Abstract
Acetylcholinesterase (AChE) in the serum of fetal cow is a tetramer. The related enzyme, butyrylcholinesterase (BChE), in the sera of humans and horse requires polyproline peptides for assembly into tetramers. Our goal was to determine whether soluble tetrameric AChE includes tetramer organizing peptides in its structure. Fetal bovine serum AChE was denatured by boiling to release non-covalently bound peptides. Bulk protein was separated from peptides by filtration and by high performance liquid chromatography. Peptide mass and amino acid sequence of the released peptides were determined by MALDI-TOF-TOF and LTQ-Orbitrap mass spectrometry. Twenty polyproline peptides, divided into 5 families, were identified. The longest peptide contained 25 consecutive prolines and no other amino acid. Other polyproline peptides included one non-proline amino acid, for example serine at the C-terminus of 20 prolines. A search of the mammalian proteome database suggested that this assortment of polyproline peptides originated from at least 5 different precursor proteins, none of which were the ColQ or PRiMA of membrane-anchored AChE. To date, AChE and BChE are the only proteins known that include polyproline tetramer organizing peptides in their tetrameric structure.
Keywords: fetal bovine serum acetylcholinesterase, polyproline peptide, mass spectrometry, PAGE gel electrophoresis, tetramer organization
1. Introduction
Vertebrate acetylcholinesterase (AChE; EC 3.1.1.7) belongs to the serine-esterase protein family [1]. AChE terminates the action of acetylcholine at the post-synaptic membrane in the neuromuscular junction. AChE is also found in a variety of other tissues where its function is unclear. For example, AChE is abundant in fetal bovine serum, but not in the serum of the adult cow where its concentration is only 10% of that in fetal serum [2]. In the neuromuscular junction, tetrameric forms of AChE are anchored to the basal lamina via a collagenous subunit referred to as ColQ [3]. In the brain, tetrameric forms of AChE are bound to membranes via a proline-rich membrane anchor (PRiMA) [4]. Both ColQ and PRiMA contain a proline-rich domain (PRAD) that is credited with organizing AChE monomers into tetramers. The crystal structure of the 40-residue AChE tetramerization domain in complex with the 15-residue PRAD peptide shows four highly conserved tryptophan amphiphilic α-helices from the C-terminus of AChE wrapped around an antiparallel left-handed PRAD helix resembling polyproline II [5]. The essential feature of PRAD is the presence of long stretches of consecutive proline residues. Even synthetic polyproline can function to organize AChE monomers into tetramers [6].
AChE can also be found in soluble form. Co-expression of AChE with the proline-rich fragment from ColQ promotes secretion of stable AChE tetramers in cell culture. In the absence of the proline-rich fragment, only monomers and dimers are secreted, indicating that the proline-rich fragment organizes assembly of the soluble tetramer [7]. It was found that soluble, tetrameric butyrylcholinesterase in human serum is stabilized by a proline-rich peptides derived from the lamellipodin protein [8] in addition to several other proteins. This strongly suggests that the tetramer-organizing peptides used for soluble cholinesterases are different from those employed by membrane-bound forms, but also suggests that they share a common polyproline motif. However, the identity of the proline-rich peptides present in naturally-occurring, soluble AChE has not been determined.
Fetal bovine serum (FBS) is a particularly rich source of soluble AChE. FBS AChE is a symmetric tetrameric globule with a molecular mass of 340,000 Da. The four subunits are associated in a dimer-of-dimers motif. That is, two pairs of disulfide crosslinked monomers are held together by hydrophobic interactions [9]. Based on the foregoing discussion it is to be expected that the tetrameric assembly of FBS AChE is organized around a proline-rich peptide.
In the present study we extracted and characterized the polyproline peptides of FBS AChE. The protein was denatured by boiling to release the peptides. Released peptides, separated from AChE subunits by HPLC or by filtration, were analyzed by mass spectrometry. We identified polyproline-rich peptides none of which matched the PRAD sequence from either ColQ or PRiMA.
2. Methods and Materials
2.1 Materials
AChE from fetal bovine serum was purified from serum pooled from multiple animals as described [10] and stored at −80°C in 50% glycerol until use. The FBS AChE was 99% pure as determined by Coomassie blue staining of an SDS gel and confirmed by measurement of specific activity. α-Cyano-4-hydroxy cinnamic acid (α-CHCA, catalog number 70990) from Fluka (a member of the Sigma-Aldrich group, St. Louis, MO) was prepared as a saturated solution (10 mg/ml) in 50% acetonitrile/water plus 0.3% trifluoroacetic acid. Acetonitrile was of DNA sequencing grade from Fisher (Pittsburgh, PA, catalog number BP-1170), and trifluoroacetic acid was of sequencing grade (>99.9%) from Beckman (Brea, CA, catalog number 290204). 5,5′-dithiobis-(2-nitrobenzoic acid) was from Sigma-Aldrich (catalog number D8130). Formic acid (puriss p.a. for mass spectrometry, catalog number 94318) was from Sigma-Aldrich. All other chemicals were of biochemical grade.
2.2 AChE activity
Activity was assayed with 1 mM acetylthiocholine in 0.1 M potassium phosphate buffer (pH 7.0) at 25°C, in the presence of 0.5 mM 5,5′-dithiobis-(2-nitrobenzoic acid) [11]. One unit of activity is defined as one micromole of acetylthiocholine hydrolyzed per minute. The specific activity of 99% pure FBS AChE at pH 7.0, 25°C is 2160 units/mg where protein concentration is calculated from absorbance at 280 nm. The same preparation has a specific activity of 3000–3300 units/mg when activity is assayed at pH 8.0 [9].
2.3 HPLC purification of tetramer organizing peptides
In preparing a sample for HPLC purification of the organizing peptides, the buffer for 14.1 ml of FBS AChE (350 U/ml) was changed from 50 mM sodium phosphate pH 8.0 containing 50% glycerol to 10 mM ammonium bicarbonate pH 8.1 plus 0.01% sodium azide by concentrating the sample to 1 ml in a 10-ml, Amicon stirred-cell (Millipore, Billerica, MA, catalog # 5121) using a YM 30 membrane (Millipore, catalog #13612) and then diluting it to 9 ml. This was repeated 3-times. Finally, the sample was concentrated to 1.4 ml (A280 = 2.4 with a measured activity of 2800 U/ml). Assuming 100% purity and using an A280 value of 1.1 for 1 mg/ml FBS AChE, a protein concentration of 2.2 mg/ml could be calculated for this preparation.
The polyproline peptides were released from the FBS AChE preparation (1.4 ml at 1.3 mg AChE/ml in 10 mM ammonium bicarbonate pH 8.1 plus 0.01% azide) by heating the AChE in a boiling water bath for 5 min. The boiled AChE was acidified to pH 2 with trifluoroacetic acid, filtered through a 0.2 micron syringe filter, and injected onto a Zorbax 300 SB C-18 reverse-phase column (Agilent Technologies, Santa Clara, CA, U.S.A.) attached to a Waters 625 HPLC system (Milford, MA, U.S.A). The HPLC was operated at room temperature (22° C) at a flow rate of 0.5 ml/min. Buffer A was 0.1 % trifluoroacetic acid in water; buffer B was 0.09 % trifluoroacetic acid in acetonitrile. Peptides were eluted with a gradient of 0–60 % buffer B in 60 min. The absorbance was monitored at 210 nm. The HPLC eluent was collected into 1 min fractions (0.5 ml each). Each fraction was concentrated to approximately 50 μl in a vacuum centrifuge (Savant SpeedVac), and saved at 4°C for mass spectrometry analysis.
