Abstract
Cells face a constant challenge as they produce new proteins. The newly synthesized polypeptides must be folded properly to avoid aggregation. If proteins do misfold, they must be cleared to ensure a functional and healthy proteome. Recent work is revealing the complex mechanisms that work cotranslationally to ensure protein quality control during biogenesis at the ribosome. Indeed, the ribosome is emerging as a central hub in coordinating these processes, particularly in sensing the nature of the nascent protein chain, recruiting protein folding and translocation components, and integrating mRNA and nascent chain quality control. The tiered and complementary nature of these decision-making processes confers robustness and fidelity to protein homeostasis during protein synthesis.
Introduction
Accurate protein folding is a pre-requisite for all cellular processes, and protein homeostasis disfunction is linked to an expanding list of human diseases (Powers et al., 2009). This is challenging for cells because proteins are continuously made by the approximately 106–107 ribosomes in a eukaryotic cell. Accurately folding newly made polypeptides is a complex task, which must be achieved in the face of the vectorial nature of translation, whereby the N-terminal end is available for folding before the other end has been synthesized, as well as the crowded cellular environment awaiting the unfolded emerging chains (Gershenson and Gierasch, 2011; Hartl et al., 2011). The cell must not only promote accurate folding but also must prevent the accumulation of misfolded species that may arise from inefficient folding, errors in translation, and aberrant mRNAs. Misfolded proteins can overwhelm the quality control machinery, leading to the accumulation of aggregated species that are often cytotoxic and linked to severe misfolding diseases that include Alzheimer’s and Parkinson’s (Chiti and Dobson, 2006).
To ensure proper folding and quality control of the nascent proteome, the cell depends on the concerted action of sophisticated pathways to both guide translated proteins into their functional conformation and prevent the accumulation of defective polypeptides. Molecular chaperones are central elements of these quality control systems, as they facilitate protein biogenesis by assisting polypeptide folding, translocation, and assembly of newly made proteins in the crowded cellular (Gershenson and Gierasch, 2011; Hartl et al., 2011). The cellular quality control machinery also responds to errors during protein synthesis by either buffering destabilizing mutations (Fares et al., 2002; Jarosz et al., 2010), or targeting polypeptides for degradation (Kaganovich et al., 2008; Tyedmers et al., 2010).
Interestingly, the ribosome itself is emerging as a central actor in protein folding and quality control processes, acting as a proofreader during protein biogenesis and initiating downstream events that regulate the fate of the nascent chains. Here, we review recent insights into the quality control of newly made proteins, and sketch an emerging integrated picture of tightly co-evolved, dynamic, and hierarchical safety mechanisms that guarantee high fidelity in nascent chain protein biogenesis.
Biophysical constraints on nascent chain folding
Emerging from the ribosome, nascent chains have to avoid off-pathway misfolding and aggregation (Jahn and Radford, 2008). Because proteins vary substantially in their propensities to fold and aggregate, the intrinsic properties of the protein sequences will dramatically influence their ability to fold upon translation. Studies of the in vitro folding kinetics of small proteins have established that the folding rate increases with the number of native contacts in the folded structure (Eaton et al., 2000), as well as with the number of potential non-native contacts (Bowman et al., 2011). These analyses implicate sequence length as the strongest determinant of folding kinetics for small single domain proteins (Ouyang and Liang, 2008). With this in mind, the vectorial nature of polypeptide synthesis at the ribosome can both assist and complicate successful folding. The number of possible structural conformations is initially constrained, thus reducing the chance of misfolding of N-terminal domains through interaction with C-terminal sequences that have not yet emerged from the ribosome (Cabrita et al., 2010; Komar, 2009; Zhang and Ignatova, 2011). On the other hand, folding of long multi-domain proteins may require the formation of contacts distant in polypeptide sequence, which must leave the ribosome before completing folding (O'Brien et al., 2012; Zhang and Ignatova, 2011). Early experiments demonstrated that N-terminal domains can fold cotranslationally, thus enhancing the rate and yield of folding in vivo compared to the in vitro folding kinetics (Frydman et al., 1999). The constraints of cotranslational folding likely influence protein evolution; individual protein domains are often maintained as independent folding units (Han et al., 2007), and the overall distribution of their lengths in proteomes reflects a length limit for folding on a biologically relevant time scale (Lin and Zewail, 2012).
The amino acid sequences of proteins also determine their intrinsic propensity to aggregate (Chiti and Dobson, 2006; Goldschmidt et al., 2010; Sawaya et al., 2007; Tartaglia et al., 2007). Non-specific interactions between aggregation-prone and promiscuous proteins can be deleterious (Vavouri et al., 2009), and aggregation is often cytotoxic (Chiti and Dobson, 2006). However, aggregation-prone sequences are also clearly important for protein structure and function, and are therefore ubiquitously present in the proteome (Fowler et al., 2007; Goldschmidt et al., 2010). For instance, all cellular signaling pathways involve interactions between proteins (Levy and Pereira-Leal, 2008), and central hubs in protein networks often rely on large proteins with complex folds, exposed hydrophobic, aggregation-prone surfaces, and long unstructured regions (Tsai et al., 2009). As discussed below, chaperones are an important cellular response to the inevitable presence of aggregation-prone sequences in the proteome.
Cellular strategies to optimize de novo folding
How are the biophysical constraints on successful cotranslational folding balanced with the cellular need for long, more complex, and slowly folding proteins? It appears that the challenge is met through a two-fold strategy; first, through the evolutionary optimization of protein sequences to prevent aggregation and maximize solubility, and second, through the action of a ribosome-anchored chaperone machinery.
Several studies suggest that protein sequences have evolved to minimize the folding burden of the cell. Most proteins are short, and longer aggregation-prone proteins tend to be on average less abundant (Tartaglia et al., 2007). In turn, highly expressed proteins have a lower propensity to aggregate (Tartaglia et al., 2007), and are generally more conserved (Drummond et al., 2005); these features help to maintain solubility and reduce the risk of erroneous mutations that can promote aggregation (Gsponer and Babu, 2012; Pechmann and Vendruscolo, 2010). Aggregation prone regions are often flanked by charged amino acids or proline residues that act as “gatekeeper residues” also to maintain solubility and minimize aggregation (Buell et al., 2009). Since aggregation can be strongly promoted by specific interaction of complementary sequence motifs (Sawaya et al., 2007), multidomain proteins with homologous domains generally maintain low sequence identities to prevent co-aggregation of their domains (Wright et al., 2005)
As discussed below, recent studies of cotranslational chaperone function suggest a direct relationship between biophysical constraints on cellular folding and the need for energetically costly quality control mechanisms (Calloni et al., 2012; Willmund et al., 2013). Thus, maintaining protein homeostasis relies on a tiered risk management strategy. The organization and evolutionary dynamics of the proteome provide a first line of defense in protein quality control, where sequences have evolved to avoid aggregation, and most proteins tend to be short and thus fold efficiently. This allows the energy dependent chaperone systems to preferentially protect long, aggregation-prone but functionally important proteins.
Role of the mRNA in translational fidelity and cotranslational folding
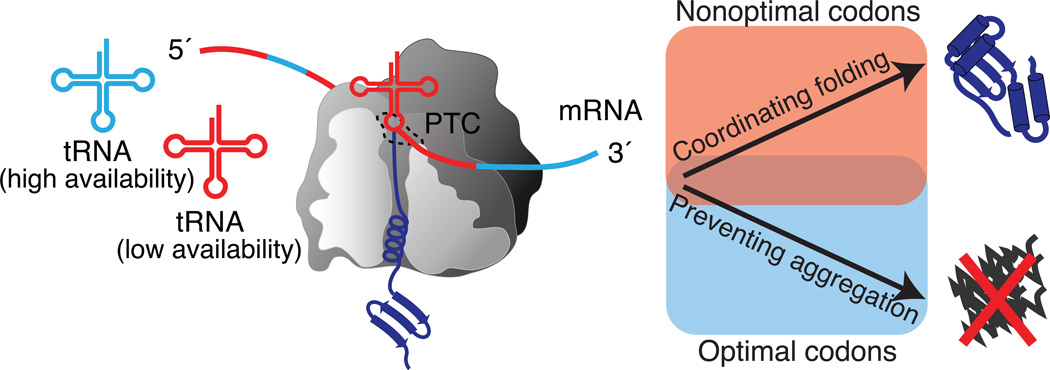
Natively folded proteins are only marginally stable in the cell (Baldwin et al., 2011; DePristo et al., 2005). The canonical error rate during protein synthesis results in approximately one per 104 mis-incorporated amino acids (Zaher and Green, 2009a) that could tip the balance away from correct folding. Recent work indicates that the mRNAs sequences, and not just the encoded amino acid sequences, also influence the cellular synthesis and folding of their corresponding proteins (Figure 1). Almost all amino acids can be encoded by several synonymous codons that are under selection for translational speed (dos Reis et al., 2004) and accuracy (Akashi, 1994). Translationally optimal codons are recognized by tRNAs with higher availability, and thus are translated more rapidly; they also confer higher translational accuracy because their corresponding tRNAs deliver the encoded amino acid more efficiently to the peptidyltransferase center (PTC) (Figure 1). Optimal codons have been found to associate with evolutionarily conserved, thus likely functionally important sites in the corresponding proteins (Drummond and Wilke, 2008). Moreover, optimal codons could be found preferentially at structurally sensitive and aggregation prone sites in proteins (Lee et al., 2010), i.e. positions that are particularly susceptible to translation errors disrupting folding and function. This suggests that selection for optimal codons to enhance translational fidelity is an important determinant of coding sequence evolution (Figure 1).
Figure 1. Translational fidelity and coordinated cotranslational folding at the ribosome.
