Dmfit version 2.0 and Microfit version 1.0 programs, based on the models of Baranyi and Roberts (37).
Statistical analysis.
Data analysis was performed using GraphPad Prism 6.0 software. The statistical significance of differences in inlA genotypes based on isolate source (FPE foods, RUF, and RTE foods) was assessed using chi-square and Fisher's exact tests (RUF and RTE foods). The Student t test was used to compare invasive inlA genotypes to control strains (08-5578, EGD-SmR, or BUG5) and to examine whether differences exist in LPD and EGR between food and environmental strains. Mutability, as indicated by the numbers of rifampin-resistant (RIFr) colonies, among serotypes (1/2a, 1/2c, 3a, and 4b) was compared using the Kruskal-Wallis test for nonparametric data, followed by Dunn's multiple-comparisons test, while differences between inlA genotypes (no PMSCs versus PMSCs) were assessed by the Mann-Whitney test. Fisher's exact test was performed to assess whether differences exist between cold-adapting groups (fast, intermediate) and inlA genotypes (no PMSCs versus PMSCs). For all analyses, differences were considered significant if the P value was <0.05.
Nucleotide sequence accession numbers.
The nucleotide sequences from the isolates in this study have been deposited in GenBank under numbers KC433332 to KC433385.
RESULTS
inlA genotypes and mutability among L. monocytogenes strains.
DNA sequencing of inlA in 54 L. monocytogenes strains originating from food and FPE samples recovered from dairy-, fish-, and meat-processing facilities revealed PMSCs in 35% of isolates (GenBank accession numbers KC433332 to KC433385). Isolates possessing truncated InlA due to PMSC mutations are hereafter referred to as PMSC isolates. Type 3 mutations (aa position 700) were the most common PMSC mutation in this collection (53%), followed by type 4 (32%) (aa position 9) and type 11 (10%) (aa position 685), and only a single isolate (5%) possessed a type 1 mutation (aa position 606) (Table 2) (19). Overall, 41% of isolates carried a full-length inlA, while 24% had a 3-codon deletion in aa positions 738 to 740 (aspartic acid, threonine, and serine), hereafter referred to as the 3-codon deletion.
Table 2.
Number of L. monocytogenes isolates recovered from food-processing environments, raw unprocessed foods (RUF), and ready-to-eat (RTE) foods with inlA mutations resulting in premature stop codons or 3-codon deletions
| inlA genotype | No. (%) of environmental isolates (n = 29) | No. (%) of food isolates |
Serotype(s) (no. of strains) | Facility IDb | |
|---|---|---|---|---|---|
| RUF (n = 6) | RTE (n = 19) | ||||
| Without PMSCsa | 16 (55) | 2 (33) | 17 (89) | 1/2a (14), 4b (21) | D5, D7, D11, F20, F21, F28, M38, M44, M46, M49 |
| With 3-codon deletion (aa 738–740) | 4 (14) | 0 | 9 (47) | 1/2a (1), 4b (12) | F20, F31 |
| With PMSCs | 13 (45) | 4 (67) | 2 (11) | ||
| Type 1 (aa 606) | 1 (3) | 0 | 0 | 1/2b (1) | F49 |
| Type 3 (aa 700) | 7 (24) | 2 (33) | 1 (5) | 1/2a (6), 3a (4) | F19, F21, F49, F50 |
| Type 4 (aa 9) | 5 (17) | 0 | 1 (5) | 1/2c (6) | F19 |
| Type 11 (aa 685) | 0 | 2 (33) | 0 | 1/2c (2) | F38, F50 |
Includes isolates with the 3-codon deletion (aa 738 to 740).
ID, identification.
Since PMSCs have been frequently reported in serotype 1/2a strains, we compared the mutability of 1/2a isolates to that of other serotypes, including 4b. Point mutations occurring in the RIF resistance-determining region of rpoB may afford resistance to RIF (RIFr) (38). Following plating in the presence of 100 μg/ml RIF, the number of RIFr colonies ranged from 1.3 ± 0.7 to 52.8 ± 7.9 per plate. Irrespective of serotype, significantly more RIFr colonies were observed in strains not possessing PMSCs than in those with PMSC mutations (P = 0.001) (Fig. 1A). Correspondingly, significantly more RIFr colonies were observed for 4b serotype strains than for 1/2a, 1/2c, and 3a strains (P = 0.0002) (Fig. 1B).
Fig 1.
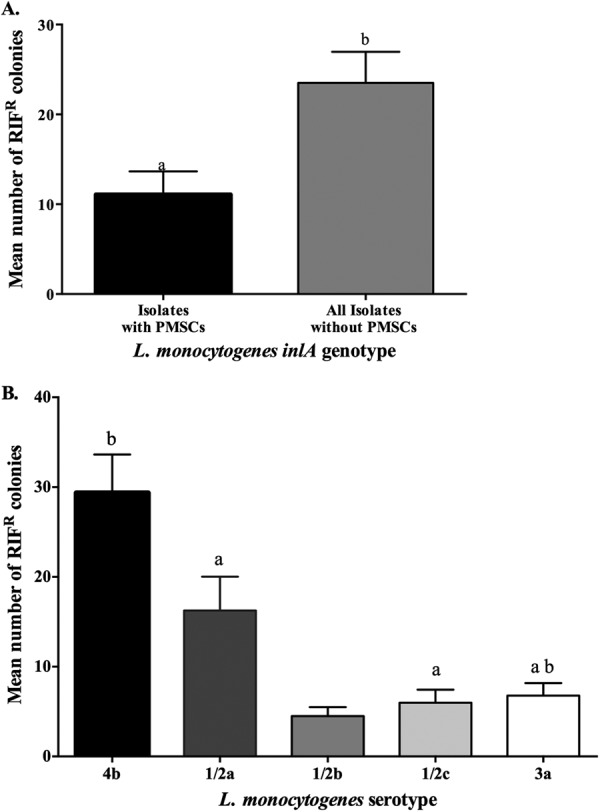
Mutability of different L. monocytogenes inlA genotypes (A) and serotypes (B), assessed by the number of rifampin-resistant colonies after 48 h of growth at 35°C in the presence of 100 μg/ml rifampin. Mutability of each isolate was assayed in triplicate in each experiment, and two independent experiments were performed. Bars show mean numbers of colonies, and error bars indicate standard errors of the mean. Different letters above the bars represent significant differences (P < 0.05) between geno- and serotype groups, determined using the Mann-Whitney test (A) or the Kruskal-Wallis test followed by Dunn's multiple-comparisons test (B). Serotype 1/2b was excluded from statistical analysis, as only one isolate was recovered.