2.4 Filtration to separate tetramer organizing peptides from FBS AChE
In preparing a sample for direct application of the organizing peptides onto the MALDI target plate or for Orbitrap analysis, the buffer for 1.5 ml of FBS AChE (2070 U/ml) was replaced with water using an Amicon stirred cell as described above (process was repeated 4-times). The sample volume was reduced to 0.4 ml. The sample was boiled for 5 min to release the polyproline peptides, and then filtered through a Microcon ultracel YM 10 spin filter (MW cutoff 10,000 Da) to separate the released peptides from residual protein. The volume of the filtrate was concentrated to 0.1 ml making the final concentration of peptides equivalent to that expected from a 25 mg/ml sample of AChE.
2.5 MALDI TOF-TOF mass spectrometry
All MALDI TOF-TOF experiments were performed on an Applied Biosystems MALDI TOF-TOF 4800 mass spectrometer equipped with a 355 nm Nd-YAG laser (Applied Biosystems, Framingham, MA). The samples (0.5 μl) were spotted on a MALDI target plate (Opti-TOF 384 well Insert from Applied Biosystems), air-dried and overlaid with 0.5 μl of α-cyano-4-hydroxy cinnamic acid matrix. Mass spectra were acquired in positive ion reflector mode, under delayed extraction conditions (500 ns), using an acceleration voltage of 20 kV, with laser intensity of 4000–6000 V, a mass range of 1000–4000 Da, a detector voltage multiplier of 0.75, with low mass gate on and low mass gate offset equal to zero. Mass spectra shown are the average of 500 laser shots collected from randomly selected locations on the target spot (50 pulses per location). Selected ions were fragmented by collision-induced dissociation, using a 1 kV method, with air as the collision gas at 2×10−6 Torr, a precursor mass window of ± 1 Da, metastable suppression on, a detector voltage multiplier of 0.95, using factory calculated delayed extraction values (DE1 = 370 ns, DE2 = 37,271 ns), and with timed ion selector activated. The identity of the fragments in the MSMS spectra were assigned manually using the Data Explorer software (version 4.9 from Applied Biosystems) with the aid of the Proteomics Toolkit (http://db.systemsbiology.net) and the MS Product algorithm in Protein Prospector v 5.9.4 from the University of California, San Francisco mass spectrometry facility (prospector.ucsf.edu/prospector/mshome.htm).
2.6 LC/MSMS Orbitrap mass spectrometry
The sample was prepared for the Orbitrap by diluting a 3.6 μl aliquot of the Microcon YM 10 filtrate from boiled FBS AChE (made as described above: 25 mg/ml in water) with 6.4 μl of 0.1% formic acid to make a 116 μM solution of peptides.
LC/MSMS analysis was performed on an LTQ Orbitrap quadrupole mass spectrometer (Thermo Scientific a part of Thermo Fisher Scientific, Rockland, IL) using electrospray ionization. Five microliters of the sample were loaded onto a C18 reverse phase trap column (CapTrap Peptide from Michrom BioResources, Auburn, CA, catalog # TRI/25109/32) and washed with 2% acetonitrile/water plus 0.1% formic acid, on an Eksigent nanoLC 2D HPLC system. Then, the peptides were transferred to a C18 reverse phase separation column (Picofrit BioBasics C-18 from New Objective, Woburn, MA, catalog # PF 7515-100H052) and eluted with a 45 min linear gradient starting with 98% solvent A (2% acetonitrile/water plus 0.1% formic acid) and 2% solvent B (98% acetonitrile/water plus 0.1% formic acid) and ending with 35% solvent A and 65% solvent B. The flow rate was 250 nl/min and the effluent was electrosprayed directly into the mass spectrometer. Data were collected in a data dependent manner with each cycle consisting of one high-resolution mass spectrum (over a 300 to 2000 Da mass range) taken with the Orbitrap and five MSMS fragmentation spectra taken with the LTQ ion trap. Collision-induced dissociation was used for fragmentation with helium as the collision gas (at 1×10−3 Torr) and a normalized collision energy of 35 (or about 35% of the maximum collision energy of 5 volts). The activation time was 30 msec, and the activation Q was 0.25. Parent ions of any charge state with a minimum signal intensity of 50,000 counts/s were accepted for fragmentation. After two data collections for a given mass were completed, that mass was excluded from further analysis for 60 s. Fragmentation spectra were analyzed manually using the Qual Browser feature of Xcalibur software (v 4.9 from Thermo Scientific).
3. Results
3.1 MS analysis
The peptides released from heat-denatured FBS AChE are proline-rich. We found that freezing in the absence of glycerol also caused release of proline-rich peptides from FBS AChE. Figure 1 is a MALDI TOF mass spectrum showing the masses of the released peptides. They range in mass from 1037 to 2764 Da. The three most prominent peaks (1949.9, 1971.9 and 1993.9 Da) represent the same peptide in 1) the protonated form, 2) the single-sodium adduct form and 3) the double-sodium adduct form, respectively. All major peptides from this sample were subjected to collision-induced fragmentation in the MALDI mass spectrometer. Qualitatively, the same information was obtained from the samples before and after HPLC purification, however better quality fragmentation spectra were obtained from the HPLC purified peptides.
Figure 1.
MALDI mass spectrum of the peptides released from FBS AChE by boiling. Peptides were separated from bulk protein by filtration through a YM10 membrane (10,000 MW cutoff). The right-hand y-axis indicates the signal intensity in counts-per-second.
For the peptides with the more intense signals, collision-induced dissociation in either the MALDI or Orbitrap mass spectrometers provided enough fragmentation information to determine their amino acid sequences. These peptides are listed in Table 1. Fragmentation series of the y-, b- and a-ion types were observed. In most cases, the peptides observed with the MALDI mass spectrometer were in complex with a single sodium atom (sodium replacing the added proton which makes the peptide positively charged) while those in the Orbitrap were sodium free.
Table 1. Summary of the peptides released from FBS AChE.
Peptides were extracted from serum acetylcholinesterase by simple denaturation and identified by mass spectrometry. Sequences were assigned by manual evaluation of the mass spectral fragmentation patterns.