The ribosome checks many aspects of protein synthesis: proofreading of tRNA binding and peptide bond formation, and sensing conformations inside the exit tunnel. Optimal codons recognized by highly available tRNAs are translated fast and are under selection for higher translational accuracy. They are found preferentially at sites where high fidelity is important to prevent aggregation. Non-optimal codons are recognized by less abundant tRNAs, thus slow down translation. Clusters of nonoptimal codons are evolutionarily conserved and enriched in secondary-structure elements that can fold cotranslationally, suggesting a general role in coordinating cotranslational folding. [Adapted from (Pechmann and Frydman, 2012)].
Non-optimal codons are recognized by lower availability tRNAs and thus slow down translation elongation; their positioning along the mRNA coding sequence could in principle allow the emerging nascent chain more time to begin to fold cotranslationally (Figure 1) (O'Brien et al., 2012; Zhang and Ignatova, 2011). Because proteins start folding as they emerge from the ribosome, native contacts in secondary or tertiary structures between distant sites in the polypeptide chain, such as in beta-sheets or between protein domains, can only form once they have completely left the ribosome exit tunnel (Cabrita et al., 2010; Fedyukina and Cavagnero, 2011). Indeed, clusters of non-optimal codons are maintained under direct selection (Clarke and Clark, 2008), and their substitution has been found to affect protein stability (Zhang et al., 2010) and function (Kimchi-Sarfaty et al., 2007). The role of optimal and non-optimal codons in cotranslational protein folding is supported by recent findings that clusters of optimal and non-optimal codons are evolutionarily conserved across eukaryotic organisms in a site-specific manner (Pechmann and Frydman, 2012). Of note, both optimal and non-optimal codons are conserved in equal measure, highlighting the importance of maintaining non-optimal codons for correct protein biogenesis (Pechmann and Frydman, 2012). The evolutionarily conserved patterns of optimal and non-optimal codons directly correlate with the secondary structure of the encoded polypeptides. Structural elements that can fold cotranslationally, such as alpha helices, contain conserved clusters of non-optimal codons, which likely slow down elongation to enhance cotranslational folding. These findings indicate that the rhythm of elongation is directly attuned to the needs of cotranslational folding (Figure 1). An even stronger attenuation of translation is achieved by specific stalling sequences recognized inside the ribosome (Tanner et al., 2009). In bacteria, ribosome profiling found that the anti-Shine-Delgarno sequence, known for its role in regulating translation initiation, is strategically placed throughout coding sequences to shape the translation of bacterial genomes (Li et al., 2012). In depth analysis of conserved mRNA sequence signatures together with powerful ribosome profiling experiments are primed to further decipher the layer of information contained within the coding mRNA sequences that coordinates protein synthesis and cotranslational folding.
The ribosome as a platform for folding and quality control
The ribosome is emerging as an active and dynamic hub in protein and mRNA quality control (Kramer et al., 2009; Wilson and Beckmann, 2011). Low stability of mRNA secondary structures at the very beginning of coding sequences facilitate the threading of mRNAs during translation initiation (Gu et al., 2010), but, if needed, the ribosome itself can act as helicase to unravel mRNA structures (Takyar et al., 2005). During translation elongation, the binding energy of cognate tRNAs induce subtle conformational changes in the ribosome itself that facilitate tRNA recognition and proofreading during translation elongation (Johansson et al., 2012; Ogle and Ramakrishnan, 2005). An additional quality control mechanism is implemented after peptidyl transfer and during peptide bond formation to avoid the incorporation of an amino acid from a non-cognate tRNA through termination of nascent chain synthesis (Zaher and Green, 2009b).
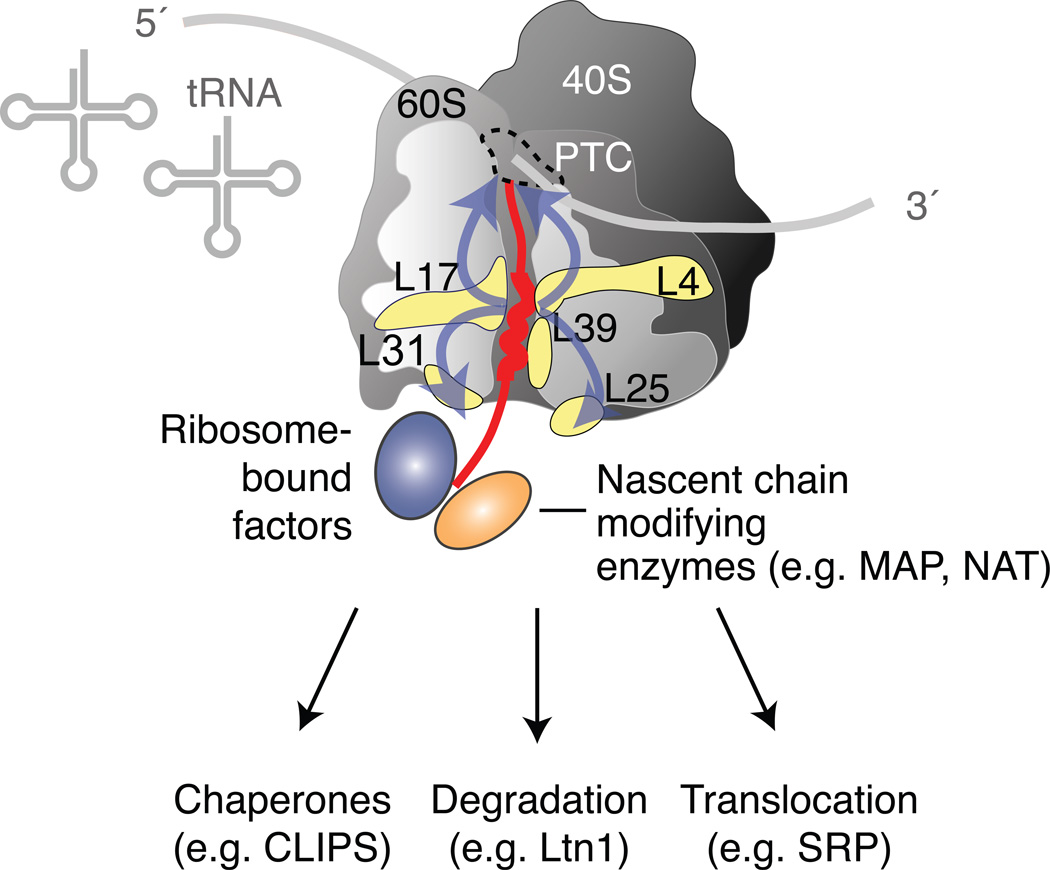
All ribosomes have an exit tunnel spanning the distance from the peptidyl transferase site (PTC) at the subunit interface to the exit site at the other end of the large subunit [Figure2, for reviews see (Melnikov et al., 2012; Klinge et al., 2012; Schmeing and Ramakrishnan, 2009)]. The nature of the tunnel, and the structural conformations of proteins and RNAs surrounding the exit site differ between eukaryotic and prokaryotic ribosomes (Melnikov et al., 2012; Klinge et al., 2012; Schmeing and Ramakrishnan, 2009). Nascent chains start to fold at the ribosomal exit tunnel port (Bhushan et al., 2010a; O'Brien et al., 2010), and they can even form helices while still deep inside the ribosome (Lu and Deutsch, 2005; Woolhead et al., 2004). There is a constriction site in the exit tunnel, approximately 10 amino acids from the PTC. In S. cerevisiae, this site consists of the ribosomal proteins L4, L17, and L39 (Figure 2, in yellow). The constriction site can sense nascent chain conformations and has been implicated in communication pathways from the inside to the outside of the ribosome (Wilson and Beckmann, 2011). The ability of the ribosomal tunnel to sense polypeptides during synthesis was initially identified in bacteria (Gong and Yanofsky, 2002; Nakatogawa and Ito, 2002). Specific sequence motifs, present for instance in SecM and TnaC, caused the translating polypeptide to stall at this constriction point in the tunnel in response to SecA or Tryptophan levels respectively (Cruz-Vera et al., 2006; Nakatogawa and Ito, 2001). Similar stalling sequences have been identified in eukaryotes (Bhushan et al., 2010b; Spevak et al., 2010). However, the constriction site within the ribosomal tunnel is likely attentive to even more subtle cues. Nascent helix interactions with the tunnel have been shown to induce structural rearrangements in the ribosome itself, triggering downstream translocation factors that control subsequent trafficking (Figure 2) (Bornemann et al., 2008; Lin et al., 2011). An evolutionarily conserved region of low translational efficiency at the very beginning of mRNA coding sequences helps to prevent ribosome traffic jams during translation elongation (Tuller et al., 2010). Interestingly, this region codes for a protein sequence long enough to span the distance from the PTC to the constriction site (Pechmann and Frydman, 2012), perhaps regulating how quickly and in which conformation the new polypeptide reaches the constriction site. An important and exciting question for future research is the extent to which the ribosome tunnel communicates with factors at the ribosome exit site, which include nascent chain modifying enzymes such as Methionyl Aminopeptidases and N-acetyl transferases, molecular chaperones, translocation factors and quality control components (Kramer et al., 2009), to influence nascent chain folding and cellular fate (Figure 2).
Figure 2. The ribosome as control center of nascent chain fate.
The ribosome exit tunnel interacts with new polypeptides. A constriction site, in S. cerevisiae comprised of ribosome proteins L4, L17, and L39, recognizes nascent chain conformations, and communicates with both the peptidyl transferase site and with the exit site (blue arrows). Ribosome associated chaperones and nascent chain modifying enzymes bind the ribosome near the tunnel exit. The nascent chain fate can be predetermined from inside the ribosome. Chaperones promote structural maturation, E3 ligases facilitate degradation, and the SRP is recruited to the ribosome before the polypeptide has exited.
