Abstract
To maintain genomic integrity, double-strand breaks (DSBs) in chromosomal DNA must be repaired. In mammalian systems, the analysis of the repair of chromosomal DSBs has been limited by the inability to introduce well-defined DSBs in genomic DNA. In this study, we created specific DSBs in mouse chromosomes for the first time, using an expression system for a rare-cutting endonuclease, I-SceI. A genetic assay has been devised to monitor the repair of DSBs, whereby cleavage sites for I-SceI have been integrated into the mouse genome in two tandem neomycin phosphotransferase genes. We find that cleavage of the I-SceI sites is very efficient, with at least 12% of stably transfected cells having at least one cleavage event and, of these, more than 70% have undergone cleavage at both I-SceI sites. Cleavage of both sites in a fraction of clones deletes 3.8 kb of intervening chromosomal sequences. We find that the DSBs are repaired by both homologous and nonhomologous mechanisms. Nonhomologous repair events frequently result in small deletions after rejoining of the two DNA ends. Some of these appear to occur by simple blunt-ended ligation, whereas several others may occur through annealing of short regions of terminal homology. The DSBs are apparently recombinogenic, stimulating gene targeting of a homologous fragment by more than 2 orders of magnitude. Whereas gene-targeted clones are nearly undetectable without endonuclease expression, they represent approximately 10% of cells transfected with the I-SceI expression vector. Gene targeted clones are of two major types, those that occur by two-sided homologous recombination with the homologous fragment and those that occur by one-sided homologous recombination. Our results are expected to impact a number of areas in the study of mammalian genome dynamics, including the analysis of the repair of DSBs and homologous recombination and, potentially, molecular genetic analyses of mammalian genomes.
Full text
PDF



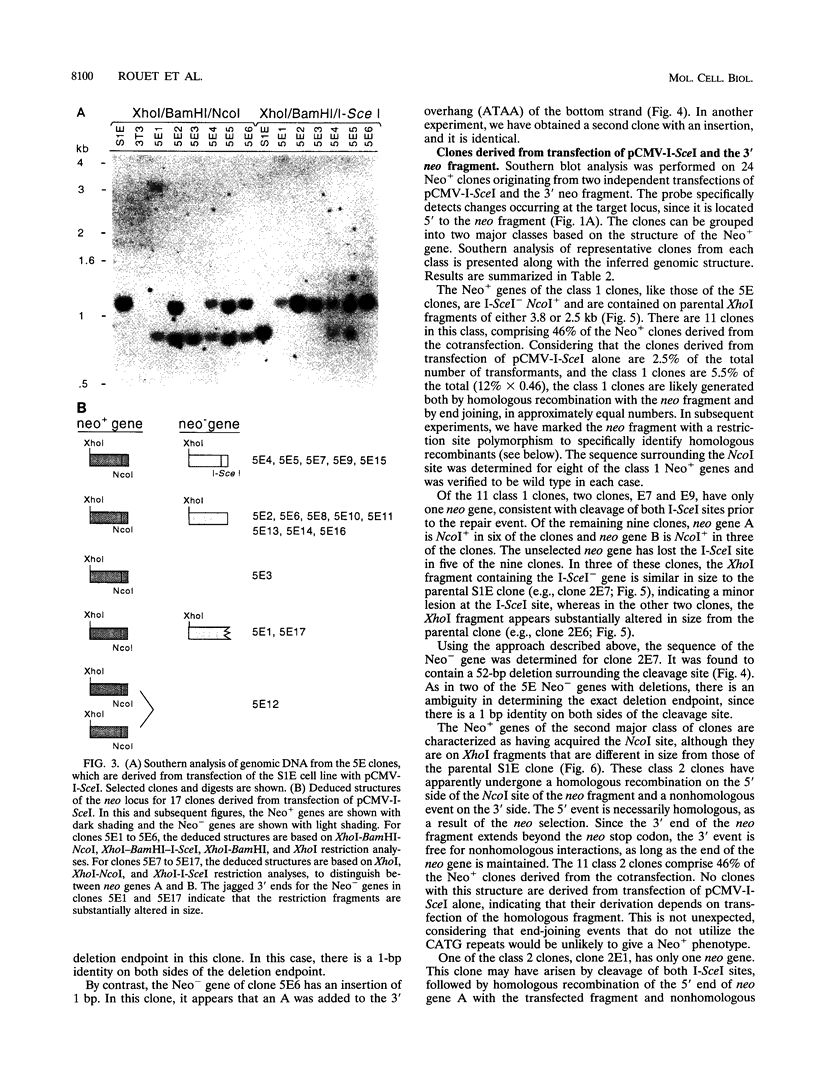






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Andersson S., Davis D. L., Dahlbäck H., Jörnvall H., Russell D. W. Cloning, structure, and expression of the mitochondrial cytochrome P-450 sterol 26-hydroxylase, a bile acid biosynthetic enzyme. J Biol Chem. 1989 May 15;264(14):8222–8229. [PubMed] [Google Scholar]
- Barnett M. A., Buckle V. J., Evans E. P., Porter A. C., Rout D., Smith A. G., Brown W. R. Telomere directed fragmentation of mammalian chromosomes. Nucleic Acids Res. 1993 Jan 11;21(1):27–36. doi: 10.1093/nar/21.1.27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berinstein N., Pennell N., Ottaway C. A., Shulman M. J. Gene replacement with one-sided homologous recombination. Mol Cell Biol. 1992 Jan;12(1):360–367. doi: 10.1128/mcb.12.1.360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cao L., Alani E., Kleckner N. A pathway for generation and processing of double-strand breaks during meiotic recombination in S. cerevisiae. Cell. 1990 Jun 15;61(6):1089–1101. doi: 10.1016/0092-8674(90)90072-m. [DOI] [PubMed] [Google Scholar]