| Sequence a | # residues | Series Type b | Instrument | Mass, Da |
|---|---|---|---|---|
| Family 1 | ||||
| (PPP)PP PPPPP P(PP) | 13 | y-, b-ion | Orbitrap | 1280.7 |
| (PPPP)P PPPPP PPPP | 14 | b-ion | Orbitrap | 1377.7 |
| (PPPPP P) PPPP PPP(PP) | 15 | y-ion | Orbitrap | 1474.8 |
| (PPPPP PP)PPP (PP)P(PP P) | 16 | y-, b-, a-ion | Orbitrap | 1571.0 |
| (PPPPP) PPPPP PPPPP PP | 17 | b-, a-ion | Orbitrap | 1668.9 |
| PPPPP PPPPP PPP(PP PPP) Na+ | 18 | y-, b-, a-ion | MALDI | 1788.3 |
| PPPPP PPPPP PP(PPP PPPP) Na+ | 19 | y-, b-, a-ion | MALDI | 1885.0 |
| (PP)PPP P(PP)PP P(PPPP PPPPP PPPP) | 24 | y-ion | MALDI | 2348.3 |
| (PP)PPP PPPPP PPPPP (PPPPP PPPPP) | 25 | y-ion | MALDI | 2445.2 |
| Family 2 | ||||
| (APPP)P PPPPP PP(PP) | 14 | y-, b-ion | Orbitrap | 1351.7 |
| (APPPP P)PPPP (PP)(PPP P) | 16 | y-ion | Orbitrap | 1545.8 |
| Family 3 | ||||
| (L/I/Hyp PPPP) PPPPP P(PPP)(PP) | 16 | y-ion | Orbitrap | 1587.9 |
| Family 4 | ||||
| PPPQ/KP PPPPP P(PPPP PP) Na+ | 17 | y-, b-ion | MALDI Orbitrap | 1721.9 |
| Family 5 | ||||
| PPPPP PPPPP PP(PPP PPPPP S) Na+ | 21 | y-, a-ion | MALDI | 2069.1 |
Peptide sequences are written with the N-terminus on the left and the C-terminus on the right. Residues enclosed in brackets indicate that separate signals for the masses of each residue were not resolved in the mass spectrum. However, masses of the bracketed fragments exactly fit the indicated sequence. Leucine, isoleucine, and hydroxyproline (L/I/Hyp) cannot be distinguished because they have the same mass. Glutamine and lysine (Q/K) cannot be distinguished because they have the same mass.
The “Series Type” column lists the major fragment series observed in the MSMS spectrum of each peptide. Peptide sequences to the left of each series type indicate the specific sequence associated with each type.
Five different families of peptides are shown in Table 1, all of which contain extended stretches of proline residues. Family 1 has nine peptides composed solely of proline (from 13 to 25 residues long). Families 2, 3, and 4 contain a non-proline residue at the N-terminus (alanine, leucine/isoleucine/hydroxyproline, or glutamine/lysine), while Family 5 has serine at the C-terminus. The parent ions in families 1 and 2 include peptides with different numbers of prolines. Different length peptides could arise either naturally via exopeptidase action on the tetramerization peptide while it was attached to AChE in the circulation or via in-source fragmentation of the liberated peptide during introduction into the mass spectrometer. We have seen in-source fragmentation of model polyprolines, in the MALDI mass spectrometer, at laser energies of 4000–6000 volts, such as were used in the current experiments (LMS unpublished observations). However, the electrospray ionization energies employed to introduce samples into the Orbitrap are substantially lower making it less likely that in-source fragmentation is responsible for the variation in parent ion length seen in the Orbitrap experiments. Thus, it is likely that both possibilities for shortening the peptide are operative.
Examples of MSMS spectra for these peptides are given in Figures 2, 3 and 4. The principal ion series detected were a-, b- and y-ion series. The a-series were seen exclusively from peptides in complex with sodium. The b-ion series were found exclusively in the Orbitrap data. There were also major peaks at high mass due to sodium-promoted C-terminal fragmentation. There were stretches of minor peaks corresponding to series of prolines that were not associated with a-, b- or y-ion series. Finally, it was common to find a series of fragments associated with internal fragmentation at proline. Internal fragmentation at the relatively labile proline linkage is a well established phenomenon in mass spectrometry and would be expected under the current circumstances where many prolines are present in the peptides. Most of the unannotated peaks in the MSMS spectra could be accounted for by one-or-another of these types of fragmentation.
Figure 2.
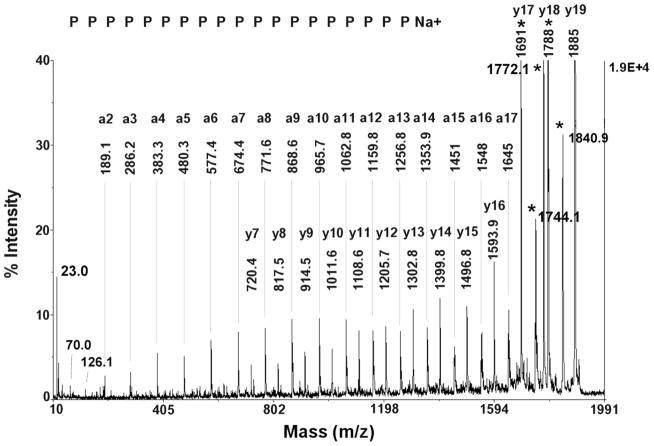
MALDI MSMS spectrum of the 1885.0 Da, 19-proline peptide (PPP)PP PPPPP PPPPP PPPP Na+ that was isolated from boiled FBS AChE by HPLC. Amino acid sequences are marked to show the y-ion series and a-ion series. The left-hand y-axis shows the signal intensity as a percentage of the most intense peak. The maximum value has been set to 40% to accentuate the minor peaks. Consequently, the most intense peaks (those at high mass) have been truncated. The right-hand y-axis indicates the signal intensity in counts-per-second. Ions from sodium-promoted fragmentation are identified by an asterisk (*). The immonium ion of proline is at 70 Da and a characteristic ion for proline is at 126 Da. The mass at 23 Da is sodium.
Figure 3.
MALDI MSMS spectrum of the 1721.9 Da peptide PPPQP PPPPP PPPPP PP Na+ isolated from boiled FBS AChE by HPLC. The right-hand y-axis indicates the signal intensity in counts-per-second. Ions from sodium-promoted fragmentation are identified by an asterisk (*). The immonium ion of proline is at 70 Da. The sodium atom is at 23 Da.
Figure 4.
Orbitrap MSMS spectrum of peptide PPPPP PPPPP PPP isolated from boiled FBS AChE by HPLC. The doubly-charged parent ion has a mass of 640.85 m/z. Amino acid sequences are marked to show singly-charged y-ion series, singly-charge b-ion series, a doubly-charged y-ion, and a doubly-charged b-ion series. The y-axis shows the signal intensity relative to the most intense peak in the spectrum.
3.2 MSMS fragmentation analysis, the first example
Figure 2 shows MSMS data for the 1885.0 Da peptide, taken with the MALDI mass spectrometer. The parent ion mass is consistent with the presence of 19 prolines plus a sodium atom. Presence of the sodium atom is confirmed by the appearance of a mass at 23 Da. The y-axis maximum has been lowered to emphasize the weaker signals. An array of peaks that defines a y-ion series and another array that define an a-ion series are annotated. The sodium atom remained associated with the N-terminal fragment in the a-ion series and with the C-terminal fragment in the y-ion series. There is also an array of peaks 1 Da larger than the a-ion series (an a+1 ion series) which were not annotated in order to avoid over-crowding of the spectrum. Seven residues in the low mass portion of the y-series gave no signals. However, the y7 mass (720.4 Da) is equal to the mass of a C-terminal proline (116 Da) plus the mass of 6 dehydro-prolines (6 × 97.05 Da), plus the mass of sodium (23 Da), and minus the mass of the proton displaced by the sodium (1 Da). There were no individual signals for the last three residues in the high mass portion of the a-ion series. However, the delta mass of 326.9 Da was equal to the C-terminal mass of a proline (143 Da) plus the dehydro-masses of proline (97 Da) and serine (87 Da).
