Abstract
To isolate Saccharomyces cerevisiae mutants defective in recombinational DNA repair, we constructed a strain that contains duplicated ura3 alleles that flank LEU2 and ADE5 genes at the ura3 locus on chromosome V. When a HO endonuclease cleavage site is located within one of the ura3 alleles, Ura+ recombination is increased over 100-fold in wild-type strains following HO induction from the GAL1, 10 promoter. This strain was used to screen for mutants that exhibited reduced levels of HO-induced intrachromosomal recombination without significantly affecting the spontaneous frequency of Ura+ recombination. One of the mutations isolated through this screen was found to affect the essential gene CDC1. This mutation, cdc1-100, completely eliminated HO-induced Ura+ recombination yet maintained both spontaneous preinduced recombination levels and cell viability, cdc1-100 mutants were moderately sensitive to killing by methyl methanesulfonate and gamma irradiation. The effect of the cdc1-100 mutation on recombinational double-strand break repair indicates that a recombinationally silent mechanism other than sister chromatid exchange was responsible for the efficient repair of DNA double-strand breaks.
Full text
PDF





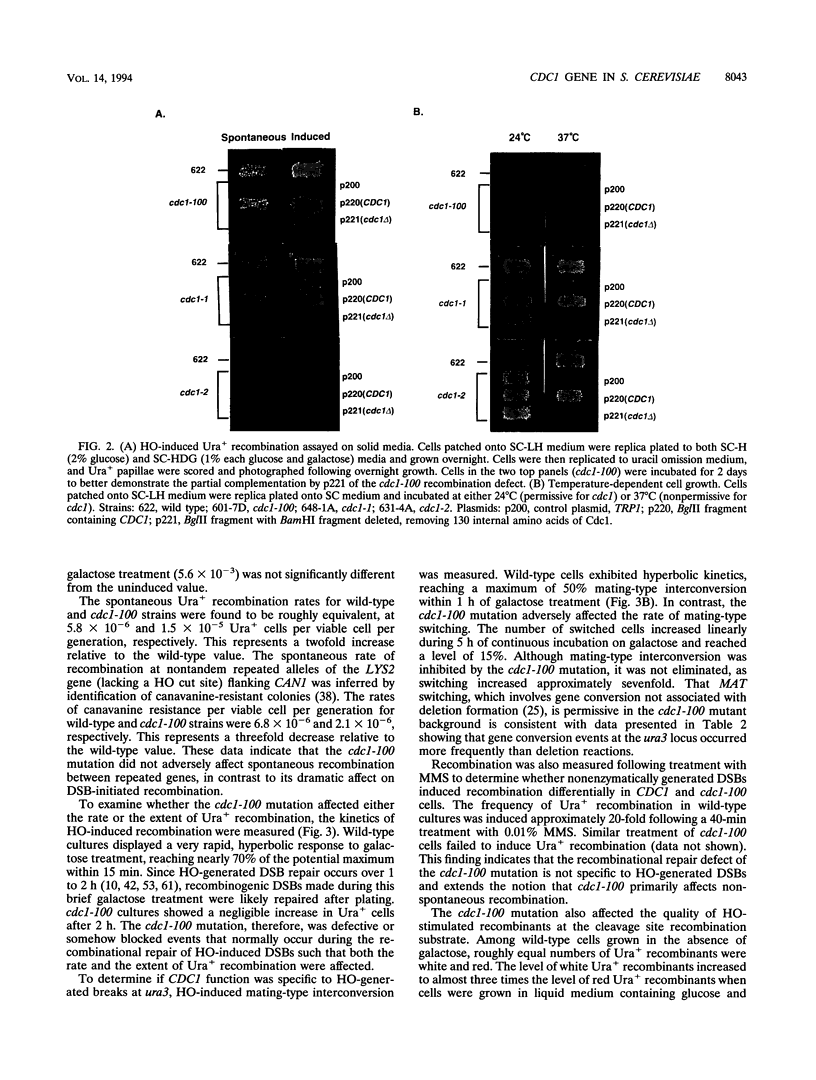







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aguilera A., Klein H. L. Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics. 1988 Aug;119(4):779–790. doi: 10.1093/genetics/119.4.779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Brunborg G., Resnick M. A., Williamson D. H. Cell-cycle-specific repair of DNA double strand breaks in Saccharomyces cerevisiae. Radiat Res. 1980 Jun;82(3):547–558. [PubMed] [Google Scholar]
- Brunborg G., Williamson D. H. The relevance of the nuclear division cycle to radiosensitivity in yeast. Mol Gen Genet. 1978 Jul 4;162(3):277–286. doi: 10.1007/BF00268853. [DOI] [PubMed] [Google Scholar]
- Byers B., Goetsch L. Duplication of spindle plaques and integration of the yeast cell cycle. Cold Spring Harb Symp Quant Biol. 1974;38:123–131. doi: 10.1101/sqb.1974.038.01.016. [DOI] [PubMed] [Google Scholar]
- Chen J., Derfler B., Samson L. Saccharomyces cerevisiae 3-methyladenine DNA glycosylase has homology to the AlkA glycosylase of E. coli and is induced in response to DNA alkylation damage. EMBO J. 1990 Dec;9(13):4569–4575. doi: 10.1002/j.1460-2075.1990.tb07910.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chlebowicz E., Jachymczyk W. J. Repair of MMS-induced DNA double-strand breaks in haploid cells of Saccharomyces cerevisiae, which requires the presence of a duplicate genome. Mol Gen Genet. 1979 Jan 2;167(3):279–286. doi: 10.1007/BF00267420. [DOI] [PubMed] [Google Scholar]
- Connolly B., White C. I., Haber J. E. Physical monitoring of mating type switching in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jun;8(6):2342–2349. doi: 10.1128/mcb.8.6.2342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dhillon N., Hoekstra M. F. Characterization of two protein kinases from Schizosaccharomyces pombe involved in the regulation of DNA repair. EMBO J. 1994 Jun 15;13(12):2777–2788. doi: 10.1002/j.1460-2075.1994.tb06571.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elble R. A simple and efficient procedure for transformation of yeasts. Biotechniques. 1992 Jul;13(1):18–20. [PubMed] [Google Scholar]