Distribution of inlA genotypes across different food production facilities.
No PMSCs were seen in inlA of L. monocytogenes isolates recovered from DPF, while 33% and 60% of isolates from FPF and MPF, respectively, had PMSCs. Of five FPFs examined, three facilities had isolates lacking PMSCs while two facilities had PMSCs in all (10/10) or 75% of recovered isolates (Table 3). More MPFs (n = 3) had L. monocytogenes isolates possessing inlA PMSCs than had isolates without mutations. Two MPFs had no PMSC mutations in their isolates, one had PMSCs in all isolates (n = 3), and two facilities had 50% and 67% of isolates with mutations (Table 3). Type 3 mutations were seen among isolates from FPF and MPF, while type 4 was seen only in isolates from FPF and types 1 and 11 were seen only in L. monocytogenes strains from MPF. Type 3 mutations were the most common among isolates possessing PMSCs from FPF, followed by type 4 mutations. Similarly, type 3 mutations were most commonly seen in isolates from MFP, followed by type 11 and type 1 mutations (see Table S1 in the supplemental material).
Table 3.
Distribution of L. monocytogenes isolates (n = 54) and the presence of inlA premature stop codons across dairy-processing (DPF), fish-processing (FPF), and meat-processing (MPF) facilities (n = 13)
| Facility ID | Facility type | Total no. of isolates | No. of isolates from each sourcea |
inlA presence (no. positive/no. tested) |
|||
|---|---|---|---|---|---|---|---|
| FPE | RUF | RTE | No PMSCsb | With PMSCs | |||
| D5 | DPF | 3 | 3 | 3/3 | |||
| D7 | DPF | 1 | 1 | 1/1 | |||
| D11 | DPF | 1 | 1 | 1/1 | |||
| F19 | FPF | 10 | 9 | 1 | 10/10 | ||
| F20 | FPF | 8 | 4 | 4 | 8/8 | ||
| F21 | FPF | 4 | 2 | 2 | 1/4 | 3/4 | |
| F28 | FPF | 7 | 4 | 3 | 7/7 | ||
| F31 | FPF | 10 | 1 | 9 | 10/10 | ||
| M38 | MPF | 2 | 2 | 1/2 | 1/2 | ||
| M44 | MPF | 1 | 1 | 1/1 | |||
| M46 | MPF | 1 | 1 | 1/1 | |||
| M49 | MPF | 3 | 3 | 1/3 | 2/3 | ||
| M50 | MPF | 3 | 3 | 3/3 | |||
FPE, food-processing environment; RUF, raw unprocessed food; RTE, ready-to-eat food.
PMSCs, premature stop codons.
Isolates possessing a 3-codon deletion were seen in two FPFs (F20, F31) but not in DPFs or MPFs (Table 2). In one of the facilities (F20), all but one isolate (88%) had this deletion, while 60% of samples had the same codons missing in the other facility (F31). PFGE typing showed that these isolates were not clonal (data not shown). None of the isolates from facilities F20 and F31 possessed PMSCs.
inlA mutations within different serotypes.
PMSC mutations were seen in four of the five serotypes examined, including all 1/2c (n = 8) and 3a (n = 4) isolates, 1/2a (30%; n = 6), and the only 1/2b (n = 1) isolate examined. Serotypes 1/2a and 3a carried only type 3 mutations, while serotype 1/2b had only a type 1 mutation. The only serotype with more than one type of mutation (i.e., types 4 and 11) was 1/2c.
A 3-codon deletion was observed in 13 strains derived from FPFs. With the exception of one 1/2a isolate, this deletion was observed exclusively in 4b serotype strains. Overall, 57% of 4b serotype isolates possessed this 3-codon deletion, though no PMSC mutations were seen in serotype 4b isolates.
Occurrence of inlA PMSCs in isolates recovered from different sources.
Isolates with inlA PMSC mutations were seen more commonly in FPE samples than RUF and RTE foods (P = 0.01). Further, more isolates from RUF (67%) carried PMSCs than those isolated from RTE (16%) foods (P = 0.02). Nine of the 13 3-codon deletion mutants were recovered from RTE food (69%), with the remaining isolates being environmental. Isolates encoding full-length InlA were seen predominantly in FPE samples (55%), followed by RTE (36%) and RUF (9%) samples.