The five most prominent peaks in the high mass region can be ascribed to sodium-promoted fragmentation at the C-terminus. Sodium-promoted fragmentation will be described in more detail in the section entitled “Sodium-promoted fragmentation”. The peaks from the sodium-promoted fragmentation are truncated in Figure 2. Their full height is 10-fold greater than most of the ions in the low mass region. The sodium-promoted fragment ions are marked with an asterisk (*). They include 1) the most intense signal, at 1788.1 Da, that is consistent with a neutral loss of the dehydro-mass for the C-terminal proline (97 Da); 2) an intense signal at 1691.0 Da that is consistent with loss of a second proline (97 Da) from the 1788.1 Da mass; 3) a mass at 1840.9 Da that is consistent with the loss of CO2 (44 Da) from the parent ion; 4) a mass at 1772.1 Da that is 16 Da smaller than the 1788.1 Da mass; and 5) a mass at 1744.1 Da which is consistent with loss of CO2 from the 1788.1 Da mass. The 1788.1 and 1691.0 Da peaks overlap with the y17 and y18 ions from the y-ion series, however the intensities of these peaks are much higher than that for the rest of the y-ion series indicating that there is a second source for ions at these positions. The masses at 1744.1, 1772.1 and 1841.8 Da do not correspond to any sequence ion. Identification and assignment of these fragments was made by consulting the literature and from unpublished observations on fragmentation of pure polyproline.
Finally, there are small, but distinct signals for the immonium ion of proline at 70 Da [12] and for a characteristic proline fragment at 126 Da [listed on the Protein Prospector website; http://prospector.ucsf.edu] (refer to Figure 2). We suggest that this 19 residue peptide is a fragment of the 25 residue polyproline peptide that is also listed in Table 1.
3.3 Sodium-promoted fragmentation
A sodium ion (Na+) can form a complex with peptides in which the Na+ displaces a proton to create what is referred to as an alkali metal cationized peptide. The most prominent ion that results from fragmentation of cationized peptides in the mass spectrometer is due to the loss of the dehydro-mass of the C-terminal residue. This yields a new peptide that is one amino acid shorter than the original peptide [13–17]. The mechanism for this fragmentation “involves the transfer of the hydroxyl group from the C-terminus of the peptide to the adjacent amino acid and subsequent loss of the residue mass of the C-terminal amino acid [97 Da for proline] leading ultimately to the production of a new alkali cationized peptide lacking the original C-terminal residue” [17]. Additional fragmentation can occur for the newly formed alkali cationized peptide resulting in ions which are two and even three amino acids shorter than the original peptide. The signal intensity of the ion from the second fragmentation can be comparable to that from the first, but the signal intensity for the ion from third fragmentation is generally much smaller [18].
In Figure 2, the first cycle of sodium-promoted fragmentation is consistent with formation of the intense, 1788.1 Da fragment. A second cycle of sodium-promoted fragmentation would be consistent with the intense signal at 1691.0 Da. However, assigning subsequent fragments to sodium-promoted fragmentation is complicated by overlapping fragments from the weaker, y-ion series. The complication arises from two facts. First, the delta mass for loss of the N-terminal residue from a y-ion series (the high mass end of the spectrum) is equal to the dehydro-mass of the residue. The delta mass for sodium-promoted fragmentation of the C-terminal residue is also equal to the dehydro-mass. Second, the peptides in this study have proline residues at both the C-terminus and the N-terminus (refer to Table 1). Consequently, the delta mass for release of the N-terminal residue for a y-ion series is the same as the delta mass for release of the C-terminal residue by sodium-promoted fragmentation. Discrimination between the two pathways relies on the relative intensities of the fragments. The principal contribution to high intensity signals can be ascribed to sodium-promoted fragmentation, with minor contribution coming from y-ion fragments when the masses from the two pathways overlap.
Additional sodium-promoted fragmentation can result in a prominent loss of 44 Da from the parent ion (probably due to loss of CO2) [15]. A prominent mass at 1840.9 Da in Figure 2 corresponds to this fragmentation. Another loss of 44 Da is associated with the 1788.1 Da fragment. Finally, for polyproline peptides, a novel mass, 113 Da smaller than the parent ion (equal to 97 Da plus 16) is typically seen in sodium-promoted fragmentation spectra (L.M. Schopfer, unpublished observations). The 1772.1 Da fragment in Figure 2 corresponds to this ion.
3.4 MSMS fragmentation, additional examples
Figure 3 shows MSMS data for the 1721.9 Da peptide, taken with the MALDI mass spectrometer. The parent ion mass is consistent with the presence of 16 prolines plus a glutamine (or lysine), and a sodium atom. Presence of the sodium atom is confirmed by the mass at 23 Da. Fragmentation is again dominated by sodium ion promoted processes. Masses at 1677.7, 1624.9, 1608.9, 1581.0 and 1527.9 Da are consistent with 1) loss of CO2 from the parent ion, 2) loss of a proline from the C-terminus, 3) loss of 16 Da from the 1624.9 Da ion, 4) loss of a CO2 from the 1624.9 Da mass, and 5) loss of a proline from the 1624.9 Da mass, respectively. An extensive y-ion series (that includes 13 consecutive prolines) can be extracted from the minor signals. Once again, the y-ion series overlaps the cation promoted fragments. A sequence interval of 128 Da for the difference between 1302.8 and 1430.8 Da corresponds to either a lysine or a glutamine residue. Y-ion and b-ion sequences from two analogous peptides (obtained with the Orbitrap mass spectrometer, see Table 1) indicated the presence of a glutamine/lysine residue on the N-terminus of a long string of prolines confirming the presence of this non-proline residue in the sequence (data not shown). We have assigned the residue to glutamine on the basis of BLAST search results (see the section entitled “BLAST search”).
The high intensity of the 1302.8 Da peak is also consistent with the presence of a non-proline residue at this position. The 1302.8 Da mass arises from cleavage on the N-terminal side of proline. The well known “Proline Effect” [19] is renowned for enhancing the intensity of fragments on the N-terminal side of proline. Finally, a distinct peak for the proline immonium ion was observed at 70 Da [12].
Figure 4 shows MSMS data for a doubly-charged peptide ([M+2H]+2 = 640.85 Da) taken with an LTQ Orbitrap mass spectrometer. The parent ion mass is consistent with the presence of 13 prolines. Singly-charged peaks for both y-ion and b-ion series are annotated. In addition, there are prominent peaks for doubly-charged b-ions and a y-ion. All of these ions are consistent with a peptide containing only prolines. It is noteworthy that there are strong signals for both a singly-charged y-ion series and a singly-charged b-ion series, and that there is no evidence for fragmentation favoring the loss of two prolines at a time, for either series. This can be compared to studies by Unnithan et. al on Pron-NH2 peptides (n = 4–10) who observed that “even numbers of Pro residues were lost from the N-terminal end of the peptide” to form a singly-charged y-ion series. B-ions were not detected [20]. Paired loss of prolines from proline-rich regions was supported by Wang et al. in studies of the proline-rich, 79-residue polypeptide, prophenin 1 [21]. A possible explanation for the difference in fragmentation patterns is that a much higher gas pressure was used in the fragmentation chamber of the Orbitrap (helium at 1×10−3 Torr) compared to that used by Unnithan et. al in the Brucker Esquire 3000 plus quadrupole ion trap (helium at 3×10−5 Torr).
