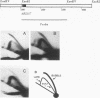
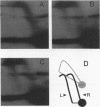
Abstract
ARS307 is highly active as a replication origin in its native location on chromosome III of Saccharomyces cerevisiae. Its ability to confer autonomous replication activity on plasmids requires the presence of an 11-bp autonomously replicating sequence (ARS) consensus sequence (ACS), which is also required for chromosomal origin function, as well as approximately 100 bp of sequence flanking the ACS called domain B. To further define the sequences required for ARS function, a linker substitution mutagenesis of domain B was carried out. The mutations defined two sequences, B1 and B2, that contribute to ARS activity. Therefore, like ARS1, domain B of ARS307 is composed of functional subdomains. Constructs carrying mutations in the B1 element were used to replace the chromosomal copy of ARS307. These mutations caused a reduction in chromosomal origin activity, demonstrating that the B1 element is required for efficient chromosomal origin function.
Full text
PDF


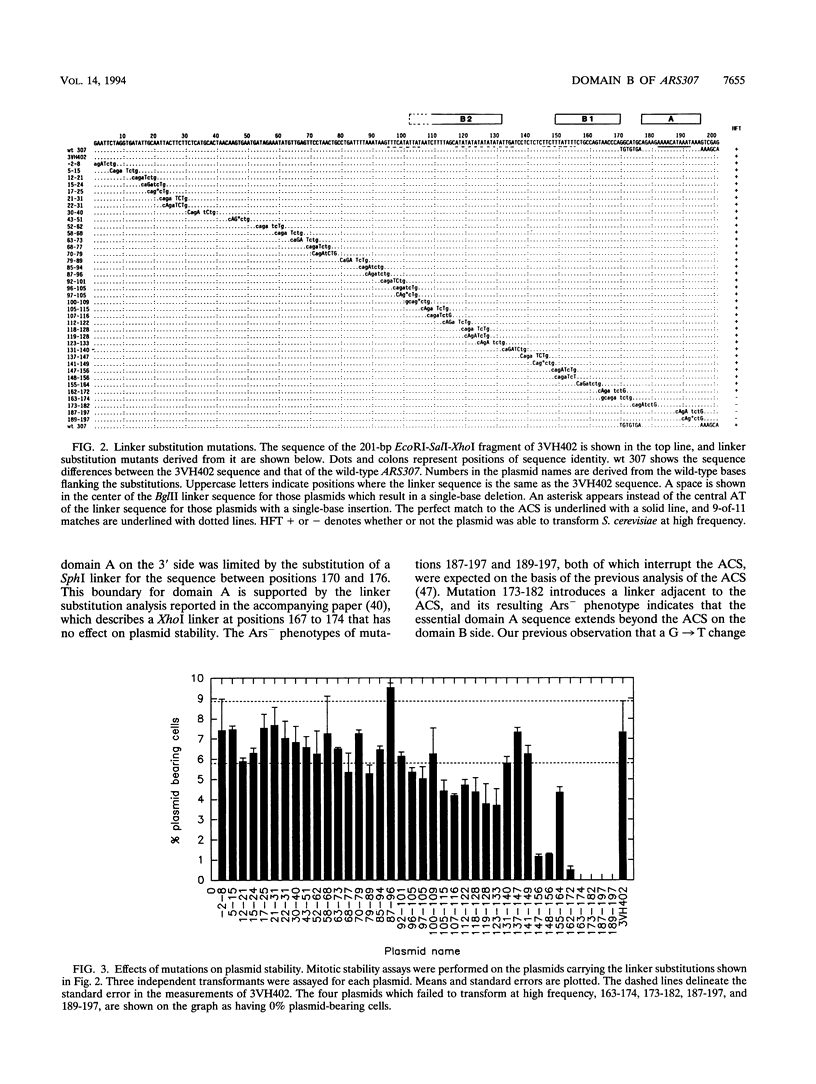




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Becker D. M., Guarente L. High-efficiency transformation of yeast by electroporation. Methods Enzymol. 1991;194:182–187. doi: 10.1016/0076-6879(91)94015-5. [DOI] [PubMed] [Google Scholar]
- Bell S. P., Kobayashi R., Stillman B. Yeast origin recognition complex functions in transcription silencing and DNA replication. Science. 1993 Dec 17;262(5141):1844–1849. doi: 10.1126/science.8266072. [DOI] [PubMed] [Google Scholar]
- Bell S. P., Stillman B. ATP-dependent recognition of eukaryotic origins of DNA replication by a multiprotein complex. Nature. 1992 May 14;357(6374):128–134. doi: 10.1038/357128a0. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., Trueheart J., Natsoulis G., Fink G. R. 5-Fluoroorotic acid as a selective agent in yeast molecular genetics. Methods Enzymol. 1987;154:164–175. doi: 10.1016/0076-6879(87)54076-9. [DOI] [PubMed] [Google Scholar]
- Bouton A. H., Smith M. M. Fine-structure analysis of the DNA sequence requirements for autonomous replication of Saccharomyces cerevisiae plasmids. Mol Cell Biol. 1986 Jul;6(7):2354–2363. doi: 10.1128/mcb.6.7.2354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brewer B. J., Chlebowicz-Sledziewska E., Fangman W. L. Cell cycle phases in the unequal mother/daughter cell cycles of Saccharomyces cerevisiae. Mol Cell Biol. 1984 Nov;4(11):2529–2531. doi: 10.1128/mcb.4.11.2529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brewer B. J., Fangman W. L. A replication fork barrier at the 3' end of yeast ribosomal RNA genes. Cell. 1988 Nov 18;55(4):637–643. doi: 10.1016/0092-8674(88)90222-x. [DOI] [PubMed] [Google Scholar]
- Brewer B. J., Fangman W. L. The localization of replication origins on ARS plasmids in S. cerevisiae. Cell. 1987 Nov 6;51(3):463–471. doi: 10.1016/0092-8674(87)90642-8. [DOI] [PubMed] [Google Scholar]
- Broach J. R., Li Y. Y., Feldman J., Jayaram M., Abraham J., Nasmyth K. A., Hicks J. B. Localization and sequence analysis of yeast origins of DNA replication. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1165–1173. doi: 10.1101/sqb.1983.047.01.132. [DOI] [PubMed] [Google Scholar]
- Deshpande A. M., Newlon C. S. The ARS consensus sequence is required for chromosomal origin function in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Oct;12(10):4305–4313. doi: 10.1128/mcb.12.10.4305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Desplan C., Theis J., O'Farrell P. H. The Drosophila developmental gene, engrailed, encodes a sequence-specific DNA binding activity. Nature. 1985 Dec 19;318(6047):630–635. doi: 10.1038/318630a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diffley J. F., Cocker J. H. Protein-DNA interactions at a yeast replication origin. Nature. 1992 May 14;357(6374):169–172. doi: 10.1038/357169a0. [DOI] [PubMed] [Google Scholar]