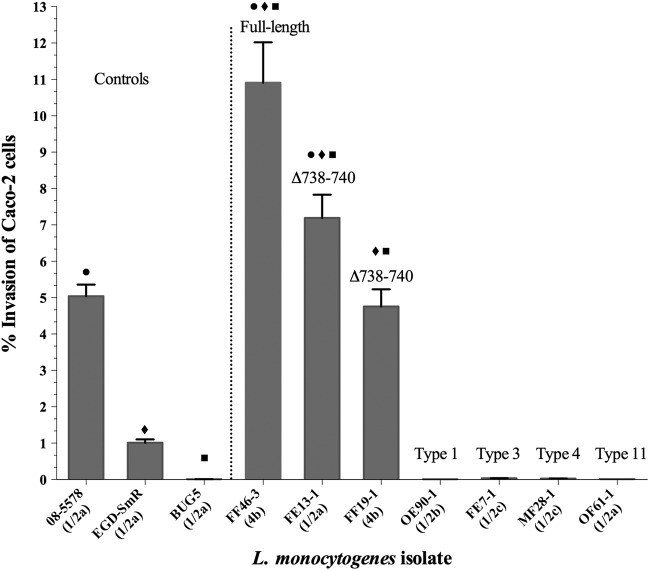
Invasion of Caco-2 cells by L. monocytogenes strains possessing truncated InlA or a 3-codon deletion.
PMSC-encoding isolates exhibited reduced Caco-2 cell invasion phenotypes (Fig. 2). A 4b isolate (FF46-3) possessing wild-type inlA was 2.2 times more invasive (P < 0.0001) than a clinical isolate that claimed 23 lives during a 2008 deli meat outbreak in Canada and 10.8 times more invasive (P < 0.0001) than the laboratory control EGD-SmR strain (Fig. 2). This phenomenon was observed for another 4b isolate (FF19-1) and a 1/2a strain (FE13-1) possessing the 3-codon deletion, which were 4.7 and 7.1 times more invasive (P < 0.0001), respectively, than the control EDG-SmR strain. Compared to the Canadian deli meat outbreak strain, FF19-1 and FE13-1 were as invasive and 1.4 times more invasive (P = 0.006), respectively (Fig. 2).
Fig 2.
Invasion efficiencies (% of bacteria recovered relative to the initial inoculum) of L. monocytogenes isolates possessing inlA PMSC mutations (types 1, 3, 4, and 11) or a 3-codon deletion at amino acid positions 738 to 740 (Δ738-740) compared to those of wild-type clinical isolates (08-5578 and EGD-SmR) and a Tn1545-induced noninvasive inlA mutant of EGD-SmR (BUG5). Assays for each isolate were carried out in triplicate and repeated two times. Bars show mean invasion efficiencies, and error bars indicate standard errors of the mean. Different symbols above the bars indicate significantly higher invasion efficiencies (P < 0.05; t test) compared to those of the controls 08-5578 (●), EGD-SmR (◆), and BUG5 (■).
Cold growth adaptation and growth of strains from different serogroups and sources.
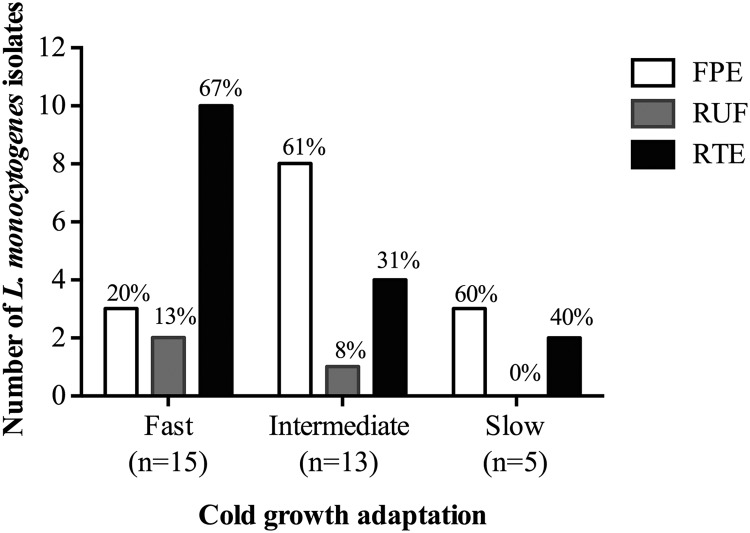
Three cold growth categories were observed among 33 isolates assessed for their ability to adapt to 4°C following a downshift from 37°C. The first category included fast-adapting strains (n = 15) possessing an LPD less than 70 h. The second group was comprised of intermediate cold growth adaptors with an LPD ranging between 70 and 200 h and included the majority of strains (n = 13). Finally, five strains adapted slowly to 4°C, possessing an LPD of >200 h. Fast-adapting strains included mainly RTE food-derived isolates and, to a lesser degree, FPE and RUF isolates, while intermediate cold growth adaptors were recovered predominantly from FPE but also included isolates from RTE foods and RUF (P > 0.05) (Fig. 3). Slow-growing strains were seen only in FPE and RTE samples (Fig. 3). No significant differences were observed in LPD or EGR between food and environmental isolates (Fig. 4).
Fig 3.
Cold growth adaptation of L. monocytogenes isolates recovered from food-processing environments (FPE), raw unprocessed foods (RUF), and ready-to-eat (RTE) foods when grown at 4°C. Differences were not statistically significant (P > 0.05; chi-square test).
Fig 4.