3.5 BLAST search on the proline-rich peptides
A total of twenty proline-rich peptides were identified from FBS AChE (Table 1). All of these are suitable candidates for tetramer-organizing peptides. These peptides could be divided into five families. The origin of these peptides is unclear. A logical assumption would be that they begin as part of another protein. In order to test this assumption, the longest representative from each family was submitted for a BLAST (Basic Logical Alignment Search Tool) search of the NCBInr database (November 2011) using the mammalian taxonomy (40674). Matches were found for each family. We had anticipated that the necessity for 15 or more prolines in a row would have imposed a severe limitation on the number of matching candidates. However, this was not the case. For example, we found 25 separate proteins that matched the APPPP PPPPP PPPPP P sequence from family 2. Attempting to sort through this abundance of candidates for those that interact specifically with AChE is beyond the scope of the current study. Therefore, we have selected representative examples for purposes of discussion, which are shown in Table 2. When possible, a protein from the Bos taurus taxonomy was chosen for Table 2. When no candidates appeared in the Bos taurus taxonomy, a candidate from the Homo sapiens taxonomy was used. The proteins listed in Table 2 should not be assumed to be the actual source of the corresponding peptide found in AChE tetramers. This issue is addressed in greater detail in the Discussion, section 4.0.
Table 2.
Blast search results on the peptides released from FBS AChE
| Family | Sequence a | Blast Result b | Species | GI number d |
|---|---|---|---|---|
| 1 | PPPPP PPPPP PPPPP PPPPP PPPPP | UDP-N-acetyl glucosamine transferase ALG13 subunit homology domain-containing protein 1 | Homo c sapiens | 153791910 |
| 2 | APPPP PPPPP PPPPP P | Leiomodin-2 | Bos taurus | 157427900 |
| 3 | LPPPP PPPPP PPPPP P e IPPPP PPPPP PPPPP P LPPPP PPPPP PPPPP P |
Zinc finger protein 318; Predicted YLP motif-containing protein 1 lamellipodin |
Bos taurus Bos taurus Homo sapiens |
194677641 297479917 82581557 |
| 4 | Q/KPPPP PPPPP PPPPP PP | Paternally expressed 10 isoform 3 | Bos taurus | 187761320 |
| 5 | PPPPP PPPPP PPPPP PPPPP S | Zinc finger homeobox protein 4 | Bos taurus | 330340393 |
Peptides in this column represent the longest members of each family given in Table 1. Residues enclosed in brackets indicate that separate signals for the masses of each residue were not resolved in the mass spectrum. However, masses of the bracketed fragments exactly fit the indicated sequence.
Data were BLAST searched using the Mammalian taxonomy (40674) in the NCBInr database.
A matching sequence was not found in the bovine proteome, therefore a corresponding match from the Homo sapiens taxonomy was used.
The GI number is the protein accession number in the NCBI database in PubMed.
When leucine (L) was used, a match was obtained with the Zinc finger protein 318 of Bos taurus and with lamellipodin of Homo sapiens. When isoleucine (I) was used, a match was obtained with the predicted YLP motif-containing protein 1 of Bos taurus. Leucine and isoleucine cannot be distinguished on the basis of mass.
Thirteen of the peptides from Table 1 consisted of only proline residues (family 1). A Blast search using the 25-residue proline peptide found seven different proteins that contained a continuous stretch of 25 prolines. There was no match from the bovine proteome so “UDP-N-acetyl glucosamine transferase subunit homology domain-containing protein 1” from the Homo sapiens taxonomy was selected for use in Table 2.
The second family was represented by APPPP PPPPP PPPPP P. There was some ambiguity in the position of the alanine in the N-terminus (see Table 1). Consequently, a BLAST search was made with the alanine at positions 1, 2, 3 and 4 from the N-terminus. All four searches yielded matches to multiple proteins. With the alanine in the N-terminal position (as shown in Table 2) 25 proteins matched the sequence. Five of these proteins were found in the Bos taurus taxonomy (Predicted R3H domain-containing protein 1; Leiomodin-2; Predicted formin-like 1; Large proline-rich protein BAG6; and Zinc finger homeobox 4). Leiomodin-2 was selected for Table 2. Progressively fewer protein matches were found each time the alanine was moved one position toward the C-terminus. The PPPAP PPPPP PPPPP P peptide matched 6 proteins, only one of which was present in the first set of 25. The APPPP PPPPP PPPPP P peptide sequence was consistent with a tetramer-organizing peptide from leiomodin-2 that was reported by Biberoglu et. al for horse butyrylcholinesterase [22].
The third family was represented by peptide L/I/Hyp PPPPP PPPPP PPPPP. Though there was some ambiguity in the exact position of the L/I/Hyp residue (see Table 1), BLAST searches were made only with the L/I/Hyp residue in the N-terminal position. Leucine (L), isoleucine (I) and hydroxyproline (Hyp) all have the same nominal mass and therefore were not distinguished by mass analysis. BLAST searches were made for both the leucine and isoleucine possibilities. It was not possible to use hydroxyproline in a BLAST search. When leucine was used as the N-terminal residue, matches were obtained to 23 proteins, 3 of which were in the Bos taurus taxonomy (Protein diaphanous homolog 3, DIAPH3; Lysine-specific demethylase 6B; and Zinc finger protein 318). “Zinc finger protein 318” was used for Table 2. When isoleucine was used as the N-terminal residue, a match was obtained for 2 proteins, only one of which (Predicted YLP motif-containing protein 1) was from the Bos taurus taxonomy. The L PPPPP PPPPP PPPPP sequence was consistent with the tetramer-organizing peptide from lamellipodin that was reported by Li et. al for human butyrylcholinesterase (PSPPL PPPPP PPPPP PPPPP PPPPP LP) [8] and confirmed by Biberoglu et. al [22]. Lamellipodin from the Homo sapiens taxonomy was included in Table 2 to draw attention to the possibility that lamellipodin might be a source for this peptide in bovine as well.
The fourth family was represented by peptide PPPQ/KP PPPPP PPPPP PP. In this case, there was no ambiguity regarding the position of the Q/K residue. Lysine (K) and glutamine (Q) both have the same nominal mass and therefore precise identification of the correct residue could not be made from the mass spectrum. A BLAST search using lysine yielded no matches, whereas a search with glutamine yielded six proteins. “Paternally expressed 10 isoform 3” was the only match from the bovine proteome. Reversing the sequence to PPPPP PPPPP PPPQP PP or PPPPP PPPPP PPPKP PP resulted in no BLAST search matches, supporting the sequence as given in Table 2.
From the fifth family, the last three residues on the C-terminus of peptide PPPPP PPPPP PPPPP PPPPP S were not resolved (see Table 1). BLAST searches found a match only when serine was the C-terminal residue. That search yielded a match to only one protein, zinc finger homeobox protein 4, which was found in the Bos taurus taxonomy, among others.
In conclusion, the blast search results indicate that peptides from many proteins could participate in the tetramerization of FBS AChE.”
4. Discussion
4.1 Comparison of proline-rich peptides in soluble and membrane-anchored tetramers
Our study identified polyproline peptides in soluble, tetrameric FBS AChE. Previously, polyproline peptides have been identified in soluble, tetrameric human and horse butyrylcholinesterase (BChE) from plasma [8], [22]. The polyproline peptides in plasma AChE and plasma BChE resemble each other but are distinctly different from the proline-rich attachment domain (PRAD) of ColQ in the collagen tailed AChE in the neuromuscular junction, and from the PRAD of the proline-rich membrane anchor (PRiMA) in the membrane-anchored AChE of brain (Table 3). The polyproline peptides in plasma AChE and BChE tetramers have long strings of prolines up to 25 residues in length, whereas the proline-rich region of ColQ has 8 proline residues and that of PRiMA has 14 proline residues. In ColQ and PRiMA, the string of prolines is interrupted by Leu, LeuPhe, MetPhe or Met. Table 3 shows only one peptide for FBS AChE; additional peptides are in Table 1.