- Fishman-Lobell J., Rudin N., Haber J. E. Two alternative pathways of double-strand break repair that are kinetically separable and independently modulated. Mol Cell Biol. 1992 Mar;12(3):1292–1303. doi: 10.1128/mcb.12.3.1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Game J. C., Mortimer R. K. A genetic study of x-ray sensitive mutants in yeast. Mutat Res. 1974 Sep;24(3):281–292. doi: 10.1016/0027-5107(74)90176-6. [DOI] [PubMed] [Google Scholar]
- Gietz R. D., Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene. 1988 Dec 30;74(2):527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H., Culotti J., Pringle J. R., Reid B. J. Genetic control of the cell division cycle in yeast. Science. 1974 Jan 11;183(4120):46–51. doi: 10.1126/science.183.4120.46. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H., Culotti J., Reid B. Genetic control of the cell-division cycle in yeast. I. Detection of mutants. Proc Natl Acad Sci U S A. 1970 Jun;66(2):352–359. doi: 10.1073/pnas.66.2.352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartwell L. H. Genetic control of the cell division cycle in yeast. IV. Genes controlling bud emergence and cytokinesis. Exp Cell Res. 1971 Dec;69(2):265–276. doi: 10.1016/0014-4827(71)90223-0. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H., Mortimer R. K., Culotti J., Culotti M. Genetic Control of the Cell Division Cycle in Yeast: V. Genetic Analysis of cdc Mutants. Genetics. 1973 Jun;74(2):267–286. doi: 10.1093/genetics/74.2.267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henner W. D., Grunberg S. M., Haseltine W. A. Sites and structure of gamma radiation-induced DNA strand breaks. J Biol Chem. 1982 Oct 10;257(19):11750–11754. [PubMed] [Google Scholar]
- Ho K. S., Mortimer R. K. Induction of dominant lethality by x-rays in radiosensitive strain of yeast. Mutat Res. 1973 Oct;20(1):45–51. doi: 10.1016/0027-5107(73)90096-1. [DOI] [PubMed] [Google Scholar]
- Ho K. S., Mortimer R. K. Two mutations which confer temperature-sensitive radiation sensitivity in the yeast Saccharomyces cerevisiae. Mutat Res. 1975 Dec;33(2-3):157–164. doi: 10.1016/0027-5107(75)90190-6. [DOI] [PubMed] [Google Scholar]
- Hovland P., Flick J., Johnston M., Sclafani R. A. Galactose as a gratuitous inducer of GAL gene expression in yeasts growing on glucose. Gene. 1989 Nov 15;83(1):57–64. doi: 10.1016/0378-1119(89)90403-4. [DOI] [PubMed] [Google Scholar]
- Huisman O., Raymond W., Froehlich K. U., Errada P., Kleckner N., Botstein D., Hoyt M. A. A Tn10-lacZ-kanR-URA3 gene fusion transposon for insertion mutagenesis and fusion analysis of yeast and bacterial genes. Genetics. 1987 Jun;116(2):191–199. doi: 10.1093/genetics/116.2.191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jensen R. E., Herskowitz I. Directionality and regulation of cassette substitution in yeast. Cold Spring Harb Symp Quant Biol. 1984;49:97–104. doi: 10.1101/sqb.1984.049.01.013. [DOI] [PubMed] [Google Scholar]