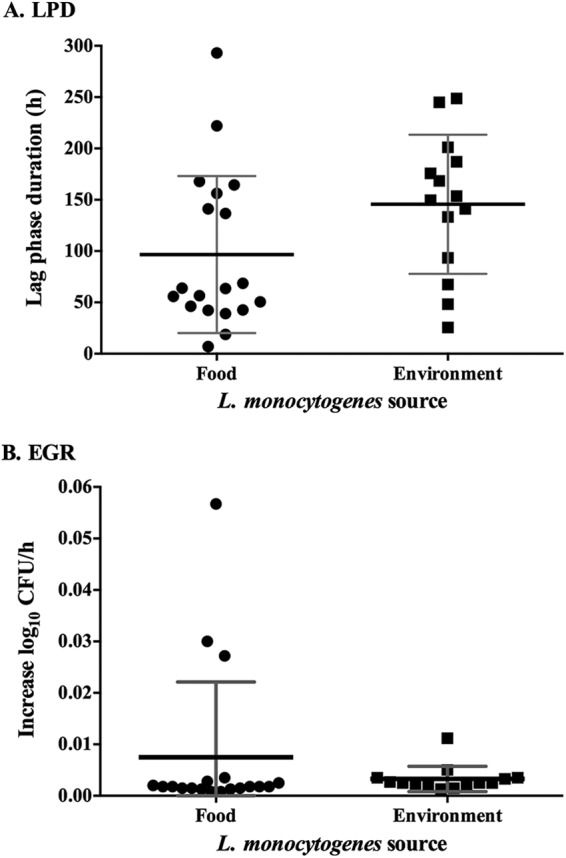
Lag phase durations (A) and exponential growth rates (B) of 33 L. monocytogenes isolates recovered from food and food-processing environments following a downshift from 37°C to 4°C in BHI. Each isolate was assayed in duplicate, and two independent growth assays were performed. Scatter plots show mean values with standard deviations.
The majority of fast-adapting strains were serotype 4b (53%), followed by 1/2a (40%) and 1/2c (7%) serotypes. Intermediate cold adaptors were represented predominantly by 1/2a strains (46%), followed by the 1/2c, 4b, 3a, and 1/2b serotypes. Of the five slow-adapting strains, three were 1/2a and two were 4b.
Cold growth adaptation of different L. monocytogenes inlA genotypes.
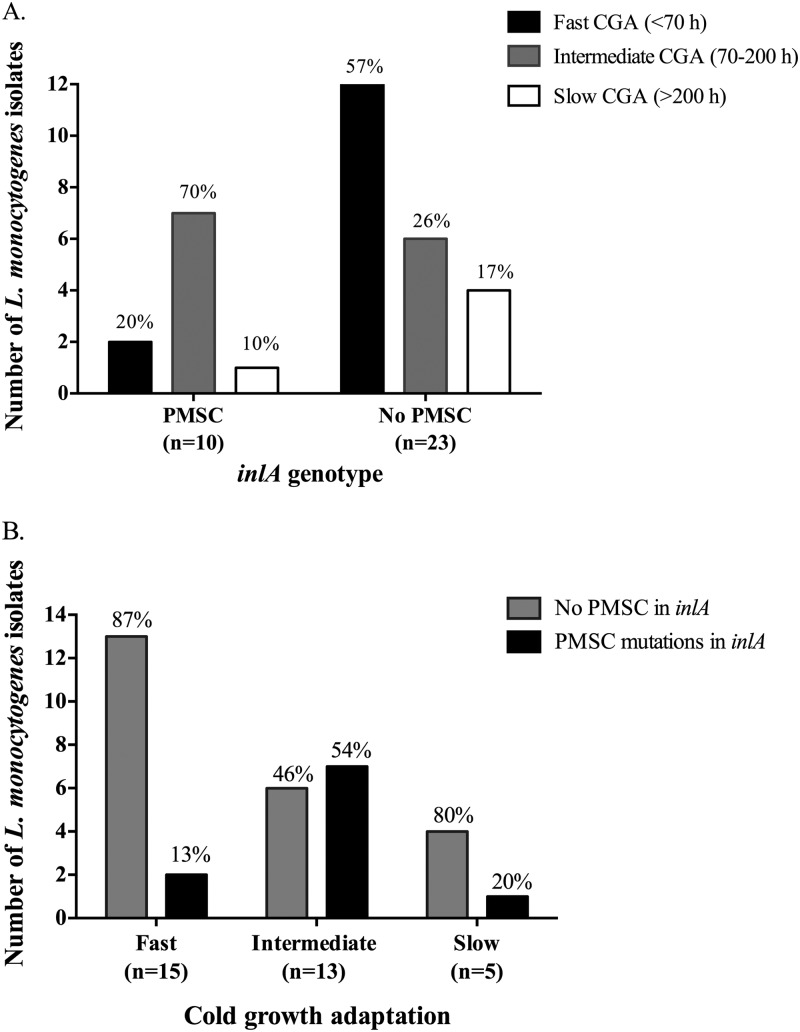
Significant differences in the ability to adapt and grow at 4°C between isolates with and without inlA PMSCs were observed (fast versus intermediate, P = 0.04). Overall, intermediate cold-adapting strains more frequently possessed inlA PMSCs (70%) than did fast (20%) and slow (10%) cold adaptors (Fig. 5A). In contrast, with the exception of two isolates (serotype 1/2c and 1/2a), fast-adapting strains lacked PMSCs (Fig. 5B). Notably, isolates possessing wild-type inlA (i.e., full-length InlA) or the 3-codon deletion comprised 57% of fast-adapting strains, 26% of intermediate strains, and 17% of slow-growing strains (Fig. 5A).
Fig 5.
Distribution of inlA genotypes (i.e., PMSCs versus no PMSCs) (A) and identification of cold-adaptive groups (i.e., fast, intermediate, or slow) (B) observed at 4°C following a downshift from 37°C in BHI. Differences in cold growth adaptation between fast and intermediate L. monocytogenes inlA genotypes were significant (Fisher's exact test; P = 0.04). Percentages of isolates within respective inlA genotypes (A) and cold-adaptive groups (B) are indicated above bars.