Table 3.
Comparison of tetramer organizing sequences in soluble and membrane-anchored tetramers of AChE and BChE
| Source | Sequence | Reference | Accession #a |
|---|---|---|---|
| fetal bovine serum AChE | PPPPPPPPPPPPPPPPPPPPPPPPP25 | this work | |
| human plasma BChE | PSPPLPPPPPPPPPPPPPPPPPPPPLP27 | [8] | gi 82581557 |
| horse plasma BChE | PSPPLPPPPPPPPPPPPPPPPPPPPLP27 | [8] | gi 82581557 |
| human ColQ | LLTPPPPPLFPPPFF15 | [26] | gi 18105016 |
| rat ColQ | LLMPPPPPLFPPPFF15 | [27, 28] | gi 9506503 |
| mouse ColQ | LLMPPPPPLFPPPFF15 | b | gi 122065161 |
| torpedo ColQ | LLTPPPPPMFPPPFF15 | [29] | gi 543769 |
| human PRiMA | RPPPPLPPPPPPPPPPRLLS20 | [4] | gi 29788778 |
| rat PRiMA | RPPPPLPPPPPPPPPPRLLS20 | b | gi 205830448 |
| mouse PRiMA | RPPPPLPPPPPPPPPPRLLS20 | [30] | gi 29824432 |
The NCBI accession numbers are for the proteins from which the proline-rich peptides are derived.
Mouse ColQ and rat PRiMA sequences were submitted to PubMed without a journal publication.
The differences in amino acid sequence between the tetramer organizing peptides from soluble AChE, BChE and the proline-rich peptides of ColQ and Prima are shown in Table 3. These differences mean that the tetramer organizing peptides of soluble cholinesterases are produced from genes that are distinct from those for ColQ and PRiMA. Thus, soluble AChE and BChE tetramers in plasma do not derive from the membrane-bound forms. For human and horse plasma BChE the source of the polyproline peptide appears to be a number of proteins including lamellipodin [8, 22]. In our first report we identified lamellipodin as the source of the tetramer organizing polyproline peptide [8] however subsequent investigations found that tetramer organizing peptides could be traced to several proteins [22]. Reasons for the differences in the results are discussed in Biberoglu et. al [22]. For FBS AChE there also appear to be multiple sources. Therefore no single gene encodes for all of the polyproline peptides identified in either soluble BChE or soluble AChE.
4.2 Noncovalent binding
The polyproline peptides are bound non-covalently to tetrameric FBS AChE. The peptides were easily released by denaturing the protein in boiling water, or by freezing the protein in the absence of a cryoprotectant. The interaction of polyproline with the tetramerization domain is very strong as the tetramer does not dissociate upon dilution down to nanomolar levels. Released peptides were visualized at 210 nm in an HPLC chromatogram and by mass spectrometry. However, the polyproline peptides were not detected on polyacrylamide gels stained with Coomassie blue or on gels reverse stained with imidazole-zinc [23].
4.3 A second gene product is required to make a tetramer
To date AChE and BChE are the only proteins known to require a polyproline peptide for assembly into tetramers. A search of the Protein Data Bank showed many tetrameric crystal structures but none included a polyproline peptide derived from a second gene. The common motif for tetramer organization is through interactions of side chains of the 4 subunits. AChE and BChE also have contacts between subunits but these are not sufficient to stabilize the tetramer. The cartoon in Figure 5 indicates the location of a polyproline peptide in the center of the tetramer. This cartoon is presented to emphasize the overall organization of the AChE tetramer and to focus attention on the role of the tetramerization peptide in organizing this structure. The planar symmetry and large contact area between the polyproline peptide and the core of the AChE are not intended to realistically represent the details of the interaction.
Figure 5.

Four identical subunits of fetal bovine serum AChE (accession number gi 108493 and gi 115497516) are linked by interactions with a polyproline peptide in the center of the tetramer. Dimers of AChE are formed by interchain disulfide bonds at Cysteine 580. The close contact between the globular subunits is intended to show that interactions between subunits contribute to the stability of the tetramer. Each AChE subunit contains 583 amino acids. The MW of the glycosylated tetramer is 340 kDa.
Muscle and brain contain not only membrane bound forms of AChE but also a small amount of soluble tetrameric AChE. It is not known what type of proline-rich peptide is present in tetrameric AChE in these tissues. We expect that future studies will identify additional tetrameric proteins that include a non-covalently bound polyproline peptide in the tetrameric structure.
4.4 The proline-rich peptides released from FBS AChE contain up to 25 residues
The crystal structure of the complex between synthetic human PRAD and 4 parallel chains representing the tetramerization domain showed that the coiled 15-residue ColQ PRAD peptide (LLTPP PPPLF PPPFF) has the same length as the helical 40-residue tetramerization domain of AChE [5]. The crystal structure of full-length tetrameric AChE complexed with a proline-rich peptide is not yet available. However, information from the crystal structure of synthetic peptides allowed Dvir et al. to produce a model of the AChE tetramer linked to the full length ColQ protein. Figure 6 (adapted from Dvir et. al.) shows that the ColQ protein traverses the entire AChE tetramer. The N-terminal region of ColQ interacts with the AChE C-terminal tetramerization domain in an anti-parallel head-to-tail configuration, passes through the center of the AChE tetramer and extends out the other side. This configuration will accommodate all sizes of polyproline peptides observed in our study, including the 25-residue polyproline peptide. A complete symmetry of the four subunit positions would not necessarily be expected since the tetramerization domains must accommodate to the helical structure of the polyproline.
Figure 6.
Model of the AChE tetramer in the presence of the full-length ColQ protein. AChE monomers are displayed as globular units with their tryptophan-rich, C-terminal, amphiphilic α-helices protruding out from the top of each unit. The ColQ protein, or alternatively the tetramer organizing polyproline peptide (yellow), projects through the middle of the AChE tetramer complex. The N-terminus of ColQ interacts with the C-terminal tetramerization domain of AChE (at the top) and extends as a single chain through the bottom of the complex. Figure reproduced from [5].
4.5 Requirement for protease activity
Protease activity is not involved in the production of the membrane-anchored forms of AChE. In contrast, the polyproline peptides in plasma AChE and plasma BChE derive from large precursor proteins that must undergo proteolysis to yield the proline-rich peptides. It is unknown whether the entire precursor protein is initially assembled together with the AChE tetramer followed by proteolysis, or whether proline-rich peptides are released from the precursor proteins before assembly with AChE subunits to yield AChE tetramers. The polyproline peptides released from FBS AChE range in size from 1280 to 2445 Da corresponding to 13 to 25 amino acid residues. The various sizes could originate from protease activity in plasma. Mass spectrometry analysis of the low molecular weight plasma proteome has demonstrated that proteases including aminopeptidase and carboxypeptidase convert large proteins to overlapping series of related peptides [24].
4.6 Assembly of the AChE tetramer
The presence of a set of different polyproline peptides extracted from the cores of soluble AChE tetramers raises a number of questions. First, could the variety of tetramer-organizing peptides reflect AChE expression from different tissues? Second, unlike membrane-bound AChE tetramers (which employ dedicated proteins that carry proline-rich tetramer organizing domains) the tetramer organizing peptides for soluble AChE appear to come from proteins designed for other purposes. How do these precursor proteins release the polyproline peptides for use in AChE? Is the polyproline peptide added to the oligomers prior to secretion or do these associations occur spontaneously in the bloodstream? Is there any evidence for the presence of putative precursor proteins in the endoplasmic reticulum and/or Golgi where interaction with developing AChE might be expected to occur? Third, since the AChE used in our experiments came from pooled serum, could the diversity in tetramer-organizing peptides that was observed represent variations between individuals?