- Capecchi M. R. Altering the genome by homologous recombination. Science. 1989 Jun 16;244(4910):1288–1292. doi: 10.1126/science.2660260. [DOI] [PubMed] [Google Scholar]
- Chen C., Okayama H. High-efficiency transformation of mammalian cells by plasmid DNA. Mol Cell Biol. 1987 Aug;7(8):2745–2752. doi: 10.1128/mcb.7.8.2745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clark J. M. Novel non-templated nucleotide addition reactions catalyzed by procaryotic and eucaryotic DNA polymerases. Nucleic Acids Res. 1988 Oct 25;16(20):9677–9686. doi: 10.1093/nar/16.20.9677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Colleaux L., D'Auriol L., Galibert F., Dujon B. Recognition and cleavage site of the intron-encoded omega transposase. Proc Natl Acad Sci U S A. 1988 Aug;85(16):6022–6026. doi: 10.1073/pnas.85.16.6022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dujon B. Group I introns as mobile genetic elements: facts and mechanistic speculations--a review. Gene. 1989 Oct 15;82(1):91–114. doi: 10.1016/0378-1119(89)90034-6. [DOI] [PubMed] [Google Scholar]
- Ellis J., Bernstein A. Gene targeting with retroviral vectors: recombination by gene conversion into regions of nonhomology. Mol Cell Biol. 1989 Apr;9(4):1621–1627. doi: 10.1128/mcb.9.4.1621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farr C. J., Stevanovic M., Thomson E. J., Goodfellow P. N., Cooke H. J. Telomere-associated chromosome fragmentation: applications in genome manipulation and analysis. Nat Genet. 1992 Dec;2(4):275–282. doi: 10.1038/ng1292-275. [DOI] [PubMed] [Google Scholar]
- Fishman-Lobell J., Rudin N., Haber J. E. Two alternative pathways of double-strand break repair that are kinetically separable and independently modulated. Mol Cell Biol. 1992 Mar;12(3):1292–1303. doi: 10.1128/mcb.12.3.1292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gloor G. B., Nassif N. A., Johnson-Schlitz D. M., Preston C. R., Engels W. R. Targeted gene replacement in Drosophila via P element-induced gap repair. Science. 1991 Sep 6;253(5024):1110–1117. doi: 10.1126/science.1653452. [DOI] [PubMed] [Google Scholar]
- Goldway M., Sherman A., Zenvirth D., Arbel T., Simchen G. A short chromosomal region with major roles in yeast chromosome III meiotic disjunction, recombination and double strand breaks. Genetics. 1993 Feb;133(2):159–169. doi: 10.1093/genetics/133.2.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haber J. E. Exploring the pathways of homologous recombination. Curr Opin Cell Biol. 1992 Jun;4(3):401–412. doi: 10.1016/0955-0674(92)90005-w. [DOI] [PubMed] [Google Scholar]
- Hasty P., Rivera-Pérez J., Bradley A. The role and fate of DNA ends for homologous recombination in embryonic stem cells. Mol Cell Biol. 1992 Jun;12(6):2464–2474. doi: 10.1128/mcb.12.6.2464. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Itzhaki J. E., Barnett M. A., MacCarthy A. B., Buckle V. J., Brown W. R., Porter A. C. Targeted breakage of a human chromosome mediated by cloned human telomeric DNA. Nat Genet. 1992 Dec;2(4):283–287. doi: 10.1038/ng1292-283. [DOI] [PubMed] [Google Scholar]