The network of ribosome-bound chaperones
Early studies indicated that chaperones cotranslationally bind to nascent chains as they emerge from the ribosome (Frydman et al., 1994). The extent of cotranslational chaperone action is very different in eukaryotic and prokaryotic cells (Figure 3) (Albanèse et al., 2006). The bacterial Trigger factor (TF) is a single ATP-independent ribosome-bound chaperone, associating with many nascent chains (Oh et al., 2011; Preissler and Deuerling, 2012). Subsequent de novo folding is mostly post-translational (Agashe et al., 2004), and supported by the same ATP-dependent chaperone systems that protect the proteome from stress. These include the Hsp70 system composed of DnaJ, DnaK and GrpE, and the chaperonin system composed of GroEL-GroES (Figure 3) (Agashe et al., 2004; Hartl et al., 2011). In eukaryotes, a significant amount of folding occurs cotranslationally (Duncan and Mata, 2011; Frydman et al., 1999; Khushoo et al., 2011). The eukaryotic chaperone machinery comprises two independently regulated networks with distinct functions (Albanèse et al., 2006). A set of Chaperones Linked to Protein Synthesis (CLIPS) associates specifically with ribosomes to facilitate de novo folding and is transcriptionally co-regulated with the translation apparatus. In contrast, Heat Shock chaperones are induced under stress conditions to protect the proteome and mediate either refolding or quality control (Albanèse et al., 2006). CLIPS include structurally and mechanistically diverse chaperones, such as ATP-independent nascent polypeptide-associated complex (NAC), and GimC/Prefoldin; as well as ATP-dependent chaperones, including Hsp70 family members such as SSB in yeast, and the eukaryotic chaperonin TRiC/CCT (Figure 3) (Albanèse et al., 2006). Why eukaryotes require such a complex folding machinery, and the specificity and interplay between these different chaperones at the ribosome are questions that remain to be addressed.
Figure 3. Co- and post-translationally acting chaperone networks.
Prokaryotic and eukaryotic chaperone networks differ in their complexity and division of labor. In prokayotes, Trigger factor is the main ribosome-associated chaperone, folding new proteins orpassing them to the Hsp70 system of DnaK/DnaJ or the chaperonin GroEL/ES to facilitate de novo and stress-induced folding. In eukaryotes, ribosome-bound chaperones and co-chaperones like SSB, RAC, SRP, and NAC compete for overlapping ribosomal binding sites. More diverse and specialized networks of downstream chaperones split the task of mediating de novo and stress-induced folding Prefoldin and TRiC are directly coupled to protein synthesis, while Hsp90 is the primary heat shock chaperone.
The best-studied ribosome-bound chaperone is the bacterial TF, which provides a paradigm for how chaperones may influence nascent chains as they exit the ribosome. TF binds ribosomes close to the tunnel exit site (Ferbitz et al., 2004; Lakshmipathy et al., 2007). Global profiling of TF-bound ribosomes indicated that TF interacts with cytoplasmic and outer membrane-bound nascent polypeptides (Oh et al., 2011). Despite its broad specificity, biophysical studies showed that the residence time of TF on translating ribosomes correlates with sequence hydrophobicity of the emerging nascent chains (Kaiser et al., 2006). Nascent polypeptides are accommodated in the interior cleft of TF where multiple hydrophobic and hydrophilic binding sites support nascent chain interactions (Martinez-Hackert and Hendrickson, 2009). Biochemical experiments indicate that TF binding to nascent chains prevents their aggregation, protects them from proteolytic cleavage, delays premature folding of translated nascent polypeptide and reduces the formation of early misfolding intermediates of nascent polypeptide (Agashe et al., 2004; Kaiser et al., 2006, Hoffmann, 2012 #11).
The chaperones in the more complex eukaryotic CLIPS network show neither sequence nor structural homology to prokaryotic TF (Kramer et al., 2009; Preissler and Deuerling, 2012). Their increased complexity likely responds to the larger and more complex nature of eukaryotic proteomes (Albanèse et al., 2006). Recent work has begun to illuminate mechanistic and functional aspects of two early acting CLIPS, the nascent polypeptide-associated complex NAC and the Hsp70 SSB. These two chaperones appear to have overlapping but distinct functions, as highlighted by their strong negative genetic interactions (Koplin et al., 2010).
The ribosome–associated nascent polypeptide-associated complex (NAC) is a highly conserved dimeric complex that can be crosslinked to very short nascent chains (Raue et al., 2007). Yeast cells contain alpha and beta isoforms that can form hetero- and homo-oligomers (Beatrix et al., 2000). Ribosome binding appears primarily mediated through the N-terminus of beta NAC to a site proximal to ribosomal protein Rpl31 close to the exit tunnel (Pech et al., 2010; Zhang et al., 2012). While NAC is dispensable for viability in yeast, it is essential in metazoans (Markesich et al., 2000). The exact role of NAC is still under debate, but recent studies showed it is a cotranslational binding factor for most nascent chains (Alamo et al., 2011). Interestingly, while NAC interacts very broadly with translating ribosomes, including those translating precursors of secretory and mitochondrial proteins, the different isoforms of NAC interact with different subsets of nascent chains, suggesting they may differ in their binding specificity and function (Alamo et al., 2011). These experiments also began to reveal the complex interplay between different nascent chain binding factors at the ribosome. Thus, deletion of NAC causes a relaxation in the specificity of another cotranslationally acting factor, the signal recognition particle (SRP), which recognizes nascent chains carrying a signal sequence for translocation to the Endoplasmic Reticulum (ER) (Alamo et al., 2011). Since NAC and SRP can both bind to ribosomes carrying ER-bound nascent chains, the relaxation in specificity likely arises from the interplay of these two factors at the ribosome. Consistent with this idea, biochemical studies indicated that the presence of a signal sequence in the nascent chain is sensed from within the ribosomal exit tunnel to preemptively recruit SRP to those ribosomes (Berndt et al., 2009) in a manner modulated by NAC (Zhang et al., 2012). Future studies should further determine how NAC modulates the fate of cytoplasmic proteins, as well as mitochondrial precursors (George et al., 1998).
Hsp70s were observed to bind cotranslationally to nascent chains over 20 years ago (Beckmann et al., 1990; Frydman et al., 1994) but the function of this interaction is only now beginning to emerge. The cotranslational Hsp70 cycle is best characterized for the S. cerevisiae Hsp70 SSB, comprising the functionally redundant Ssb1 and Ssb2 isoforms. SSB associates with ribosomes and directly binds a large fraction of newly translated polypeptides (Willmund et al., 2013). ATP hydrolysis by SSB is stimulated by the Ribosome-Associated Complex RAC, which is composed by the atypical Hsp70 Ssz1 and the J-domain protein Zuo1 (Huang et al., 2005). RAC binds directly to ribosomes but does not interact with nascent chains itself (Yam et al., 2005). Instead, RAC regulates the cotranslational substrate binding and specificity of SSB (Willmund et al., 2013). A recent characterization of the interaction of RAC with ribosomes showed that RAC contacts the ribosome close to Rpl31, bending in a crouched conformation over the exit tunnel (Leidig et al., 2013). Structural analysis suggests that the atypical Hsp70 component of RAC, Ssz1, is inactive due to a permanently locked conformation, which prevents cycling through ATP and thus substrate binding and release (Leidig et al., 2013). The other cofactor completing the regulation of the ATPase cycle of SSB is the nucleotide exchange factor Sse1/Hsp110, a large Hsp70-like chaperone, which may itself bind and fold substrates (Raviol et al., 2006; Shaner et al., 2005; Yam et al., 2005). Interestingly, Sse1, which promotes substrate release from SSB (Yam et al., 2005) associates with SSB off the ribosome (Willmund et al., 2013), suggesting that the cycle of SSB substrate binding and ATP hydrolysis is spatially regulated by the differential interaction between its cofactors and the ribosome. Importantly, RAC and Sse1 are conserved in mammalian cells, indicating their role is conserved across eukaryotes (Jaiswal et al., 2011).
A global analysis of the function and specificity of SSB illuminates the role of cotranslational Hsp70 in eukaryotic protein biogenesis (Willmund et al., 2013). SSB binds preferentially to a subset nascent chains encoding nuclear and cytosolic proteins. Importantly, SSB, unlike NAC, does not bind SRP substrates, which indicates an early sorting mechanism at the ribosome, (Willmund et al., 2013). Nascent polypeptides that associate with SSB tend to be longer that those that do not bind Hsp70; furthermore, they have properties that pose a significant challenge to cotranslational folding, such as enhanced aggregation propensity, regions of intrinsic disorder and complex domain architectures (Willmund et al., 2013). Interestingly, deletion of SSB, while viable in yeast, causes widespread aggregation of newly made proteins with exactly those folding-challenging characteristics. Thus, Hsp70 plays a critical role in preventing cotranslational misfolding and downstream aggregation.
These recent studies reveal an intricate network of early acting ribosome-bound chaperones choreographing the early sorting of nascent chains (Figure 3). The generalist NAC both protects newly made cytosolic proteins, and promotes the specificity of SRP recognition of ER-bound nascent polypeptides. Hsp70 in turn appears to more specifically associate with folding-challenged cytoplasmic and nuclear proteins to prevent their aggregation and promote folding. Interestingly, the specificity of Hsp70 is modulated by RAC, whose binding site on the ribosome is close to that of NAC, thus raising the question regarding the competition for ribosomal binding sites of co-chaperones. Taken together, the highly dynamic and partially redundant nature of this network of ribosome-bound chaperones increases both the plasticity as well as robustness of the quality control of polypeptides emerging from the ribosome.