DISCUSSION
Our data demonstrate variability in inlA genotypes among L. monocytogenes strains recovered from foods and food-processing environments in BC, Canada, that were often unique within food-processing facilities. Overall, 35% of strains possessed mutations in inlA due to PMSCs, a rate which is lower than that reported for food chain isolates in the United States (45%) (20, 39) but similar to levels reported in France (40). In addition to previously described inlA mutations, including types 1, 3, 4, and 11, we observed inlA genotypes with a consecutive 3-codon deletion in the amino acid positions 738 to 740, a phenomenon to date reported only in a single isolate from a meat facility in Portugal (41). It has been suggested that certain PMSC mutations accumulate at the population level, with notable differences between PMSCs occurring in North America and those in European countries (19, 20, 25). Interestingly, we observed isolates with a type 11 (aa 685) PMSC mutation, which to date has not been reported outside France (19–21, 39).
It is well established that frameshift and transition/transversion mutations in inlA can lead to PMSCs, resulting in a truncated or nonsecreted InlA. Strains possessing these genotypes are associated with attenuated virulence (23, 24, 42) and are seen predominantly in L. monocytogenes strains adapted to environmental and food-processing niches (i.e., 1/2a serotype strains) and, to a lesser degree, in clinical strains overrepresented by 4b serotypes (20, 40). In line with this, strains possessing PMSC mutations that were examined in our study were 1/2a, 1/2c and 3a serotypes. In general, 4b strains are typically more conserved in their genetic content, exhibiting lower recombination rates, and are less likely to possess plasmids and extrachromosomal elements (28, 43, 44). Interestingly, when we compared different serotypes in their ability to acquire point mutations leading to RIFr, 4b strains were significantly (P = 0.0002) more likely to gain mutations conferring resistance than were all other serotypes. Considering that 1/2a serotype strains are known to possess mutations in several virulence loci, including actA, inlA, and prfA (reviewed by Orsi et al. [28]), we hypothesized that 1/2a serotype strains would have higher mutability in this assay. In fact, the opposite was observed, although reasons for this are not clear. This is particularly interesting since positive selection, resulting from the acquisition of advantageous mutations, has been reported to contribute to the evolution of numerous genes in 1/2a strains but is less often reported for 4b serotypes (45, 46). Although this assay has not previously been used to examine mutability in L. monocytogenes, it has been used to examine mutation rates in Enterobacteriaceae (34, 35), in which comparison to reference strains was made to identify hypermutability. In our study, we examined the relative abilities of strains to acquire mutations leading to RIFr. Although further work is needed to explore this phenomenon, our results suggest that serotype 4b strains may acquire mutations more readily than 1/2a strains, demonstrating that there must exist selection pressure for the maintenance of internalin A genes encoding full-length InlA for serotype 4b.
Within our collection, we observed highly invasive isolates possessing a 3-codon deletion (aa 738 to 740), which is contrary to a previous report (41). These isolates exhibited invasion efficiencies equivalent to or surpassing that of the deli meat outbreak strain (08-5578) that contributed to the deaths of 23 individuals in 2008 (7) and that of the EGD-SmR strain (Fig. 2). In recent years, strains possessing truncated InlA proteins have been identified as strains with lower invasiveness and, accordingly, suggested to present reduced public risk (18, 24). Although this seems prudent for most inlA genotypes in which PMSCs lead to truncated proteins, this does not apply to the 3-codon deletion we observed. In contrast, although the InlA protein is truncated by three amino acids, isolates possessing it remain equally invasive as or more invasive than control strains, indicating that they are of considerable risk and public health concern. Considering that 12 of 13 3-codon deletions were serotype 4b, we plan on comparing the internalin gene complement of the single 1/2a isolate with this deletion to those of the 4b strains harboring the same inlA genotype. It has been shown experimentally that the deletion of aa 714 to 766, corresponding to the preanchor region, did not reduce the invasiveness when the modified inlA gene had been transferred to Listeria innocua (47). However, the impact of a consecutive deletion of aspartic acid, threonine, and serine in positions 738 to 740 in naturally occurring L. monocytogenes strains has not been described. It is possible that this deletion may affect protein folding in a manner enhancing bacterial interaction with its human cell surface receptor, E-cadherin. However, it is also possible that other virulence-related factors are contributing to invasion. In particular, a host of other internalin genes (inlB, inlC2, inlD, inlE, inlF, inlG, inlH) have been implicated in invasive behavior (28, 48, 49). Of these, inlC2, inlD, inlE, and inlJ are common to lineage I and II strains, while inlF, inlG, and inlH have been observed only in lineage II (28, 49).
In general, it has been proposed that overrepresentation of serotype 1/2a (lineage II) strains in isolates originating from food and FPEs derives from enhanced capacity to survive food chain conditions, although data substantiating this assertion are often conflicting and limited (28). It has been reported, however, that 4b strains incubated at 4°C for 4 weeks and subsequently upshifted to 37°C possessed shorter LPDs than did 1/2a isolates (50). This implies that 4b strains present in foods may quickly adapt to host temperature and, correspondingly, may be more likely to cause disease. In this study, we assessed the ability of various serotypes to adapt and subsequently grow at 4°C following a downshift from 37°C. In the 33 strains examined, we observed three cold-adapting groups, similar to observations previously made among L. monocytogenes strains derived from different origins (36). When sample origin was examined, the majority of fast-adapting strains were recovered from RTE foods, although differences were not significant. It is tempting to speculate that wild-type inlA genotypes (i.e., full-length) may be an indicator of food chain strain fitness. Support for this stems from observations centering on the absence/presence of inlA PMSCs in respective cold growth groups. Significantly more intermediate cold-adapting isolates possessed PMSCs (70%) than did fast-adapting isolates (P = 0.04), with only two fast-adapting isolates shown to encode a PMSC. This observation lends support to the use of inlA as a suitable biomarker to identify high-risk strains, though in this light, it may be used as an indicator of increased ability to adapt and grow at refrigeration temperatures. Considering that cold temperatures are used in RTE-food-processing facilities and relied on to ensure product quality throughout the food supply chain, these strains may possess enhanced ability to persist in FPE. Further, if present in food, strains having an inlA gene producing a full-length InlA may have increased ability to grow to unacceptable and potentially dangerous levels during cold storage, particularly if abusive conditions are encountered.