- Jasin M., Berg P. Homologous integration in mammalian cells without target gene selection. Genes Dev. 1988 Nov;2(11):1353–1363. doi: 10.1101/gad.2.11.1353. [DOI] [PubMed] [Google Scholar]
- Jasin M., Elledge S. J., Davis R. W., Berg P. Gene targeting at the human CD4 locus by epitope addition. Genes Dev. 1990 Feb;4(2):157–166. doi: 10.1101/gad.4.2.157. [DOI] [PubMed] [Google Scholar]
- Jasin M., Liang F. Mouse embryonic stem cells exhibit high levels of extrachromosomal homologous recombination in a chloramphenicol acetyltransferase assay system. Nucleic Acids Res. 1991 Dec;19(25):7171–7175. doi: 10.1093/nar/19.25.7171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jasin M., de Villiers J., Weber F., Schaffner W. High frequency of homologous recombination in mammalian cells between endogenous and introduced SV40 genomes. Cell. 1985 Dec;43(3 Pt 2):695–703. doi: 10.1016/0092-8674(85)90242-9. [DOI] [PubMed] [Google Scholar]
- Kolodkin A. L., Klar A. J., Stahl F. W. Double-strand breaks can initiate meiotic recombination in S. cerevisiae. Cell. 1986 Aug 29;46(5):733–740. doi: 10.1016/0092-8674(86)90349-1. [DOI] [PubMed] [Google Scholar]
- Kostriken R., Heffron F. The product of the HO gene is a nuclease: purification and characterization of the enzyme. Cold Spring Harb Symp Quant Biol. 1984;49:89–96. doi: 10.1101/sqb.1984.049.01.012. [DOI] [PubMed] [Google Scholar]
- Kramer K. M., Brock J. A., Bloom K., Moore J. K., Haber J. E. Two different types of double-strand breaks in Saccharomyces cerevisiae are repaired by similar RAD52-independent, nonhomologous recombination events. Mol Cell Biol. 1994 Feb;14(2):1293–1301. doi: 10.1128/mcb.14.2.1293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuerbitz S. J., Plunkett B. S., Walsh W. V., Kastan M. B. Wild-type p53 is a cell cycle checkpoint determinant following irradiation. Proc Natl Acad Sci U S A. 1992 Aug 15;89(16):7491–7495. doi: 10.1073/pnas.89.16.7491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lewis S. M. The mechanism of V(D)J joining: lessons from molecular, immunological, and comparative analyses. Adv Immunol. 1994;56:27–150. doi: 10.1016/s0065-2776(08)60450-2. [DOI] [PubMed] [Google Scholar]
- Lin F. L., Sperle K., Sternberg N. Intermolecular recombination between DNAs introduced into mouse L cells is mediated by a nonconservative pathway that leads to crossover products. Mol Cell Biol. 1990 Jan;10(1):103–112. doi: 10.1128/mcb.10.1.103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin F. L., Sperle K., Sternberg N. Model for homologous recombination during transfer of DNA into mouse L cells: role for DNA ends in the recombination process. Mol Cell Biol. 1984 Jun;4(6):1020–1034. doi: 10.1128/mcb.4.6.1020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin F. L., Sperle K., Sternberg N. Repair of double-stranded DNA breaks by homologous DNA fragments during transfer of DNA into mouse L cells. Mol Cell Biol. 1990 Jan;10(1):113–119. doi: 10.1128/mcb.10.1.113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maryon E., Carroll D. Degradation of linear DNA by a strand-specific exonuclease activity in Xenopus laevis oocytes. Mol Cell Biol. 1989 Nov;9(11):4862–4871. doi: 10.1128/mcb.9.11.4862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nickoloff J. A., Chen E. Y., Heffron F. A 24-base-pair DNA sequence from the MAT locus stimulates intergenic recombination in yeast. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7831–7835. doi: 10.1073/pnas.83.20.7831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nickoloff J. A., Singer J. D., Hoekstra M. F., Heffron F. Double-strand breaks stimulate alternative mechanisms of recombination repair. J Mol Biol. 1989 Jun 5;207(3):527–541. doi: 10.1016/0022-2836(89)90462-2. [DOI] [PubMed] [Google Scholar]
- Nicolás A. L., Young C. S. Characterization of DNA end joining in a mammalian cell nuclear extract: junction formation is accompanied by nucleotide loss, which is limited and uniform but not site specific. Mol Cell Biol. 1994 Jan;14(1):170–180. doi: 10.1128/mcb.14.1.170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ozenberger B. A., Roeder G. S. A unique pathway of double-strand break repair operates in tandemly repeated genes. Mol Cell Biol. 1991 Mar;11(3):1222–1231. doi: 10.1128/mcb.11.3.1222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perrin A., Buckle M., Dujon B. Asymmetrical recognition and activity of the I-SceI endonuclease on its site and on intron-exon junctions. EMBO J. 1993 Jul;12(7):2939–2947. doi: 10.1002/j.1460-2075.1993.tb05956.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfeiffer P., Thode S., Hancke J., Vielmetter W. Mechanisms of overlap formation in nonhomologous DNA end joining. Mol Cell Biol. 1994 Feb;14(2):888–895. doi: 10.1128/mcb.14.2.888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plasterk R. H. The origin of footprints of the Tc1 transposon of Caenorhabditis elegans. EMBO J. 1991 Jul;10(7):1919–1925. doi: 10.1002/j.1460-2075.1991.tb07718.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plessis A., Perrin A., Haber J. E., Dujon B. Site-specific recombination determined by I-SceI, a mitochondrial group I intron-encoded endonuclease expressed in the yeast nucleus. Genetics. 1992 Mar;130(3):451–460. doi: 10.1093/genetics/130.3.451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puchta H., Dujon B., Hohn B. Homologous recombination in plant cells is enhanced by in vivo induction of double strand breaks into DNA by a site-specific endonuclease. Nucleic Acids Res. 1993 Nov 11;21(22):5034–5040. doi: 10.1093/nar/21.22.5034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray A., Siddiqi I., Kolodkin A. L., Stahl F. W. Intra-chromosomal gene conversion induced by a DNA double-strand break in Saccharomyces cerevisiae. J Mol Biol. 1988 May 20;201(2):247–260. doi: 10.1016/0022-2836(88)90136-2. [DOI] [PubMed] [Google Scholar]