Co- and post-translational folding pathways
Many cellular proteins require additional chaperone assistance to fold, beyond the action offered by early acting ribosome-bound chaperones. In prokaryotes, the Hsp70/DnaK system and the GroEL/GroES chaperonin cooperate to promote folding. They interact with substrates primarily post-translationally, although DnaK can also associate cotranslationally (Deuerling et al., 1999; Teter et al., 1999). DnaK and GroEL have distinct, but overlapping substrates (Houry et al., 1999), as do DnaK and TF (Deuerling et al., 1999; Teter et al., 1999). GroEL is essential in most prokaryotes and is an obligate chaperone for the folding of topologically complex and aggregation prone proteins (Kerner et al., 2005; Tartaglia et al., 2010). The functional overlap between TF and DnaK confers robustness to the network, and deletions of either component are viable (Deuerling et al., 1999) (Teter et al., 1999). However, loss of both TF and DnaK severely affects growth and disrupts access of newly translated proteins to GroEL, thus resulting in wide-spread aggregation (Calloni et al., 2012).
In eukaryotes, several chaperones act co- and post-translationally to facilitate protein folding and assembly, including the Gim Complex/Prefoldin (GimC/PFD), the ring-shaped chaperonin TRiC/CCT, the Hsp70 SSA, and Hsp90 and its cofactors (McClellan et al., 2007; Taipale et al., 2012; Yam et al., 2008). These chaperones interact with smaller subsets of nascent polypeptides compared to ribosome-bound SSB/Hsp70 and NAC (McClellan et al., 2007; Taipale et al., 2012; Yam et al., 2008). While the organization and specificity of these downstream chaperones remains relativively unexplored, it is clear that TRiC/CCT and Hsp90 have distinct and obligate sets of substrates. TRiC/CCT associates co- and postranslationally with approximately 5–10% of newly made proteins (Yam et al., 2008), predominantly beta-sheet rich and topologically complex proteins that likely require the protected environment of its central cavity to fold (Douglas et al., 2011). Substrates appear to reach TRiC with the help of upstream-acting chaperones such as Hsp70 and GimC/PFD (Melville et al., 2003; Vainberg et al., 1998). Hsp90 is specialized in assisting the structural maturation and conformational regulation of numerous oligomeric complexes, structurally labile proteins, as well signal-transduction components (McClellan et al., 2007; Taipale et al., 2012). The soluble SSA isoforms of Hsp70 are predominantly responsible for cytoplasmic folding (Kim et al., 1998; Melville et al., 2003) and assisting translocation to the ER and mitochondria (McClellan and Brodsky, 2000; Young et al., 2003). This eukaryotic chaperone network for de novo folding is very robust, as deletions of single components like GimC/PFD, NAC or SSB are not lethal (Albanèse et al., 2010; Albanèse et al., 2006; Koplin et al., 2010). The network appears to be organized hierarchically, where Hsp70 and GimC act upstream of TRiC and Hsp90 (Hartl et al., 2011). This functional organization of hierarchical chaperone systems and dynamically regulated co-factors brings increased plasticity while maintaining specificity in chaperone mediated folding.
Quality control of nascent polypeptides at the ribosome
Newly made polypeptides can be ubiquitinated during or shortly after synthesis (Sato et al., 1998; Turner and Varshavsky, 2000). The extent of cotranslational ubiquitination and degradation has been intensely debated over the last 15 years (Vabulas and Hartl, 2005; Yewdell and Nicchitta, 2006). On the one hand, the misfolding potential of nascent, partially unfolded polypeptides suggests that significant fraction may be immediately degraded (Yewdell and Nicchitta, 2006). On the other hand, protein synthesis is energetically expensive and, the cell devotes many resources to regulating it. Several lines of evidence suggest that newly made proteins are largely protected from quality control (Frydman and Hartl, 1996; Vabulas and Hartl, 2005).
Translation of damaged mRNAs, such as those lacking a stop codon have been useful for understanding cotranslational protein quality control (Ito-Harashima et al., 2007). Translation of non-stop mRNAs (NS-mRNAs) generates proteins with poly-lysine tracts coded by the poly-A tail (Isken and Maquat, 2007; Klauer and van Hoof, 2012). The poly-lysine peptides stall the ribosome, likely by interacting with the negatively charged ribosomal tunnel wall. These aberrant, stalled complexes are recognized by a quality control pathway that dissociates the ribosomal subunits, recruits the exosome to degrade non-stop mRNAs (Tomecki et al., 2010; van Hoof and Parker, 2002), and targets the aberrant non-stop polypeptides stalled in the 60S subunit for ubiquitination and degradation (Ito-Harashima et al., 2007). Ubiquitination of the stalled nascent chain is carried out by the ribosome-associated Ubiquitin Ligase Listerin/Rkr1, (Bengtson and Joazeiro, 2010). Recently, Listerin/Rkr1 has been shown to form a complex with the AAA+ ATPase Cdc48, Tae2, and Ydr333C called the ribosome quality control complex (RQC) (Brandman et al., 2012). RQC contributes to the dissociation of the ribosomal subunits, and to coordinate clearance of the aberrant mRNA and nascent polypeptides (Brandman et al., 2012). Interestingly, RAC and SSB, while not required for polyubiquitination and degradation of poly-lysine containing chains, contribute to the stabilization of stalled complexes (Chiabudini et al., 2012). Because this pathway helps ensure faithful polypeptide synthesis by eliminating aberrant mRNAs, presumably after a single round of translation, it serves as a safeguard rather a general mechanism of nascent chain quality control. This function is underscored by the exceedingly low levels of Listerin in the cell, which is only present at ca. 200 copies per yeast cell. Disfunction of mRNA quality control is clearly deleterious for protein homeostasis, since Listerin mutants lead to neurodegeneration (Chu et al., 2009) and the RQC complex is linked to the signaling of the stress response by HSF1 (Brandman et al., 2012). Another E3 ligase routinely recruited to the ribosome is Not4 of the Ccr4/NOT complex, this ligase may play a role in either nascent chain ubiquitination or mRNA quality control (Collart and Panasenko, 2012).
Proteome homeostasis in health and disease
Nascent protein quality control will ultimately have to be understood as a systems level process impacting all aspects of cellular and organismal biology. As quality control capacity is limited, its overload can lead to proteostatic collapse (Powers et al., 2009). Excessive aggregation can directly impair protein quality control, impacting protein function in many cellular networks (Olzscha et al., 2011). An impairment of proteostatic capacity is a general hallmark of aging (Morimoto, 2008; Taylor and Dillin, 2011), often accompanied by widespread aggregation, particularly of ribosomal proteins (David et al., 2010). The immediate consequence of failed clearance of aggregated proteins can be the onset of neurodegenerative protein misfolding diseases that are especially prevalent during aging (Chiti and Dobson, 2006). In addition, deregulation of chaperone networks is exploited for uncontrolled cell proliferation in cancers (Whitesell and Lindquist, 2005) and helps drive aneuploidy (Oromendia et al., 2012). Molecular chaperones also play critical roles in virus replication (Geller et al., 2012). Advances in our understanding of individual mechanisms and pathways are prerequisite to piece together a more nuanced global understanding of protein homeostasis.
Perspectives
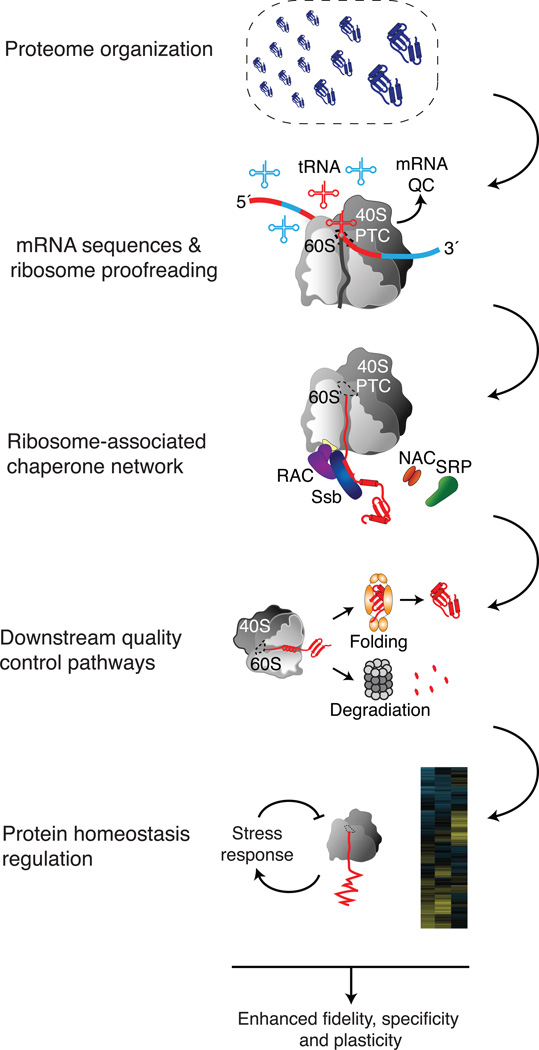
The cell uses diverse, dynamic, and adaptive strategies to ensure quality and fidelity of new proteins. (Figure 4). For example, genomic sequences have generally evolved to avoid aggregation, limit misincorporation of critical amino acids, and promote cotranslational folding. During translation, the ribosome actively monitors all aspects of protein biosynthesis, dynamically responding to errors, and influencing the fate of the nascent protein. The cell manages important evolutionary trade-offs with its quality control strategies (Wagner, 2008). Stringent quality control is more energetically expensive, and more static than less stringent control, complicating adaptation to changing conditions. Conversely, less stringent quality control may be more adaptable, but allows more damaged proteins to reach the cellular milieu, thus increasing the risk of protein aggregation. The combination of mechanisms provides an elegant way to maximize fidelity, specificity, and plasticity of protein quality control(Figure 4).