Conclusions.
In summary, we observed inlA mutations in four L. monocytogenes serotypes recovered from the BC food continuum. Notably, when the mutability of isolates was examined, serotype 4b isolates were shown to acquire point mutations more frequently than all other serotypes. Also, isolates with a 3-codon inlA deletion possessed highly invasive phenotypes, suggesting that this inlA genotype may be a public health concern. When we examined the ability of L. monocytogenes isolates to adapt to cold temperature, we demonstrated that isolates possessing rapid cold adaption were more likely to carry an inlA gene lacking PMSCs. Our findings substantiate in new ways the assertion that isolates lacking inlA PMSCs are a significant concern. In light of our findings, which show that these isolates were more commonly recovered from RTE food, possessed the capacity to more-rapidly adapt and grow at refrigeration than isolates with PMSCs, and genetically possess a causally linked virulence determinant, these isolates are of significant concern to food processors and public health officials.
ACKNOWLEDGMENTS
This work was supported by the National Sciences and Engineering Research Council of Canada, for which we are grateful.
We acknowledge the British Columbia Centre for Disease Control for donating bacterial isolates and for PFGE typing and the Canadian National Microbiology Laboratory for performing serotyping. We also extend our thanks to B. Brett Finlay and Kristie Keeney for their assistance and Pascale Cossart and Matthew Gilmour for donation of L. monocytogenes controls.
Footnotes
Published ahead of print 11 January 2013
Supplemental material for this article may be found at http://dx.doi.org/10.1128/AEM.03341-12.
REFERENCES
- 1. Mead PS, Slutsker L, Dietz V, McCaig LF, Bresee JS, Shapiro C, Griffin PM, Tauxe RV. 1999. Food-related illness and death in the United States. Emerg. Infect. Dis. 5:607–625 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Scallan E, Hoekstra RM, Angulo FJ, Tauxe RV, Widdowson MA, Roy SL, Jones JL, Griffin PM. 2011. Foodborne illness acquired in the United States—major pathogens. Emerg. Infect. Dis. 17:7–15 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Swaminathan B, Gerner-Smidt P. 2007. The epidemiology of human listeriosis. Microbes Infect. 9:1236–1243 [DOI] [PubMed] [Google Scholar]
- 4. McLauchlin J, Mitchell RT, Smerdon WJ, Jewell K. 2004. Listeria monocytogenes and listeriosis: a review of hazard characterisation for use in microbiological risk assessment of foods. Int. J. Food Microbiol. 92:15–33 [DOI] [PubMed] [Google Scholar]
- 5. Farber JM, Peterkin PI. 1991. Listeria monocytogenes, a food-borne pathogen. Microbiol. Rev. 55:476–511 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Hof H, Nichterlein T, Kretschmar M. 1997. Management of listeriosis. Clin. Microbiol. Rev. 10:345–357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Weatherill S. 2009, posting date Listeriosis investigative review. Report of the independent investigator into the 2008 listeriosis outbreak. Agriculture and Agri-Food Canada, Ottawa, Ontario, Canada [Google Scholar]
- 8. Chenal-Francisque V, Lopez J, Cantinelli T, Caro V, Tran C, Leclercq A, Lecuit M, Brisse S. 2011. Worldwide distribution of major clones of Listeria monocytogenes. Emerg. Infect. Dis. 17:1110–1112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Gray MJ, Zadoks RN, Fortes ED, Dogan B, Cai S, Chen Y, Scott VN, Gombas DE, Boor KJ, Wiedmann M. 2004. Listeria monocytogenes isolates from foods and humans form distinct but overlapping populations. Appl. Environ. Microbiol. 70:5833–5841 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Jeffers GT, Bruce JL, McDonough PL, Scarlett J, Boor KJ, Wiedmann M. 2001. Comparative genetic characterization of Listeria monocytogenes isolates from human and animal listeriosis cases. Microbiology 147:1095–1104 [DOI] [PubMed] [Google Scholar]
- 11. Chen J, Zhang X, Mei L, Jiang L, Fang W. 2009. Prevalence of Listeria in Chinese food products from 13 provinces between 2000 and 2007 and virulence characterization of Listeria monocytogenes isolates. Foodborne Pathog. Dis. 6:7–14 [DOI] [PubMed] [Google Scholar]
- 12. Bille J, Blanc DS, Schmid H, Boubaker K, Baumgartner A, Siegrist HH, Tritten ML, Lienhard R, Berner D, Anderau R, Treboux M, Ducommun JM, Malinverni R, Genne D, Erard PH, Waespi U. 2006. Outbreak of human listeriosis associated with tomme cheese in northwest Switzerland, 2005. Euro Surveill. 11:91–93 [PubMed] [Google Scholar]
- 13. Gaulin C, Ramsay D, Bekal S. 2012. Widespread listeriosis outbreak attributable to pasteurized cheese, which led to extensive cross-contamination affecting cheese retailers, Quebec, Canada, 2008. J. Food Prot. 75:71–78 [DOI] [PubMed] [Google Scholar]
- 14. Clark CG, Farber J, Pagotto F, Ciampa N, Dore K, Nadon C, Bernard K, Ng LK, CPHLN 2010. Surveillance for Listeria monocytogenes and listeriosis, 1995-2004. Epidemiol. Infect. 138:559–572 [DOI] [PubMed] [Google Scholar]