- Kadyk L. C., Hartwell L. H. Sister chromatids are preferred over homologs as substrates for recombinational repair in Saccharomyces cerevisiae. Genetics. 1992 Oct;132(2):387–402. doi: 10.1093/genetics/132.2.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kostriken R., Heffron F. The product of the HO gene is a nuclease: purification and characterization of the enzyme. Cold Spring Harb Symp Quant Biol. 1984;49:89–96. doi: 10.1101/sqb.1984.049.01.012. [DOI] [PubMed] [Google Scholar]
- Kramer K. M., Brock J. A., Bloom K., Moore J. K., Haber J. E. Two different types of double-strand breaks in Saccharomyces cerevisiae are repaired by similar RAD52-independent, nonhomologous recombination events. Mol Cell Biol. 1994 Feb;14(2):1293–1301. doi: 10.1128/mcb.14.2.1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunz B. A., Haynes R. H. Phenomenology and genetic control of mitotic recombination in yeast. Annu Rev Genet. 1981;15:57–89. doi: 10.1146/annurev.ge.15.120181.000421. [DOI] [PubMed] [Google Scholar]
- Lin Y. H., Keil R. L. Mutations affecting RNA polymerase I-stimulated exchange and rDNA recombination in yeast. Genetics. 1991 Jan;127(1):31–38. doi: 10.1093/genetics/127.1.31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luchnik A. N., Glaser V. M., Shestakov S. V. Repair of DNA double-strand breaks requires two homologous DNA duplexes. Mol Biol Rep. 1977 Dec;3(6):437–442. doi: 10.1007/BF00808385. [DOI] [PubMed] [Google Scholar]
- Malone R. E., Esposito R. E. The RAD52 gene is required for homothallic interconversion of mating types and spontaneous mitotic recombination in yeast. Proc Natl Acad Sci U S A. 1980 Jan;77(1):503–507. doi: 10.1073/pnas.77.1.503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumoto K., Yoshimatsu T., Oshima Y. Recessive mutations conferring resistance to carbon catabolite repression of galactokinase synthesis in Saccharomyces cerevisiae. J Bacteriol. 1983 Mar;153(3):1405–1414. doi: 10.1128/jb.153.3.1405-1414.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nickoloff J. A., Singer J. D., Hoekstra M. F., Heffron F. Double-strand breaks stimulate alternative mechanisms of recombination repair. J Mol Biol. 1989 Jun 5;207(3):527–541. doi: 10.1016/0022-2836(89)90462-2. [DOI] [PubMed] [Google Scholar]
- Ozenberger B. A., Roeder G. S. A unique pathway of double-strand break repair operates in tandemly repeated genes. Mol Cell Biol. 1991 Mar;11(3):1222–1231. doi: 10.1128/mcb.11.3.1222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray A., Siddiqi I., Kolodkin A. L., Stahl F. W. Intra-chromosomal gene conversion induced by a DNA double-strand break in Saccharomyces cerevisiae. J Mol Biol. 1988 May 20;201(2):247–260. doi: 10.1016/0022-2836(88)90136-2. [DOI] [PubMed] [Google Scholar]