Figure 4. Sequential quality control of newly made proteins.
The cell relies on tiered quality control mechanisms. Most proteins are short and fold readily, while long and folding-challenged but often functionally important proteins can rely on chaperone assistance. Individual sequences are optimized for translational fidelity, thus reducing the risk of phenotypic missense mutations, as well as to facilitate cotranslational folding. A selective ribosome-associated chaperone network binds nascent polypeptides to promote their folding or translocation. Co-chaperones add specificity and plasticity to the selection of substrates and downstream folding/degradation pathways. Stresses like heat shock can temporarily rebalance the burden of newly made proteins and chaperone capacity. This sequential organization achieves high fidelity, specificity, and plasticity.
Many exciting and pressing questions remain. Does the ribosome sense the nascent chain inside the ribosome exit tunnel to influence translocation, degradation, and folding through the recruitment of ribosome-associated chaperones and related factors? If so, how? We are only beginning to understand the signaling from within the exit tunnel and its consequences on protein fate. Furthermore, we still don’t understand how aspects of the cellular environment, such as the metabolic state of the cell, are signaled to folding networks. It would also be interesting to explore the spatial control of translation and protein quality. Ribosomes translating proteins that are destined for export into the ER or membrane are often pre-localized near their functional environment. Currently, it is not clear if and how these specialized ribosomes are marked and targeted to specific locations,. Finally, it will be useful to examine the the kinetics of translation elongation and how it might be attuned to the cotranslational folding, as this will be important for understanding the connections between these processes.. It remains to be seen if translation elongation directly coordinates the cotranslational binding of chaperones. A better understanding of these questions will open the door to a general integrated framework of protein folding in the cell. The organization and execution of the quality control of newly made proteins in the cell is a remarkable feat waiting to be further uncovered.
Acknowledgements
We thank the Frydman Lab for helpful discussions. We gratefully acknowledge support from EMBO Long-Term Fellowships to S.P. and F.W., and NIH grant GM56433 and AI91575 to J.F.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Agashe V, Guha S, Chang H-C, Genevaux P, Hayer-Hartl M, Stemp M, Georgopoulos C, Hartl F, Barral J. Function of trigger factor and DnaK in multidomain protein folding: increase in yield at the expense of folding speed. Cell. 2004;117:199–209. doi: 10.1016/s0092-8674(04)00299-5. [DOI] [PubMed] [Google Scholar]
- Akashi H. Synonymous codon usage in Drosophila melanogaster: natural selection and translational accuracy. Genetics. 1994;136:927–935. doi: 10.1093/genetics/136.3.927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alamo MD, Hogan DJ, Pechmann S, Albanese V, Brown PO, Frydman J. Defining the Specificity of Cotranslationally Acting Chaperones by Systematic Analysis of mRNAs Associated with Ribosome-Nascent Chain Complexes. PLoS Biol. 2011;9:e1001100. doi: 10.1371/journal.pbio.1001100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albanèse V, Reissmann S, Frydman J. A ribosome-anchored chaperone network that facilitates eukaryotic ribosome biogenesis. The Journal of Cell Biology. 2010;189:69–81. doi: 10.1083/jcb.201001054. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Albanèse V, Yam AY-W, Baughman J, Parnot C, Frydman J. Systems analyses reveal two chaperone networks with distinct functions in eukaryotic cells. Cell. 2006;124:75–88. doi: 10.1016/j.cell.2005.11.039. [DOI] [PubMed] [Google Scholar]
- Baldwin AJ, Knowles TPJ, Tartaglia GG, Fitzpatrick AW, Devlin GL, Shammas SL, Waudby CA, Mossuto MF, Meehan S, Gras SL, et al. Metastability of native proteins and the phenomenon of amyloid formation. Journal of the American Chemical Society. 2011;133:14160–14163. doi: 10.1021/ja2017703. [DOI] [PubMed] [Google Scholar]
- Beatrix B, Sakai H, Wiedmann M. The alpha and beta subunit of the nascent polypeptide-associated complex have distinct functions. J Biol Chem. 2000;275:37838–37845. doi: 10.1074/jbc.M006368200. [DOI] [PubMed] [Google Scholar]
- Beckmann RP, Mizzen LE, Welch WJ. Interaction of Hsp 70 with newly synthesized proteins: implications for protein folding and assembly. Science. 1990;248:850–854. doi: 10.1126/science.2188360. [DOI] [PubMed] [Google Scholar]
- Bengtson MH, Joazeiro CAP. Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature. 2010;467:470–473. doi: 10.1038/nature09371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berndt U, Oellerer S, Zhang Y, Johnson AE, Rospert S. A signal-anchor sequence stimulates signal recognition particle binding to ribosomes from inside the exit tunnel. Proc Natl Acad Sci USA. 2009;106:1398–1403. doi: 10.1073/pnas.0808584106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhushan S, Gartmann M, Halic M, Armache J-P, Jarasch A, Mielke T, Berninghausen O, Wilson DN, Beckmann R. alpha-Helical nascent polypeptide chains visualized within distinct regions of the ribosomal exit tunnel. Nature Structural & Molecular Biology. 2010a;17:313–317. doi: 10.1038/nsmb.1756. [DOI] [PubMed] [Google Scholar]
- Bhushan S, Meyer H, Starosta AL, Becker T, Mielke T, Berninghausen O, Sattler M, Wilson DN, Beckmann R. Structural basis for translational stalling by human cytomegalovirus and fungal arginine attenuator peptide. Molecular Cell. 2010b;40:138–146. doi: 10.1016/j.molcel.2010.09.009. [DOI] [PubMed] [Google Scholar]
- Bornemann T, Jöckel J, Rodnina MV, Wintermeyer W. Signal sequence-independent membrane targeting of ribosomes containing short nascent peptides within the exit tunnel. Nat Struct Mol Biol. 2008;15:494–499. doi: 10.1038/nsmb.1402. [DOI] [PubMed] [Google Scholar]
- Bowman GR, Voelz VA, Pande VS. Taming the complexity of protein folding. Curr Opin Struct Biol. 2011;21:4–11. doi: 10.1016/j.sbi.2010.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brandman O, Stewart-Ornstein J, Wong D, Larson A, Williams CC, Li G-W, Zhou S, King D, Shen PS, Weibezahn J, et al. A Ribosome-Bound Quality Control Complex Triggers Degradation of Nascent Peptides and Signals Translation Stress Cell. 2012;151:1042–10154. doi: 10.1016/j.cell.2012.10.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buell AK, Tartaglia GG, Birkett NR, Waudby CA, Vendruscolo M, Salvatella X, Welland ME, Dobson CM, Knowles TPJ. Position-dependent electrostatic protection against protein aggregation. ChemBioChem. 2009;10:1309–1312. doi: 10.1002/cbic.200900144. [DOI] [PubMed] [Google Scholar]
- Cabrita LD, Dobson CM, Christodoulou J. Protein folding on the ribosome. Curr Opin Struct Biol. 2010;20:33–45. doi: 10.1016/j.sbi.2010.01.005. [DOI] [PubMed] [Google Scholar]
- Calloni G, Chen T, Schermann SM, Chang H-C, Genevaux P, Agostini F, Tartaglia GG, Hayer-Hartl M, Hartl FU. DnaK functions as a central hub in the E. coli chaperone network. CellReports. 2012;1:251–264. doi: 10.1016/j.celrep.2011.12.007. [DOI] [PubMed] [Google Scholar]
- Chiabudini M, Conz C, Reckmann F, Rospert S. Ribosome-Associated Complex and Ssb Are Required for Translational Repression Induced by Polylysine Segments within Nascent Chains. Molecular and Cellular Biology. 2012;32:4769–4779. doi: 10.1128/MCB.00809-12. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chiti F, Dobson CM. Protein misfolding, functional amyloid, and human disease. Annu Rev Biochem. 2006;75:333–366. doi: 10.1146/annurev.biochem.75.101304.123901. [DOI] [PubMed] [Google Scholar]
- Chu J, Hong NA, Masuda CA, Jenkins BV, Nelms KA, Goodnow CC, Glynne RJ, Wu H, Masliah E, Joazeiro CAP, et al. A mouse forward genetics screen identifies LISTERIN as an E3 ubiquitin ligase involved in neurodegeneration. Proc Natl Acad Sci USA. 2009;106:2097–2103. doi: 10.1073/pnas.0812819106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarke TF, Clark PL. Rare codons cluster. PLoS ONE. 2008;3:e3412. doi: 10.1371/journal.pone.0003412. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collart MA, Panasenko OO. The Ccr4--not complex. Gene. 2012;492:42–53. doi: 10.1016/j.gene.2011.09.033. [DOI] [PubMed] [Google Scholar]