- 15. Knabel SJ, Reimer A, Verghese B, Lok M, Ziegler J, Farber J, Pagotto F, Graham M, Nadon CA, Canadian Public Health Laboratory Network, Gilmour MW. 2012. Sequence typing confirms that a predominant Listeria monocytogenes clone caused human listeriosis cases and outbreaks in Canada from 1988 to 2010. J. Clin. Microbiol. 50:1748–1751 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Gilmour MW, Graham M, Van Domselaar G, Tyler S, Kent H, Trout-Yakel KM, Larios O, Allen V, Lee B, Nadon C. 2010. High-throughput genome sequencing of two Listeria monocytogenes clinical isolates during a large foodborne outbreak. BMC Genomics 11:120 doi:10.1186/1471-2164-11-120 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Kovacevic J, Mesak LR, Allen KJ. 2012. Occurrence and characterization of Listeria spp. in ready-to-eat retail foods from Vancouver, British Columbia. Food Microbiol. 30:372–378 [DOI] [PubMed] [Google Scholar]
- 18. Nightingale KK, Ivy RA, Ho AJ, Fortes ED, Njaa BL, Peters RM, Wiedmann M. 2008. inlA premature stop codons are common among Listeria monocytogenes isolates from foods and yield virulence-attenuated strains that confer protection against fully virulent strains. Appl. Environ. Microbiol. 74:6570–6583 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Van Stelten A, Simpson JM, Ward TJ, Nightingale KK. 2010. Revelation by single-nucleotide polymorphism genotyping that mutations leading to a premature stop codon in inlA are common among Listeria monocytogenes isolates from ready-to-eat foods but not human listeriosis cases. Appl. Environ. Microbiol. 76:2783–2790 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Ward TJ, Evans P, Wiedmann M, Usgaard T, Roof SE, Stroika SG, Hise K. 2010. Molecular and phenotypic characterization of Listeria monocytogenes from U.S. Department of Agriculture Food Safety and Inspection Service surveillance of ready-to-eat foods and processing facilities. J. Food Prot. 73:861–869 [DOI] [PubMed] [Google Scholar]
- 21. Felicio MT, Hogg T, Gibbs P, Teixeira P, Wiedmann M. 2007. Recurrent and sporadic Listeria monocytogenes contamination in alheiras represents considerable diversity, including virulence-attenuated isolates. Appl. Environ. Microbiol. 73:3887–3895 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Handa-Miya S, Kimura B, Takahashi H, Sato M, Ishikawa T, Igarashi K, Fujii T. 2007. Nonsense-mutated inlA and prfA not widely distributed in Listeria monocytogenes isolates from ready-to-eat seafood products in Japan. Int. J. Food Microbiol. 117:312–318 [DOI] [PubMed] [Google Scholar]
- 23. Jonquieres R, Bierne H, Mengaud J, Cossart P. 1998. The inlA gene of Listeria monocytogenes LO28 harbors a nonsense mutation resulting in release of internalin. Infect. Immun. 66:3420–3422 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Nightingale K, Windham K, Martin K, Yeung M, Wiedmann M. 2005. Select Listeria monocytogenes subtypes commonly found in foods carry distinct nonsense mutations in inlA, leading to expression of truncated and secreted internalin A, and are associated with a reduced invasion phenotype for human intestinal epithelial cells. Appl. Environ. Microbiol. 71:8764–8772 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Rousseaux S, Olier M, Lemaitre JP, Piveteau P, Guzzo J. 2004. Use of PCR-restriction fragment length polymorphism of inlA for rapid screening of Listeria monocytogenes strains deficient in the ability to invade Caco-2 cells. Appl. Environ. Microbiol. 70:2180–2185 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Roldgaard BB, Andersen JB, Hansen TB, Christensen BB, Licht TR. 2009. Comparison of three Listeria monocytogenes strains in a guinea-pig model simulating food-borne exposure. FEMS Microbiol. Lett. 291:88–94 [DOI] [PubMed] [Google Scholar]
- 27. Van Stelten A, Simpson JM, Chen Y, Scott VN, Whiting RC, Ross WH, Nightingale KK. 2011. Significant shift in median guinea pig infectious dose shown by an outbreak-associated Listeria monocytogenes epidemic clone strain and a strain carrying a premature stop codon mutation in inlA. Appl. Environ. Microbiol. 77:2479–2487 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Orsi RH, den Bakker HC, Wiedmann M. 2011. Listeria monocytogenes lineages: genomics, evolution, ecology, and phenotypic characteristics. Int. J. Med. Microbiol. 301:79–96 [DOI] [PubMed] [Google Scholar]
- 29. Kovacevic J, McIntyre LF, Henderson SB, Kosatsky T. 2012. Occurrence and distribution of Listeria species in facilities producing ready-to-eat foods in British Columbia, Canada. J. Food Prot. 75:216–224 [DOI] [PubMed] [Google Scholar]
- 30. Seeliger H, Höhne K. 1979. Serotyping of Listeria monocytogenes and related species, p 31–50 In Bergan T, Norris JR. (ed), Methods in microbiology, vol 13 Academic Press, London, United Kingdom [Google Scholar]