Question 1 The origin of soluble AChE in fetal bovine serum is unknown. However, it is firmly established that soluble butyrylcholinesterase (BChE) in human serum originates in the liver [25]. It is widely accepted that the origin of mammalian serum cholinesterases, in general, is the liver. Though fetal bovine could differ, it would be unexpected to find serum AChE coming from tissues other than the liver.
Question 2 All of the bovine proteins that were found to match the tetramerization peptides were examined in detail with the goal of determining a mechanism by which the peptide could become associated with AChE. Their sequences were obtained from the NCBInr Protein resource. Their functions, subcellular locations, and tissue distributions were obtained from UniProtKB protein knowledgebase.
In all, 11 bovine and 1 human protein(s), associated with the 5 peptide families, were examined. The proline-rich sequences from families 2, 3 and 5 are all located deep within the protein structure (97 to 1556 residues from the nearest terminus).
Nuclear proteins included: 1) Zinc finger protein 318 (gi 194677641) an endocrine regulator protein expressed in endocrine tissue (containing a family 3 peptide). 2) Lysine-specific demethylase 6B (gi 359076756) a histone demethylase with no specific tissue location (containing a family 3 peptide). 3) YLP motif-containing protein 1 (gi 297479917) involved in reduction of telomerase activity (containing a family 3 peptide). 4) Zinc finger homeobox protein 4 (gi296480588) a transcription regulator involved in neural and muscular differentiation expressed in brain, skeletal muscle, liver and stomach (containing both a peptide from family 2 and a peptide from family 5; different peptides). 5) R3H domain-containing protein 1 (gi 194664704) a nucleic acid binding protein (containing a family 2 peptide). and 6) myb-Related p42POP (gi 359075807) a transcriptional repressor and DNA binding protein (containing a family 2 peptide).
Cytoplasmic proteins (all interacting with the cytoskeleton) included: 7) Leiomodin-2 (gi 157427900) an actin and tropomyosin binding protein found in heart and skeletal muscle (containing a family 2 peptide). 8) Protein diaphanous homolog 3, DIAPH3 (gi 296481927) a profilin and Rho binding protein that promotes actin polymerization with no tissue specificity (containing a family 3 peptide). and 9) Formin-like 1 (gi 297487196) a regulator of cell morphology and cytoskeletal organization that binds to actin and Rho (containing a family 2 peptide).
Two of the remaining three protein matches (to families 1 & 2) are more encouraging in that they are associated with the endoplasmic reticulum and therefore could theoretically come into contact with AChE as it is being synthesized. UDP-N-acetylglucosamine transferase subunit ALG13 homology domain-containing protein 1 (gi 153791910) is involved with protein N-glycosylation in the endoplasmic reticulum. It contains the peptide from family 1. No proteins from the bovine taxonomy were matched to the family 1 peptide, therefore this protein from the human taxonomy has been taken as a credible substitute. The unfortunate feature of this protein is that the tetramerization peptide is located 192 residues from the C-terminus, and further from the N-terminus. The large proline-rich protein BAG6 (gi 161621275) is a chaperone associated with insertion of tail-anchored proteins into the endoplasmic reticulum. It contains a family 2 peptide. Like the previous candidate, the tetramerization peptide is located well from either terminus (464 residues from the C-terminus). It should be noted that both of these proteins are associated with the cytosolic, not the luminal face of the endoplasmic reticulum. Their topology presents a problem for interactions with AChE.
The most appealing candidate of the 12 is mesoderm-specific transcript homology protein (MEST) that is also referred to as paternally expressed 10 isoform 3 (gi 187761320) (to family 4). The tetramerization peptide is ideally suited for association with fetal bovine AChE in that it is located 5 residues from the C-terminus. One can easily envision the proline helix providing an anchor around which four AChE monomers could cluster. Once formed, the mature AChE tetramer could be released by proteolysis of the MEST C-terminus. A similar mechanism with other proteins similar to MEST could account for the other tetramerization peptides isolated from soluble AChE tetramers. The major drawback for this candidate is that it is found in the nucleus and cytoplasm.
Question 3 Because the AChE used in these experiments came from multiple animals, the possibility that the different polyproline peptides arose from variations between individual animals cannot be excluded.
More detailed investigation of these issues is a matter for future studies.”
5. Conclusion
Soluble, tetrameric AChE molecules in plasma a contain polyproline peptide in their quaternary structure. Five types of polyproline peptide have been identified. These polyproline peptides are bound to the C-terminal tetramerization domain where they organize four AChE subunits into a tetramer. No single protein precursor is the source of these polyproline peptides.
Highlights.
Polyproline peptides are present in fetal bovine serum acetylcholinesterase.
We isolated five types of polyproline peptide.
One type was composed solely of prolines, 13 to 25 residues in length.
Four types included a single non-proline terminal residue.
The serum acetylcholinesterase tetramer is organized around a polyproline peptide.
Acknowledgments
This work was supported by a TÜBITAK grant from the Scientific and Technological Research council of Turkey to OT, a fellowship from Hacettepe University to KB, and NIH grant P30CA36727 to the Eppley Cancer Center.
The funding sources had no role in the study design; in collection, analysis and interpretation of the data; in writing of the report; or in the decision to submit the article for publication.
Mass spectra were obtained with the support of the Mass Spectrometry and Proteomics core facility at the University of Nebraska Medical Center.
Abbreviations
- AChE
acetylcholinesterase
- BChE
butyrylcholinesterase
- MALDI-TOF-TOF
matrix assisted laser desorption/ionization time-of-flight tandem mass spectrometry
- ColQ
a collagen like protein
- PRiMA
proline rich membrane
- PRAD
proline-rich attachment domain
- FBS
fetal bovine serum
- MS
mass spectrometry
- MSMS
tandem mass spectrometry
- BLAST
basic logical alignment search tool
- LC/MSMS
liquid chromatography/mass spectrometry including a peptide fragmentation stage
- LTQ-Orbitrap
linear ion-trap quadrupole mass spectrometer
- NCBInr
National Center for Biotechnology Information non-redundant database
Footnotes
None of the authors have any conflict of interest or potential conflict of interest regarding this paper.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
Contributor Information
Kevser Biberoglu, Email: Kevser.biberoglu@gmail.com.
Lawrence M. Schopfer, Email: lmschopf@unmc.edu.
Ashima Saxena, Email: ashima.saxena@us.army.mil.
Ozden Tacal, Email: otacal@yahoo.com.
Oksana Lockridge, Email: olockrid@unmc.edu.