- Cruz-Vera LR, Gong M, Yanofsky C. Changes produced by bound tryptophan in the ribosome peptidyl transferase center in response to TnaC, a nascent leader peptide. Proc Natl Acad Sci U S A. 2006;103:3598–3603. doi: 10.1073/pnas.0600082103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- David DC, Ollikainen N, Trinidad JC, Cary MP, Burlingame AL, Kenyon C. Widespread Protein Aggregation as an Inherent Part of Aging in C. elegans. PLoS Biol. 2010;8 doi: 10.1371/journal.pbio.1000450. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DePristo MA, Weinreich DM, Hartl DL. Missense meanderings in sequence space: a biophysical view of protein evolution. Nature reviews Genetics. 2005;6:678–687. doi: 10.1038/nrg1672. [DOI] [PubMed] [Google Scholar]
- Deuerling E, Schulze-Specking A, Tomoyasu T, Mogk A, Bukau B. Trigger factor and DnaK cooperate in folding of newly synthesized proteins. Nature. 1999;400:693–696. doi: 10.1038/23301. [DOI] [PubMed] [Google Scholar]
- dos Reis M, Savva R, Wernisch L. Solving the riddle of codon usage preferences: a test for translational selection. Nucleic Acids Res. 2004;32:5036–5044. doi: 10.1093/nar/gkh834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Douglas NR, Reissmann S, Zhang J, Chen B, Jakana J, Kumar R, Chiu W, Frydman J. Dual Action of ATP Hydrolysis Couples Lid Closure to Substrate Release into the Group II Chaperonin Chamber. Cell. 2011;144:240–252. doi: 10.1016/j.cell.2010.12.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond DA, Bloom JD, Adami C, Wilke CO, Arnold FH. Why highly expressed proteins evolve slowly. Proc Natl Acad Sci USA. 2005;102:14338–14343. doi: 10.1073/pnas.0504070102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drummond DA, Wilke CO. Mistranslation-induced protein misfolding as a dominant constraint on coding-sequence evolution. Cell. 2008;134:341–352. doi: 10.1016/j.cell.2008.05.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duncan C, Mata J. Widespread cotranslational formation of protein complexes. PLoS genetics. 2011;7:e1002398. doi: 10.1371/journal.pgen.1002398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eaton WA, Muñoz V, Hagen SJ, Jas GS, Lapidus LJ, Henry ER, Hofrichter J. Fast kinetics and mechanisms in protein folding. Annu Rev Biophys Biomol Struct. 2000;29:327–359. doi: 10.1146/annurev.biophys.29.1.327. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fares MA, Ruiz-González MX, Moya A, Elena SF, Barrio E. Endosymbiotic bacteria: groEL buffers against deleterious mutations. Nature. 2002;417:398. doi: 10.1038/417398a. [DOI] [PubMed] [Google Scholar]
- Fedyukina DV, Cavagnero S. Protein folding at the exit tunnel. Annual review of biophysics. 2011;40:337–359. doi: 10.1146/annurev-biophys-042910-155338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferbitz L, Maier T, Patzelt H, Bukau B, Deuerling E, Ban N. Trigger factor in complex with the ribosome forms a molecular cradle for nascent proteins. Nature. 2004;431:590–596. doi: 10.1038/nature02899. [DOI] [PubMed] [Google Scholar]
- Fowler DM, Koulov AV, Balch WE, Kelly JW. Functional amyloid--from bacteria to humans. Trends Biochem Sci. 2007;32:217–224. doi: 10.1016/j.tibs.2007.03.003. [DOI] [PubMed] [Google Scholar]
- Frydman J, Erdjument-Bromage H, Tempst P, Hartl FU. Cotranslational domain folding as the structural basis for the rapid de novo folding of firefly luciferase. Nat Struct Biol. 1999;6:697–705. doi: 10.1038/10754. [DOI] [PubMed] [Google Scholar]
- Frydman J, Hartl FU. Principles of chaperone-assisted protein folding: differences between in vitro and in vivo mechanisms. Science. 1996;272:1497–1502. doi: 10.1126/science.272.5267.1497. [DOI] [PubMed] [Google Scholar]
- Frydman J, Nimmesgern E, Ohtsuka K, Hartl F. Folding of nascent polypeptide chains in a high molecular mass assembly with molecular chaperones. Nature. 1994;370:111–117. doi: 10.1038/370111a0. [DOI] [PubMed] [Google Scholar]
- Geller R, Taguwa S, Frydman J. Broad action of Hsp90 as a host chaperone required for viral replication. Biochim Biophys Acta. 2012;1823:698–706. doi: 10.1016/j.bbamcr.2011.11.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- George R, Beddoe T, Landl K, Lithgow T. The yeast nascent polypeptide-associated complex initiates protein targeting to mitochondria in vivo. Proc Natl Acad Sci USA. 1998;95:2296–2301. doi: 10.1073/pnas.95.5.2296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gershenson A, Gierasch LM. Protein folding in the cell: challenges and progress. Curr Opin Struct Biol. 2011;21:32–41. doi: 10.1016/j.sbi.2010.11.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldschmidt L, Teng PK, Riek R, Eisenberg D. Identifying the amylome, proteins capable of forming amyloid-like fibrils. Proc Natl Acad Sci USA. 2010;107:3487–3492. doi: 10.1073/pnas.0915166107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gong F, Yanofsky C. Instruction of translating ribosome by nascent peptide. Science. 2002;297:1864–1867. doi: 10.1126/science.1073997. [DOI] [PubMed] [Google Scholar]
- Gsponer J, Babu MM. Cellular Strategies for Regulating Functional and Nonfunctional Protein Aggregation. CellReports. 2012 doi: 10.1016/j.celrep.2012.09.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gu W, Zhou T, Wilke CO. A universal trend of reduced mRNA stability near the translation-initiation site in prokaryotes and eukaryotes. PLoS Comput Biol. 2010;6:e1000664. doi: 10.1371/journal.pcbi.1000664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han J-H, Batey S, Nickson AA, Teichmann SA, Clarke J. The folding and evolution of multidomain proteins. Nat Rev Mol Cell Biol. 2007;8:319–330. doi: 10.1038/nrm2144. [DOI] [PubMed] [Google Scholar]
- Hartl FU, Bracher A, Hayer-Hartl M. Molecular chaperones in protein folding and proteostasis. Nature. 2011;475:324–332. doi: 10.1038/nature10317. [DOI] [PubMed] [Google Scholar]
- Houry WA, Frishman D, Eckerskorn C, Lottspeich F, Hartl FU. Identification of in vivo substrates of the chaperonin GroEL. Nature. 1999;402:147–154. doi: 10.1038/45977. [DOI] [PubMed] [Google Scholar]
- Huang P, Gautschi M, Walter W, Rospert S, Craig E. The Hsp70 Ssz1 modulates the function of the ribosome-associated J-protein Zuo1. Nature Structural and Molecular Biology. 2005;12:497–504. doi: 10.1038/nsmb942. [DOI] [PubMed] [Google Scholar]
- Isken O, Maquat LE. Quality control of eukaryotic mRNA: safeguarding cells from abnormal mRNA function. Genes Dev. 2007;21:1833–1856. doi: 10.1101/gad.1566807. [DOI] [PubMed] [Google Scholar]
- Ito-Harashima S, Kuroha K, Tatematsu T, Inada T. Translation of the poly(A) tail plays crucial roles in nonstop mRNA surveillance via translation repression and protein destabilization by proteasome in yeast. Genes Dev. 2007;21:519–524. doi: 10.1101/gad.1490207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jahn TR, Radford SE. Folding versus aggregation: polypeptide conformations on competing pathways. Arch Biochem Biophys. 2008;469:100–117. doi: 10.1016/j.abb.2007.05.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaiswal H, Conz C, Otto H, Wölfle T, Fitzke E, Mayer MP, Rospert S. The chaperone network connected to human ribosome-associated complex (mRAC) Molecular and Cellular Biology. 2011;31:1160–1173. doi: 10.1128/MCB.00986-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarosz DF, Taipale M, Lindquist S. Protein homeostasis and the phenotypic manifestation of genetic diversity: principles and mechanisms. Annu Rev Genet. 2010;44:189–216. doi: 10.1146/annurev.genet.40.110405.090412. [DOI] [PubMed] [Google Scholar]
- Johansson M, Zhang J, Ehrenberg M. Genetic code translation displays a linear trade-off between efficiency and accuracy of tRNA selection. Proc Natl Acad Sci USA. 2012;109:131–136. doi: 10.1073/pnas.1116480109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaganovich D, Kopito R, Frydman J. Misfolded proteins partition between two distinct quality control compartments. Nature. 2008;454:1088–1095. doi: 10.1038/nature07195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaiser C, Chang H-C, Agashe V, Lakshmipathy S, Etchells S, Hayer-Hartl M, Hartl F, Barral J. Real-time observation of trigger factor function on translating ribosomes. Nature. 2006;444:455–460. doi: 10.1038/nature05225. [DOI] [PubMed] [Google Scholar]
- Kerner M, Naylor D, Ishihama Y, Maier T, Chang H-C, Stines A, Georgopoulos C, Frishman D, Hayer-Hartl M, Mann M, et al. Proteome-wide analysis of chaperonin-dependent protein folding in Escherichia coli. Cell. 2005;122:209–220. doi: 10.1016/j.cell.2005.05.028. [DOI] [PubMed] [Google Scholar]