- Rouet P., Smih F., Jasin M. Expression of a site-specific endonuclease stimulates homologous recombination in mammalian cells. Proc Natl Acad Sci U S A. 1994 Jun 21;91(13):6064–6068. doi: 10.1073/pnas.91.13.6064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudin N., Haber J. E. Efficient repair of HO-induced chromosomal breaks in Saccharomyces cerevisiae by recombination between flanking homologous sequences. Mol Cell Biol. 1988 Sep;8(9):3918–3928. doi: 10.1128/mcb.8.9.3918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rudin N., Sugarman E., Haber J. E. Genetic and physical analysis of double-strand break repair and recombination in Saccharomyces cerevisiae. Genetics. 1989 Jul;122(3):519–534. doi: 10.1093/genetics/122.3.519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sandell L. L., Zakian V. A. Loss of a yeast telomere: arrest, recovery, and chromosome loss. Cell. 1993 Nov 19;75(4):729–739. doi: 10.1016/0092-8674(93)90493-a. [DOI] [PubMed] [Google Scholar]
- Southern P. J., Berg P. Transformation of mammalian cells to antibiotic resistance with a bacterial gene under control of the SV40 early region promoter. J Mol Appl Genet. 1982;1(4):327–341. [PubMed] [Google Scholar]
- Strathern J. N., Klar A. J., Hicks J. B., Abraham J. A., Ivy J. M., Nasmyth K. A., McGill C. Homothallic switching of yeast mating type cassettes is initiated by a double-stranded cut in the MAT locus. Cell. 1982 Nov;31(1):183–192. doi: 10.1016/0092-8674(82)90418-4. [DOI] [PubMed] [Google Scholar]
- Sugawara N., Haber J. E. Characterization of double-strand break-induced recombination: homology requirements and single-stranded DNA formation. Mol Cell Biol. 1992 Feb;12(2):563–575. doi: 10.1128/mcb.12.2.563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun H., Treco D., Schultes N. P., Szostak J. W. Double-strand breaks at an initiation site for meiotic gene conversion. Nature. 1989 Mar 2;338(6210):87–90. doi: 10.1038/338087a0. [DOI] [PubMed] [Google Scholar]
- Szostak J. W., Orr-Weaver T. L., Rothstein R. J., Stahl F. W. The double-strand-break repair model for recombination. Cell. 1983 May;33(1):25–35. doi: 10.1016/0092-8674(83)90331-8. [DOI] [PubMed] [Google Scholar]
- Thomas K. R., Capecchi M. R. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell. 1987 Nov 6;51(3):503–512. doi: 10.1016/0092-8674(87)90646-5. [DOI] [PubMed] [Google Scholar]
- Valancius V., Smithies O. Double-strand gap repair in a mammalian gene targeting reaction. Mol Cell Biol. 1991 Sep;11(9):4389–4397. doi: 10.1128/mcb.11.9.4389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H., Liu X., Jaenisch R. Double replacement: strategy for efficient introduction of subtle mutations into the murine Col1a-1 gene by homologous recombination in embryonic stem cells. Proc Natl Acad Sci U S A. 1994 Mar 29;91(7):2819–2823. doi: 10.1073/pnas.91.7.2819. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu T. C., Lichten M. Meiosis-induced double-strand break sites determined by yeast chromatin structure. Science. 1994 Jan 28;263(5146):515–518. doi: 10.1126/science.8290959. [DOI] [PubMed] [Google Scholar]
- Zheng H., Wilson J. H. Gene targeting in normal and amplified cell lines. Nature. 1990 Mar 8;344(6262):170–173. doi: 10.1038/344170a0. [DOI] [PubMed] [Google Scholar]
- te Riele H., Maandag E. R., Clarke A., Hooper M., Berns A. Consecutive inactivation of both alleles of the pim-1 proto-oncogene by homologous recombination in embryonic stem cells. Nature. 1990 Dec 13;348(6302):649–651. doi: 10.1038/348649a0. [DOI] [PubMed] [Google Scholar]