- Dubey D. D., Davis L. R., Greenfeder S. A., Ong L. Y., Zhu J. G., Broach J. R., Newlon C. S., Huberman J. A. Evidence suggesting that the ARS elements associated with silencers of the yeast mating-type locus HML do not function as chromosomal DNA replication origins. Mol Cell Biol. 1991 Oct;11(10):5346–5355. doi: 10.1128/mcb.11.10.5346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fangman W. L., Brewer B. J. Activation of replication origins within yeast chromosomes. Annu Rev Cell Biol. 1991;7:375–402. doi: 10.1146/annurev.cb.07.110191.002111. [DOI] [PubMed] [Google Scholar]
- Ferguson B. M., Brewer B. J., Reynolds A. E., Fangman W. L. A yeast origin of replication is activated late in S phase. Cell. 1991 May 3;65(3):507–515. doi: 10.1016/0092-8674(91)90468-e. [DOI] [PubMed] [Google Scholar]
- Foss M., McNally F. J., Laurenson P., Rine J. Origin recognition complex (ORC) in transcriptional silencing and DNA replication in S. cerevisiae. Science. 1993 Dec 17;262(5141):1838–1844. doi: 10.1126/science.8266071. [DOI] [PubMed] [Google Scholar]
- Greenfeder S. A., Newlon C. S. A replication map of a 61-kb circular derivative of Saccharomyces cerevisiae chromosome III. Mol Biol Cell. 1992 Sep;3(9):999–1013. doi: 10.1091/mbc.3.9.999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III in DNA sequence analysis. Methods Enzymol. 1987;155:156–165. doi: 10.1016/0076-6879(87)55014-5. [DOI] [PubMed] [Google Scholar]
- Hieter P., Mann C., Snyder M., Davis R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell. 1985 Feb;40(2):381–392. doi: 10.1016/0092-8674(85)90152-7. [DOI] [PubMed] [Google Scholar]
- Holmes S. G., Smith M. M. Interaction of the H4 autonomously replicating sequence core consensus sequence and its 3'-flanking domain. Mol Cell Biol. 1989 Dec;9(12):5464–5472. doi: 10.1128/mcb.9.12.5464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang R. Y., Kowalski D. A DNA unwinding element and an ARS consensus comprise a replication origin within a yeast chromosome. EMBO J. 1993 Dec;12(12):4521–4531. doi: 10.1002/j.1460-2075.1993.tb06141.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huberman J. A., Spotila L. D., Nawotka K. A., el-Assouli S. M., Davis L. R. The in vivo replication origin of the yeast 2 microns plasmid. Cell. 1987 Nov 6;51(3):473–481. doi: 10.1016/0092-8674(87)90643-x. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kearsey S. Structural requirements for the function of a yeast chromosomal replicator. Cell. 1984 May;37(1):299–307. doi: 10.1016/0092-8674(84)90326-x. [DOI] [PubMed] [Google Scholar]
- Kowalski D., Eddy M. J. The DNA unwinding element: a novel, cis-acting component that facilitates opening of the Escherichia coli replication origin. EMBO J. 1989 Dec 20;8(13):4335–4344. doi: 10.1002/j.1460-2075.1989.tb08620.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li J. J., Herskowitz I. Isolation of ORC6, a component of the yeast origin recognition complex by a one-hybrid system. Science. 1993 Dec 17;262(5141):1870–1874. doi: 10.1126/science.8266075. [DOI] [PubMed] [Google Scholar]
- Lin S., Kowalski D. DNA helical instability facilitates initiation at the SV40 replication origin. J Mol Biol. 1994 Jan 14;235(2):496–507. doi: 10.1006/jmbi.1994.1009. [DOI] [PubMed] [Google Scholar]
- Linskens M. H., Huberman J. A. Organization of replication of ribosomal DNA in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Nov;8(11):4927–4935. doi: 10.1128/mcb.8.11.4927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marahrens Y., Stillman B. A yeast chromosomal origin of DNA replication defined by multiple functional elements. Science. 1992 Feb 14;255(5046):817–823. doi: 10.1126/science.1536007. [DOI] [PubMed] [Google Scholar]