- Khushoo A, Yang Z, Johnson AE, Skach WR. Ligand-driven vectorial folding of ribosome-bound human CFTR NBD1. Molecular Cell. 2011;41:682–692. doi: 10.1016/j.molcel.2011.02.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim S, Schilke B, Craig EA, Horwich AL. Folding in vivo of a newly translated yeast cytosolic enzyme is mediated by the SSA class of cytosolic yeast Hsp70 proteins. Proc Natl Acad Sci USA. 1998;95:12860–12865. doi: 10.1073/pnas.95.22.12860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kimchi-Sarfaty C, Oh JM, Kim I-W, Sauna ZE, Calcagno AM, Ambudkar SV, Gottesman MM. A "silent" polymorphism in the MDR1 gene changes substrate specificity. Science. 2007;315:525–528. doi: 10.1126/science.1135308. [DOI] [PubMed] [Google Scholar]
- Klauer AA, van Hoof A. Degradation of mRNAs that lack a stop codon: a decade of nonstop progress. Wiley Interdiscip Rev RNA. 2012;3:649–660. doi: 10.1002/wrna.1124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klinge S, Voigts-Hoffmann F, Leibundgut M, Ban N. Atomic structures of the eukaryotic ribosome. Trends Biochem Sci. 2012;37:189–198. doi: 10.1016/j.tibs.2012.02.007. [DOI] [PubMed] [Google Scholar]
- Komar AA. A pause for thought along the co-translational folding pathway. Trends Biochem Sci. 2009;34:16–24. doi: 10.1016/j.tibs.2008.10.002. [DOI] [PubMed] [Google Scholar]
- Koplin A, Preissler S, Ilina Y, Koch M, Scior A, Erhardt M, Deuerling E. A dual function for chaperones SSB-RAC and the NAC nascent polypeptide-associated complex on ribosomes. The Journal of cell biology. 2010;189:57–68. doi: 10.1083/jcb.200910074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kramer G, Boehringer D, Ban N, Bukau B. The ribosome as a platform for co-translational processing, folding and targeting of newly synthesized proteins. Nat Struct Mol Biol. 2009;16:589–597. doi: 10.1038/nsmb.1614. [DOI] [PubMed] [Google Scholar]
- Lakshmipathy SK, Tomic S, Kaiser CM, Chang H-C, Genevaux P, Georgopoulos C, Barral JM, Johnson AE, Hartl FU, Etchells SA. Identification of nascent chain interaction sites on trigger factor. The Journal of biological chemistry. 2007;282:12186–12193. doi: 10.1074/jbc.M609871200. [DOI] [PubMed] [Google Scholar]
- Lee Y, Zhou T, Tartaglia GG, Vendruscolo M, Wilke CO. Translationally optimal codons associate with aggregation-prone sites in proteins. Proteomics. 2010;10:4163–4171. doi: 10.1002/pmic.201000229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leidig C, Bange G, Kopp J, Amlacher S, Aravind A, Wickles S, Witte G, Hurt E, Beckmann R, Sinning I. Structural characterization of a eukaryotic chaperone-the ribosome-associated complex. Nat Struct Mol Biol. 2013;20:23–28. doi: 10.1038/nsmb.2447. [DOI] [PubMed] [Google Scholar]
- Levy ED, Pereira-Leal JB. Evolution and dynamics of protein interactions and networks. Curr Opin Struct Biol. 2008;18:349–357. doi: 10.1016/j.sbi.2008.03.003. [DOI] [PubMed] [Google Scholar]
- Li G-W, Oh E, Weissman JS. The anti-Shine-Dalgarno sequence drives translational pausing and codon choice in bacteria. Nature. 2012;484:538–541. doi: 10.1038/nature10965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin MM, Zewail AH. Hydrophobic forces and the length limit of foldable protein domains. Proc Natl Acad Sci USA. 2012;109:9851–9856. doi: 10.1073/pnas.1207382109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin P-J, Jongsma CG, Pool MR, Johnson AE. Polytopic membrane protein folding at L17 in the ribosome tunnel initiates cyclical changes at the translocon. The Journal of Cell Biology. 2011;195:55–70. doi: 10.1083/jcb.201103118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lu J, Deutsch C. Folding zones inside the ribosomal exit tunnel. Nat Struct Mol Biol. 2005;12:1123–1129. doi: 10.1038/nsmb1021. [DOI] [PubMed] [Google Scholar]
- Markesich DC, Gajewski KM, Nazimiec ME, Beckingham K. bicaudal encodes the Drosophila beta NAC homolog, a component of the ribosomal translational machinery. Development. 2000;127:559–572. doi: 10.1242/dev.127.3.559. [DOI] [PubMed] [Google Scholar]
- Martinez-Hackert E, Hendrickson WA. Promiscuous substrate recognition in folding and assembly activities of the trigger factor chaperone. Cell. 2009;138:923–934. doi: 10.1016/j.cell.2009.07.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClellan AJ, Brodsky JL. Mutation of the ATP-binding pocket of SSA1 indicates that a functional interaction between Ssa1p and Ydj1p is required for post-translational translocation into the yeast endoplasmic reticulum. Genetics. 2000;156:501–512. doi: 10.1093/genetics/156.2.501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McClellan AJ, Xia Y, Deutschbauer AM, Davis RW, Gerstein M, Frydman J. Diverse cellular functions of the Hsp90 molecular chaperone uncovered using systems approaches. Cell. 2007;131:121–135. doi: 10.1016/j.cell.2007.07.036. [DOI] [PubMed] [Google Scholar]
- Melnikov S, Ben-Shem A, Garreau de Loubresse N, Jenner L, Yusupova G, Yusupov M. One core, two shells: bacterial and eukaryotic ribosomes. Nat Struct Mol Biol. 2012;19:560–567. doi: 10.1038/nsmb.2313. [DOI] [PubMed] [Google Scholar]
- Melville MW, McClellan AJ, Meyer AS, Darveau A, Frydman J. The Hsp70 and TRiC/CCT chaperone systems cooperate in vivo to assemble the von Hippel-Lindau tumor suppressor complex. Mol Cell Biol. 2003;23:3141–3151. doi: 10.1128/MCB.23.9.3141-3151.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morimoto RI. Proteotoxic stress and inducible chaperone networks in neurodegenerative disease and aging. Genes Dev. 2008;22:1427–1438. doi: 10.1101/gad.1657108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakatogawa H, Ito K. Secretion monitor, SecM, undergoes self-translation arrest in the cytosol. Molecular Cell. 2001;7:185–192. doi: 10.1016/s1097-2765(01)00166-6. [DOI] [PubMed] [Google Scholar]
- Nakatogawa H, Ito K. The ribosomal exit tunnel functions as a discriminating gate. Cell. 2002;108:629–636. doi: 10.1016/s0092-8674(02)00649-9. [DOI] [PubMed] [Google Scholar]
- O'Brien EP, Hsu S-TD, Christodoulou J, Vendruscolo M, Dobson CM. Transient Tertiary Structure Formation within the Ribosome Exit Port. J Am Chem Soc. 2010;132:16928–16937. doi: 10.1021/ja106530y. [DOI] [PubMed] [Google Scholar]
- O'Brien EP, Vendruscolo M, Dobson CM. Prediction of variable translation rate effects on cotranslational protein folding. Nature Communications. 2012;3:868. doi: 10.1038/ncomms1850. [DOI] [PubMed] [Google Scholar]
- Ogle JM, Ramakrishnan V. Structural insights into translational fidelity. Annual review of biochemistry. 2005;74:129–177. doi: 10.1146/annurev.biochem.74.061903.155440. [DOI] [PubMed] [Google Scholar]
- Oh E, Becker A, Sandikci A, Huber D, Chaba R, Gloge F, Nichols R, Typas A, Gross C, Kramer G, et al. Selective Ribosome Profiling Reveals the Cotranslational Chaperone Action of Trigger Factor In Vivo. Cell. 2011;147:1295–1308. doi: 10.1016/j.cell.2011.10.044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olzscha H, Schermann SM, Woerner AC, Pinkert S, Hecht MH, Tartaglia GG, Vendruscolo M, Hayer-Hartl M, Hartl FU, Vabulas RM. Amyloid-like Aggregates Sequester Numerous Metastable Proteins with Essential Cellular Functions. Cell. 2011;144:67–78. doi: 10.1016/j.cell.2010.11.050. [DOI] [PubMed] [Google Scholar]
- Oromendia AB, Dodgson SE, Amon A. Aneuploidy causes proteotoxic stress in yeast. Genes Dev. 2012;26:2696–2708. doi: 10.1101/gad.207407.112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ouyang Z, Liang J. Predicting protein folding rates from geometric contact and amino acid sequence. Protein Sci. 2008;17:1256–1263. doi: 10.1110/ps.034660.108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pech M, Spreter T, Beckmann R, Beatrix B. Dual binding mode of the nascent polypeptide-associated complex (NAC) reveals a novel universal adapter site on the ribosome. Journal of Biological Chemistry. 2010;285:19679–19687. doi: 10.1074/jbc.M109.092536. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pechmann S, Frydman J. Evolutionary conservation of codon optimality reveals hidden signatures of co-translational folding. Nat Struct Mol Biol. 2012 doi: 10.1038/nsmb.2466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pechmann S, Vendruscolo M. Derivation of a solubility condition for proteins from an analysis of the competition between folding and aggregation. Mol BioSyst. 2010;6:2490–2497. doi: 10.1039/c005160h. [DOI] [PubMed] [Google Scholar]
- Powers ET, Morimoto RI, Dillin A, Kelly JW, Balch WE. Biological and chemical approaches to diseases of proteostasis deficiency. Annual review of biochemistry. 2009;78:959–991. doi: 10.1146/annurev.biochem.052308.114844. [DOI] [PubMed] [Google Scholar]
- Preissler S, Deuerling E. Ribosome-associated chaperones as key players in proteostasis. Trends in Biochemical Sciences. 2012;37:274–283. doi: 10.1016/j.tibs.2012.03.002. [DOI] [PubMed] [Google Scholar]