- Repine J. E., Pfenninger O. W., Talmage D. W., Berger E. M., Pettijohn D. E. Dimethyl sulfoxide prevents DNA nicking mediated by ionizing radiation or iron/hydrogen peroxide-generated hydroxyl radical. Proc Natl Acad Sci U S A. 1981 Feb;78(2):1001–1003. doi: 10.1073/pnas.78.2.1001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick M. A. Genetic control of radiation sensitivity in Saccharomyces cerevisiae. Genetics. 1969 Jul;62(3):519–531. doi: 10.1093/genetics/62.3.519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Resnick M. A., Martin P. The repair of double-strand breaks in the nuclear DNA of Saccharomyces cerevisiae and its genetic control. Mol Gen Genet. 1976 Jan 16;143(2):119–129. doi: 10.1007/BF00266917. [DOI] [PubMed] [Google Scholar]
- Resnick M. A. The repair of double-strand breaks in DNA; a model involving recombination. J Theor Biol. 1976 Jun;59(1):97–106. doi: 10.1016/s0022-5193(76)80025-2. [DOI] [PubMed] [Google Scholar]
- Rose M. D., Broach J. R. Cloning genes by complementation in yeast. Methods Enzymol. 1991;194:195–230. doi: 10.1016/0076-6879(91)94017-7. [DOI] [PubMed] [Google Scholar]
- Rothstein R. Targeting, disruption, replacement, and allele rescue: integrative DNA transformation in yeast. Methods Enzymol. 1991;194:281–301. doi: 10.1016/0076-6879(91)94022-5. [DOI] [PubMed] [Google Scholar]
- Rudin N., Haber J. E. Efficient repair of HO-induced chromosomal breaks in Saccharomyces cerevisiae by recombination between flanking homologous sequences. Mol Cell Biol. 1988 Sep;8(9):3918–3928. doi: 10.1128/mcb.8.9.3918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simchen G. Are mitotic functions required in meiosis? Genetics. 1974 Apr;76(4):745–753. doi: 10.1093/genetics/76.4.745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugawara N., Haber J. E. Characterization of double-strand break-induced recombination: homology requirements and single-stranded DNA formation. Mol Cell Biol. 1992 Feb;12(2):563–575. doi: 10.1128/mcb.12.2.563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas B. J., Rothstein R. Elevated recombination rates in transcriptionally active DNA. Cell. 1989 Feb 24;56(4):619–630. doi: 10.1016/0092-8674(89)90584-9. [DOI] [PubMed] [Google Scholar]
- Weiffenbach B., Haber J. E. Homothallic mating type switching generates lethal chromosome breaks in rad52 strains of Saccharomyces cerevisiae. Mol Cell Biol. 1981 Jun;1(6):522–534. doi: 10.1128/mcb.1.6.522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinert T. A., Hartwell L. H. The RAD9 gene controls the cell cycle response to DNA damage in Saccharomyces cerevisiae. Science. 1988 Jul 15;241(4863):317–322. doi: 10.1126/science.3291120. [DOI] [PubMed] [Google Scholar]
- White C. I., Haber J. E. Intermediates of recombination during mating type switching in Saccharomyces cerevisiae. EMBO J. 1990 Mar;9(3):663–673. doi: 10.1002/j.1460-2075.1990.tb08158.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao W., Derfler B., Chen J., Samson L. Primary sequence and biological functions of a Saccharomyces cerevisiae O6-methylguanine/O4-methylthymine DNA repair methyltransferase gene. EMBO J. 1991 Aug;10(8):2179–2186. doi: 10.1002/j.1460-2075.1991.tb07753.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zehfus B. R., McWilliams A. D., Lin Y. H., Hoekstra M. F., Keil R. L. Genetic control of RNA polymerase I-stimulated recombination in yeast. Genetics. 1990 Sep;126(1):41–52. doi: 10.1093/genetics/126.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]