- Matsumoto K., Ishimi Y. Single-stranded-DNA-binding protein-dependent DNA unwinding of the yeast ARS1 region. Mol Cell Biol. 1994 Jul;14(7):4624–4632. doi: 10.1128/mcb.14.7.4624. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Micklem G., Rowley A., Harwood J., Nasmyth K., Diffley J. F. Yeast origin recognition complex is involved in DNA replication and transcriptional silencing. Nature. 1993 Nov 4;366(6450):87–89. doi: 10.1038/366087a0. [DOI] [PubMed] [Google Scholar]
- Miller C. A., Kowalski D. cis-acting components in the replication origin from ribosomal DNA of Saccharomyces cerevisiae. Mol Cell Biol. 1993 Sep;13(9):5360–5369. doi: 10.1128/mcb.13.9.5360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Natale D. A., Schubert A. E., Kowalski D. DNA helical stability accounts for mutational defects in a yeast replication origin. Proc Natl Acad Sci U S A. 1992 Apr 1;89(7):2654–2658. doi: 10.1073/pnas.89.7.2654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Natale D. A., Umek R. M., Kowalski D. Ease of DNA unwinding is a conserved property of yeast replication origins. Nucleic Acids Res. 1993 Feb 11;21(3):555–560. doi: 10.1093/nar/21.3.555. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newlon C. S., Collins I., Dershowitz A., Deshpande A. M., Greenfeder S. A., Ong L. Y., Theis J. F. Analysis of replication origin function on chromosome III of Saccharomyces cerevisiae. Cold Spring Harb Symp Quant Biol. 1993;58:415–423. doi: 10.1101/sqb.1993.058.01.048. [DOI] [PubMed] [Google Scholar]
- Newlon C. S., Theis J. F. The structure and function of yeast ARS elements. Curr Opin Genet Dev. 1993 Oct;3(5):752–758. doi: 10.1016/s0959-437x(05)80094-2. [DOI] [PubMed] [Google Scholar]
- Palzkill T. G., Newlon C. S. A yeast replication origin consists of multiple copies of a small conserved sequence. Cell. 1988 May 6;53(3):441–450. doi: 10.1016/0092-8674(88)90164-x. [DOI] [PubMed] [Google Scholar]
- Rao H., Marahrens Y., Stillman B. Functional conservation of multiple elements in yeast chromosomal replicators. Mol Cell Biol. 1994 Nov;14(11):7643–7651. doi: 10.1128/mcb.14.11.7643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shirahige K., Iwasaki T., Rashid M. B., Ogasawara N., Yoshikawa H. Location and characterization of autonomously replicating sequences from chromosome VI of Saccharomyces cerevisiae. Mol Cell Biol. 1993 Aug;13(8):5043–5056. doi: 10.1128/mcb.13.8.5043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Theis J. F., Newlon C. S. The DNA sequence of a 762 bp fragment containing the SUP11-1 gene. Yeast. 1992 Mar;8(3):223–225. doi: 10.1002/yea.320080308. [DOI] [PubMed] [Google Scholar]
- Umek R. M., Kowalski D. The ease of DNA unwinding as a determinant of initiation at yeast replication origins. Cell. 1988 Feb 26;52(4):559–567. doi: 10.1016/0092-8674(88)90469-2. [DOI] [PubMed] [Google Scholar]
- Van Houten J. V., Newlon C. S. Mutational analysis of the consensus sequence of a replication origin from yeast chromosome III. Mol Cell Biol. 1990 Aug;10(8):3917–3925. doi: 10.1128/mcb.10.8.3917. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker S. S., Francesconi S. C., Eisenberg S. A DNA replication enhancer in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4665–4669. doi: 10.1073/pnas.87.12.4665. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker S. S., Malik A. K., Eisenberg S. Analysis of the interactions of functional domains of a nuclear origin of replication from Saccharomyces cerevisiae. Nucleic Acids Res. 1991 Nov 25;19(22):6255–6262. doi: 10.1093/nar/19.22.6255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams D. L., Kowalski D. Easily unwound DNA sequences and hairpin structures in the Epstein-Barr virus origin of plasmid replication. J Virol. 1993 May;67(5):2707–2715. doi: 10.1128/jvi.67.5.2707-2715.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]