- Raue U, Oellerer S, Rospert S. Association of protein biogenesis factors at the yeast ribosomal tunnel exit is affected by the translational status and nascent polypeptide sequence. The Journal of biological chemistry. 2007;282:7809–7816. doi: 10.1074/jbc.M611436200. [DOI] [PubMed] [Google Scholar]
- Raviol H, Sadlish H, Rodriguez F, Mayer MP, Bukau B. Chaperone network in the yeast cytosol: Hsp110 is revealed as an Hsp70 nucleotide exchange factor. The EMBO Journal. 2006;25:2510–2518. doi: 10.1038/sj.emboj.7601139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato S, Ward CL, Kopito RR. Cotranslational ubiquitination of cystic fibrosis transmembrane conductance regulator in vitro. J Biol Chem. 1998;273:7189–7192. doi: 10.1074/jbc.273.13.7189. [DOI] [PubMed] [Google Scholar]
- Sawaya MR, Sambashivan S, Nelson R, Ivanova MI, Sievers SA, Apostol MI, Thompson MJ, Balbirnie M, Wiltzius JJW, McFarlane HT, et al. Atomic structures of amyloid cross-beta spines reveal varied steric zippers. Nature. 2007;447:453–457. doi: 10.1038/nature05695. [DOI] [PubMed] [Google Scholar]
- Schmeing TM, Ramakrishnan V. What recent ribosome structures have revealed about the mechanism of translation. Nature. 2009;461:1234–1242. doi: 10.1038/nature08403. [DOI] [PubMed] [Google Scholar]
- Shaner L, Wegele H, Buchner J, Morano KA. The yeast Hsp110 Sse1 functionally interacts with the Hsp70 chaperones Ssa and Ssb. J Biol Chem. 2005;280:41262–41269. doi: 10.1074/jbc.M503614200. [DOI] [PubMed] [Google Scholar]
- Spevak CC, Ivanov IP, Sachs MS. Sequence requirements for ribosome stalling by the arginine attenuator peptide. J Biol Chem. 2010;285:40933–40942. doi: 10.1074/jbc.M110.164152. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Taipale M, Krykbaeva I, Koeva M, Kayatekin C, Westover KD, Karras GI, Lindquist S. Quantitative analysis of hsp90-client interactions reveals principles of substrate recognition. Cell. 2012;150:987–1001. doi: 10.1016/j.cell.2012.06.047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Takyar S, Hickerson RP, Noller HF. mRNA helicase activity of the ribosome. Cell. 2005;120:49–58. doi: 10.1016/j.cell.2004.11.042. [DOI] [PubMed] [Google Scholar]
- Tanner DR, Cariello DA, Woolstenhulme CJ, Broadbent MA, Buskirk AR. Genetic identification of nascent peptides that induce ribosome stalling. J Biol Chem. 2009;284:34809–34818. doi: 10.1074/jbc.M109.039040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tartaglia GG, Dobson CM, Hartl FU, Vendruscolo M. Physicochemical Determinants of Chaperone Requirements. Journal of Molecular Biology. 2010 doi: 10.1016/j.jmb.2010.03.066. [DOI] [PubMed] [Google Scholar]
- Tartaglia GG, Pechmann S, Dobson CM, Vendruscolo M. Life on the edge: a link between gene expression levels and aggregation rates of human proteins. Trends Biochem Sci. 2007;32:204–206. doi: 10.1016/j.tibs.2007.03.005. [DOI] [PubMed] [Google Scholar]
- Taylor RC, Dillin A. Aging as an event of proteostasis collapse. Cold Spring Harbor Perspectives in Biology. 2011;3 doi: 10.1101/cshperspect.a004440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Teter S, Houry W, Ang D, Tradler T, Rockabrand D, Fischer G, Blum P, Georgopoulos C, Hartl F. Polypeptide flux through bacterial Hsp70: DnaK cooperates with trigger factor in chaperoning nascent chains. Cell. 1999;97:755–765. doi: 10.1016/s0092-8674(00)80787-4. [DOI] [PubMed] [Google Scholar]
- Tomecki R, Drazkowska K, Dziembowski A. Mechanisms of RNA degradation by the eukaryotic exosome. ChemBioChem. 2010;11:938–945. doi: 10.1002/cbic.201000025. [DOI] [PubMed] [Google Scholar]
- Tsai C-J, Ma B, Nussinov R. Protein-protein interaction networks: how can a hub protein bind so many different partners? Trends in Biochemical Sciences. 2009;34:594–600. doi: 10.1016/j.tibs.2009.07.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuller T, Carmi A, Vestsigian K, Navon S, Dorfan Y, Zaborske J, Pan T, Dahan O, Furman I, Pilpel Y. An evolutionarily conserved mechanism for controlling the efficiency of protein translation. Cell. 2010;141:344–354. doi: 10.1016/j.cell.2010.03.031. [DOI] [PubMed] [Google Scholar]
- Turner GC, Varshavsky A. Detecting and measuring cotranslational protein degradation in vivo. Science. 2000;289:2117–2120. doi: 10.1126/science.289.5487.2117. [DOI] [PubMed] [Google Scholar]
- Tyedmers J, Mogk A, Bukau B. Cellular strategies for controlling protein aggregation. Nat Rev Mol Cell Biol. 2010;11:777–788. doi: 10.1038/nrm2993. [DOI] [PubMed] [Google Scholar]
- Vabulas RM, Hartl FU. Protein synthesis upon acute nutrient restriction relies on proteasome function. Science. 2005;310:1960–1963. doi: 10.1126/science.1121925. [DOI] [PubMed] [Google Scholar]
- Vainberg I, Lewis S, Rommelaere H, Ampe C, Vandekerckhove J, Klein H, Cowan N. Prefoldin, a chaperone that delivers unfolded proteins to cytosolic chaperonin. Cell. 1998;93:863–873. doi: 10.1016/s0092-8674(00)81446-4. [DOI] [PubMed] [Google Scholar]
- van Hoof A, Parker R. Messenger RNA degradation: beginning at the end. Curr Biol. 2002;12:R285–R287. doi: 10.1016/s0960-9822(02)00802-3. [DOI] [PubMed] [Google Scholar]
- Vavouri T, Semple JI, Garcia-Verdugo R, Lehner B. Intrinsic protein disorder and interaction promiscuity are widely associated with dosage sensitivity. Cell. 2009;138:198–208. doi: 10.1016/j.cell.2009.04.029. [DOI] [PubMed] [Google Scholar]
- Wagner A. Robustness and evolvability: a paradox resolved. Proc Biol Sci. 2008;275:91–100. doi: 10.1098/rspb.2007.1137. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitesell L, Lindquist SL. HSP90 and the chaperoning of cancer. Nat Rev Cancer. 2005;5:761–772. doi: 10.1038/nrc1716. [DOI] [PubMed] [Google Scholar]
- Willmund F, del Alamo M, Pechmann S, Chen T, Albanese V, Dammer EB, Peng J, Frydman J. The cotranslational function of ribosome-associated Hsp70 in eukaryotic protein homeostasis. Cell. 2013 doi: 10.1016/j.cell.2012.12.001. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson DN, Beckmann R. The ribosomal tunnel as a functional environment for nascent polypeptide folding and translational stalling. Current Opinion in Structural Biology. 2011;21:274–282. doi: 10.1016/j.sbi.2011.01.007. [DOI] [PubMed] [Google Scholar]
- Woolhead C, McCormick P, Johnson A. Nascent membrane and secretory proteins differ in FRET-detected folding far inside the ribosome and in their exposure to ribosomal proteins. Cell. 2004;116:725–736. doi: 10.1016/s0092-8674(04)00169-2. [DOI] [PubMed] [Google Scholar]
- Wright CF, Teichmann SA, Clarke J, Dobson CM. The importance of sequence diversity in the aggregation and evolution of proteins. Nature. 2005;438:878–881. doi: 10.1038/nature04195. [DOI] [PubMed] [Google Scholar]
- Yam AY, Xia Y, Lin H-TJ, Burlingame A, Gerstein M, Frydman J. Defining the TRiC/CCT interactome links chaperonin function to stabilization of newly made proteins with complex topologies. Nature Structural & Molecular Biology. 2008;15:1255–1262. doi: 10.1038/nsmb.1515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yam AY-W, Albanèse V, Lin H-TJ, Frydman J. Hsp110 cooperates with different cytosolic HSP70 systems in a pathway for de novo folding. J Biol Chem. 2005;280:41252–41261. doi: 10.1074/jbc.M503615200. [DOI] [PubMed] [Google Scholar]
- Yewdell JW, Nicchitta CV. The DRiP hypothesis decennial: support, controversy, refinement and extension. Trends Immunol. 2006;27:368–373. doi: 10.1016/j.it.2006.06.008. [DOI] [PubMed] [Google Scholar]
- Young JC, Hoogenraad NJ, Hartl FU. Molecular chaperones Hsp90 and Hsp70 deliver preproteins to the mitochondrial import receptor Tom70. Cell. 2003;112:41–50. doi: 10.1016/s0092-8674(02)01250-3. [DOI] [PubMed] [Google Scholar]
- Zaher HS, Green R. Fidelity at the molecular level: lessons from protein synthesis. Cell. 2009a;136:746–762. doi: 10.1016/j.cell.2009.01.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaher HS, Green R. Quality control by the ribosome following peptide bond formation. Nature. 2009b;457:161–166. doi: 10.1038/nature07582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang F, Saha S, Shabalina SA, Kashina A. Differential arginylation of actin isoforms is regulated by coding sequence-dependent degradation. Science. 2010;329:1534–1537. doi: 10.1126/science.1191701. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang G, Ignatova Z. Folding at the birth of the nascent chain: coordinating translation with co-translational folding. Curr Opin Struct Biol. 2011;21:25–31. doi: 10.1016/j.sbi.2010.10.008. [DOI] [PubMed] [Google Scholar]
- Zhang Y, Berndt U, Gölz H, Tais A, Oellerer S, Wölfle T, Fitzke E, Rospert S. NAC functions as a modulator of SRP during the early steps of protein targeting to the endoplasmic reticulum. Molecular biology of the cell. 2012;23:3027–3040. doi: 10.1091/mbc.E12-02-0112. [DOI] [PMC free article] [PubMed] [Google Scholar]