References
- 1.Massoulie J, Pezzementi L, Bon S, Krejci E, Vallette FM. Molecular and cellular biology of cholinesterases. Prog Neurobiol. 1993;41:31–91. doi: 10.1016/0301-0082(93)90040-y. [DOI] [PubMed] [Google Scholar]
- 2.Li B, Stribley JA, Ticu A, Xie W, Schopfer LM, Hammond P, Brimijoin S, Hinrichs SH, Lockridge O. Abundant tissue butyrylcholinesterase and its possible function in the acetylcholinesterase knockout mouse. J Neurochem. 2000;75:1320–1331. doi: 10.1046/j.1471-4159.2000.751320.x. [DOI] [PubMed] [Google Scholar]
- 3.Feng G, Krejci E, Molgo J, Cunningham JM, Massoulie J, Sanes JR. Genetic analysis of collagen Q: roles in acetylcholinesterase and butyrylcholinesterase assembly and in synaptic structure and function. J Cell Biol. 1999;144:1349–1360. doi: 10.1083/jcb.144.6.1349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Perrier AL, Massoulie J, Krejci E. PRiMA: the membrane anchor of acetylcholinesterase in the brain. Neuron. 2002;33:275–285. doi: 10.1016/s0896-6273(01)00584-0. [DOI] [PubMed] [Google Scholar]
- 5.Dvir H, Harel M, Bon S, Liu WQ, Vidal M, Garbay C, Sussman JL, Massoulie J, Silman I. The synaptic acetylcholinesterase tetramer assembles around a polyproline II helix. EMBO J. 2004;23:4394–4405. doi: 10.1038/sj.emboj.7600425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Simon S, Krejci E, Massoulie J. A four-to-one association between peptide motifs: four C-terminal domains from cholinesterase assemble with one proline-rich attachment domain (PRAD) in the secretory pathway. EMBO J. 1998;17:6178–6187. doi: 10.1093/emboj/17.21.6178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bon S, Massoulie J. Quaternary associations of acetylcholinesterase. I. Oligomeric associations of T subunits with and without the amino-terminal domain of the collagen tail. J Biol Chem. 1997;272:3007–3015. doi: 10.1074/jbc.272.5.3007. [DOI] [PubMed] [Google Scholar]
- 8.Li H, Schopfer LM, Masson P, Lockridge O. Lamellipodin proline rich peptides associated with native plasma butyrylcholinesterase tetramers. Biochem J. 2008;411:425–432. doi: 10.1042/BJ20071551. [DOI] [PubMed] [Google Scholar]
- 9.Ralston JS, Rush RS, Doctor BP, Wolfe AD. Acetylcholinesterase from fetal bovine serum. Purification and characterization of soluble G4 enzyme. J Biol Chem. 1985;260:4312–4318. [PubMed] [Google Scholar]
- 10.De la Hoz D, Doctor BP, Ralston JS, Rush RS, Wolfe AD. A simplified procedure for the purification of large quantities of fetal bovine serum acetylcholinesterase. Life Sci. 1986;39:195–199. doi: 10.1016/0024-3205(86)90530-8. [DOI] [PubMed] [Google Scholar]
- 11.Ellman GL, Courtney KD, Andres V, Jr, Featherstone RM. A new and rapid colorimetric determination of acetylcholinesterase activity. Biochem Pharmacol. 1961;7:88–95. doi: 10.1016/0006-2952(61)90145-9. [DOI] [PubMed] [Google Scholar]
- 12.Hung CW, Schlosser A, Wei J, Lehmann WD. Collision-induced reporter fragmentations for identification of covalently modified peptides. Anal Bioanal Chem. 2007;389:1003–1016. doi: 10.1007/s00216-007-1449-y. [DOI] [PubMed] [Google Scholar]
- 13.Renner D, Spiteller G. Linked scan investigation of peptide degradation initiated by liquid secondary ion mass spectrometry. Biomed Environ Mass Spectrom. 1988;15:75–77. doi: 10.1002/bms.1200150204. [DOI] [PubMed] [Google Scholar]
- 14.Grese RP, Cerny RL, Gross ML. Metal ion-peptide interactions in the gas phase: a tandem mass spectrometry study of alkali metal cationized peptides. J Am Chem Soc. 1989;111:2835–2842. [Google Scholar]
- 15.Teesch LM, Adams J. Fragmentation of gas-phase complexes between alkali metal ions and peptides: metal ion binding to carbonyl oxygens and other neutral functional groups. J Am Chem Soc. 1991;113:812–820. [Google Scholar]
- 16.Lin T, Glish GL. C-terminal peptide sequencing via multistage mass spectrometry. Anal Chem. 1998;70:5162–5165. doi: 10.1021/ac980823v. [DOI] [PubMed] [Google Scholar]
- 17.Barr JM, Van Stipdonk MJ. Multi-stage tandem mass spectrometry of metal cationized leucine enkephalin and leucine enkephalin amide. Rapid Commun Mass Spectrom. 2002;16:566–578. doi: 10.1002/rcm.612. [DOI] [PubMed] [Google Scholar]
- 18.Tang XJ, Ens W, Standing KG, Westmore JB. Daughter ion mass spectra from cationized molecules of small oligopeptides in a reflecting time-of-flight mass spectrometer. Anal Chem. 1988;60:1791–1799. doi: 10.1021/ac00168a029. [DOI] [PubMed] [Google Scholar]
- 19.Bleiholder C, Suhai S, Harrison AG, Paizs B. Towards understanding the tandem mass spectra of protonated oligopeptides. 2: The proline effect in collision-induced dissociation of protonated Ala-Ala-Xxx-Pro-Ala (Xxx = Ala, Ser, Leu, Val, Phe, and Trp) Journal of the American Society for Mass Spectrometry. 2011;22:1032–1039. doi: 10.1007/s13361-011-0092-1. [DOI] [PubMed] [Google Scholar]
- 20.Unnithan AG, Myer MJ, Veale CJ, Danell AS. MS/MS of protonated polyproline peptides: the influence of N-terminal protonation on dissociation. Journal of the American Society for Mass Spectrometry. 2007;18:2198–2203. doi: 10.1016/j.jasms.2007.09.016. [DOI] [PubMed] [Google Scholar]
- 21.Wang Y, Johansson J, Griffiths WJ. Characterisation of variant forms of prophenin: mechanistic aspects of the fragmentation of proline-rich peptides. Rapid Commun Mass Spectrom. 2000;14:2182–2202. doi: 10.1002/1097-0231(20001215)14:23<2182::AID-RCM151>3.0.CO;2-7. [DOI] [PubMed] [Google Scholar]
- 22.Biberoglu K, Schopfer LM, Tacal O, Lockridge O. The proline-rich tetramerization peptides in equine serum butyrylcholinesterase. The FEBS journal. 2012;279:3844–3858. doi: 10.1111/j.1742-4658.2012.08744.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Fernandez-Patron C, Hardy E, Sosa A, Seoane J, Castellanos L. Double staining of Coomassie blue-stained polyacrylamide gels by imidazole-sodium dodecyl sulfate-zinc reverse staining: sensitive detection of coomassie blue-undetected proteins. Anal Biochem. 1995;224:263–269. doi: 10.1006/abio.1995.1039. [DOI] [PubMed] [Google Scholar]
- 24.Koomen JM, Li D, Xiao LC, Liu TC, Coombes KR, Abbruzzese J, Kobayashi R. Direct tandem mass spectrometry reveals limitations in protein profiling experiments for plasma biomarker discovery. J Proteome Res. 2005;4:972–981. doi: 10.1021/pr050046x. [DOI] [PubMed] [Google Scholar]
- 25.Khoury GF, Brill J, Walts L, Busuttil RW. Atypical serum cholinesterase eliminated by orthotopic liver transplantation. Anesthesiology. 1987;67:273–274. doi: 10.1097/00000542-198708000-00025. [DOI] [PubMed] [Google Scholar]