- 31. Glaser P, Frangeul L, Buchrieser C, Rusniok C, Amend A, Baquero F, Berche P, Bloecker H, Brandt P, Chakraborty T, Charbit A, Chetouani F, Couve E, de Daruvar A, Dehoux P, Domann E, Dominguez-Bernal G, Duchaud E, Durant L, Dussurget O, Entian KD, Fsihi H, Garcia-del Portillo F, Garrido P, Gautier L, Goebel W, Gomez-Lopez N, Hain T, Hauf J, Jackson D, Jones LM, Kaerst U, Kreft J, Kuhn M, Kunst F, Kurapkat G, Madueno E, Maitournam A, Vicente JM, Ng E, Nedjari H, Nordsiek G, Novella S, de Pablos B, Perez-Diaz JC, Purcell R, Remmel B, Rose M, Schlueter T, Simoes N, Tierrez A, Vazquez-Boland JA, Voss H, Wehland J, Cossart P. 2001. Comparative genomics of Listeria species. Science 294:849–852 [DOI] [PubMed] [Google Scholar]
- 32. Gaillard JL, Finlay BB. 1996. Effect of cell polarization and differentiation on entry of Listeria monocytogenes into the enterocyte-like Caco-2 cell line. Infect. Immun. 64:1299–1308 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Gaillard JL, Berche P, Frehel C, Gouin E, Cossart P. 1991. Entry of L. monocytogenes into cells is mediated by internalin, a repeat protein reminiscent of surface antigens from Gram-positive cocci. Cell 65:1127–1141 [DOI] [PubMed] [Google Scholar]
- 34. Allen KJ, Poppe C. 2002. Phenotypic and genotypic characterization of food animal isolates of Salmonella with reduced sensitivity to ciprofloxacin. Microb. Drug Resist. 8:375–383 [DOI] [PubMed] [Google Scholar]
- 35. LeClerc JE, Li B, Payne WL, Cebula TA. 1996. High mutation frequencies among Escherichia coli and Salmonella pathogens. Science 274:1208–1211 [DOI] [PubMed] [Google Scholar]
- 36. Arguedas-Villa C, Stephan R, Tasara T. 2010. Evaluation of cold growth and related gene transcription responses associated with Listeria monocytogenes strains of different origins. Food Microbiol. 27:653–660 [DOI] [PubMed] [Google Scholar]
- 37. Baranyi J, Roberts TA. 1994. A dynamic approach to predicting bacterial growth in food. Int. J. Food Microbiol. 23:277–294 [DOI] [PubMed] [Google Scholar]
- 38. Wehrli W. 1983. Rifampin: mechanisms of action and resistance. Rev. Infect. Dis. 5(Suppl 3):S407–S411 [DOI] [PubMed] [Google Scholar]
- 39. Chen Y, Ross WH, Whiting RC, Van Stelten A, Nightingale KK, Wiedmann M, Scott VN. 2011. Variation in Listeria monocytogenes dose responses in relation to subtypes encoding a full-length or truncated internalin A. Appl. Environ. Microbiol. 77:1171–1180 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Jacquet C, Doumith M, Gordon JI, Martin PMV, Cossart P, Lecuit M. 2004. A molecular marker for evaluating the pathogenic potential of foodborne Listeria monocytogenes. J. Infect. Dis. 189:2094–2100 [DOI] [PubMed] [Google Scholar]
- 41. Ferreira V, Barbosa J, Stasiewicz M, Vongkamjan K, Moreno Switt A, Hogg T, Gibbs P, Teixeira P, Wiedmann M. 2011. Diverse geno- and phenotypes of persistent Listeria monocytogenes isolates from fermented meat sausage production facilities in Portugal. Appl. Environ. Microbiol. 77:2701–2715 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Bonazzi M, Lecuit M, Cossart P. 2009. Listeria monocytogenes internalin and E-cadherin: from bench to bedside. Cold Spring Harb. Perspect. Biol. 1:a003087 doi:10.1101/cshperspect.a003087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43. Orsi RH, Ripoll DR, Yeung M, Nightingale KK, Wiedmann M. 2007. Recombination and positive selection contribute to evolution of Listeria monocytogenes inlA. Microbiology 153:2666–2678 [DOI] [PubMed] [Google Scholar]
- 44. Ragon M, Wirth T, Hollandt F, Lavenir R, Lecuit M, Le Monnier A, Brisse S. 2008. A new perspective on Listeria monocytogenes evolution. PLoS Pathog. 4:e1000146 doi:10.1371/journal.ppat.1000146 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Dunn KA, Bielawski JP, Ward TJ, Urquhart C, Gu H. 2009. Reconciling ecological and genomic divergence among lineages of Listeria under an “extended mosaic genome concept.” Mol. Biol. Evol. 26:2605–2615 [DOI] [PubMed] [Google Scholar]
- 46. Orsi RH, Sun Q, Wiedmann M. 2008. Genome-wide analyses reveal lineage specific contributions of positive selection and recombination to the evolution of Listeria monocytogenes. BMC Evol. Biol. 8:233 doi:10.1186/1471-2148-8-233 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Lecuit M, Ohayon H, Braun L, Mengaud J, Cossart P. 1997. Internalin of Listeria monocytogenes with an intact leucine-rich repeat region is sufficient to promote internalization. Infect. Immun. 65:5309–5319 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Raffelsbauer D, Bubert A, Engelbrecht F, Scheinpflug J, Simm A, Hess J, Kaufmann SH, Goebel W. 1998. The gene cluster inlC2DE of Listeria monocytogenes contains additional new internalin genes and is important for virulence in mice. Mol. Gen. Genet. 260:144–158 [DOI] [PubMed] [Google Scholar]
- 49. Tsai YHL, Orsi RH, Nightingale KK, Wiedmann M. 2006. Listeria monocytogenes internalins are highly diverse and evolved by recombination and positive selection. Infect. Genet. Evol. 6:378–389 [DOI] [PubMed] [Google Scholar]
- 50. Buncic S, Avery SM, Rocourt J, Dimitrijevic M. 2001. Can food-related environmental factors induce different behaviour in two key serovars, 4b and 1/2a, of Listeria monocytogenes? Int. J. Food Microbiol. 65:201–212 [DOI] [PubMed] [Google Scholar]