Abstract
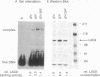
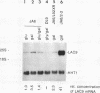
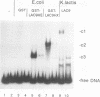
The concentration of the transcriptional activator LAC9 (KlGAL4) of Kluyveromyces lactis is moderately regulated by the carbon source as is the case for GAL4, its homolog in Saccharomyces cerevisiae. Expression of the LAC9 gene is induced about twofold in galactose. This induction is due to autoregulation. The LAC9 gene product binds to a low-affinity binding site in the LAC9 promoter and moderately activates transcription in response to galactose above a basal level. As for the LAC9-controlled metabolic genes, induction of LAC9 is inhibited in the presence of glucose. This inhibition of induction is a prerequisite for glucose repression of the lactose-galactose metabolic pathway. On the other hand, induced LAC9 levels are required for optimal growth on galactose, since mutating the LAC9 binding site in the LAC9 promoter resulted in poor growth and reduced expression of LAC9-controlled genes. Thus, in addition to the GAL80-dependent regulation by protein-protein interaction, the regulation of LAC9 gene expression is an important parameter in determining carbon source control of the LAC-GAL regulon. Although the mode of control is different, the pattern of LAC9 gene regulation resembles that of the S. cerevisiae GAL4 gene, being lower in glucose and glucose-galactose than in galactose.
Full text
PDF

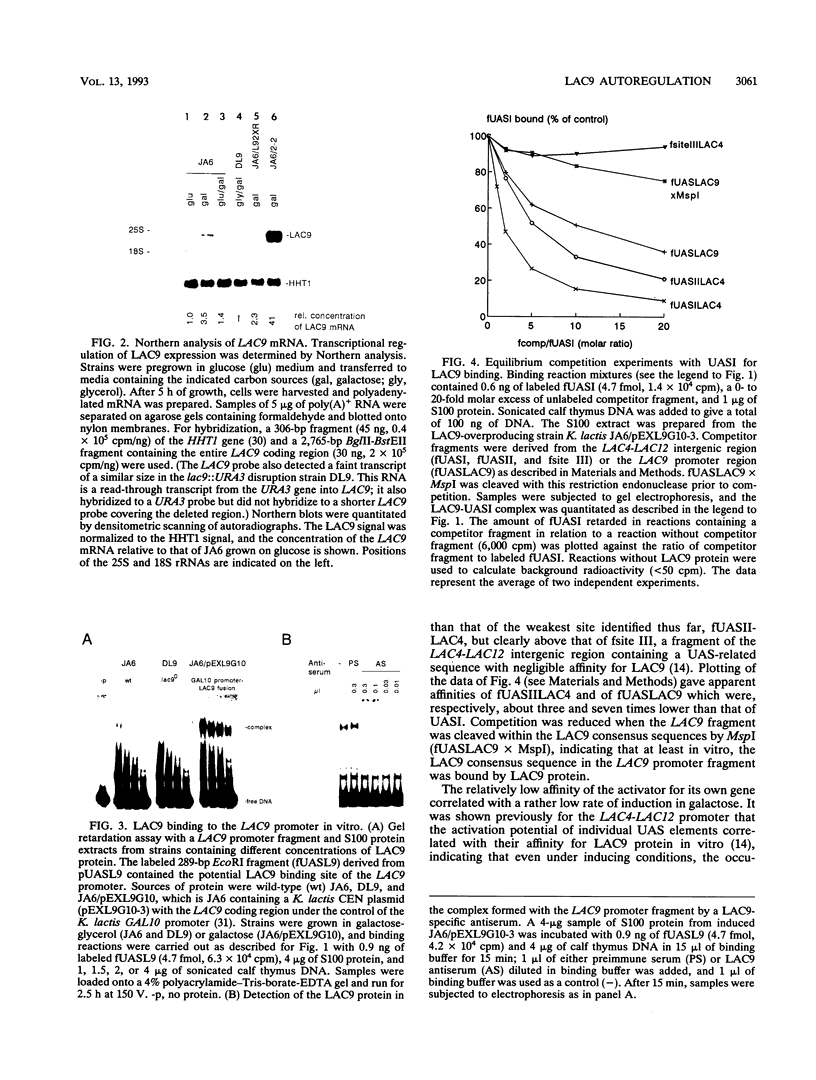

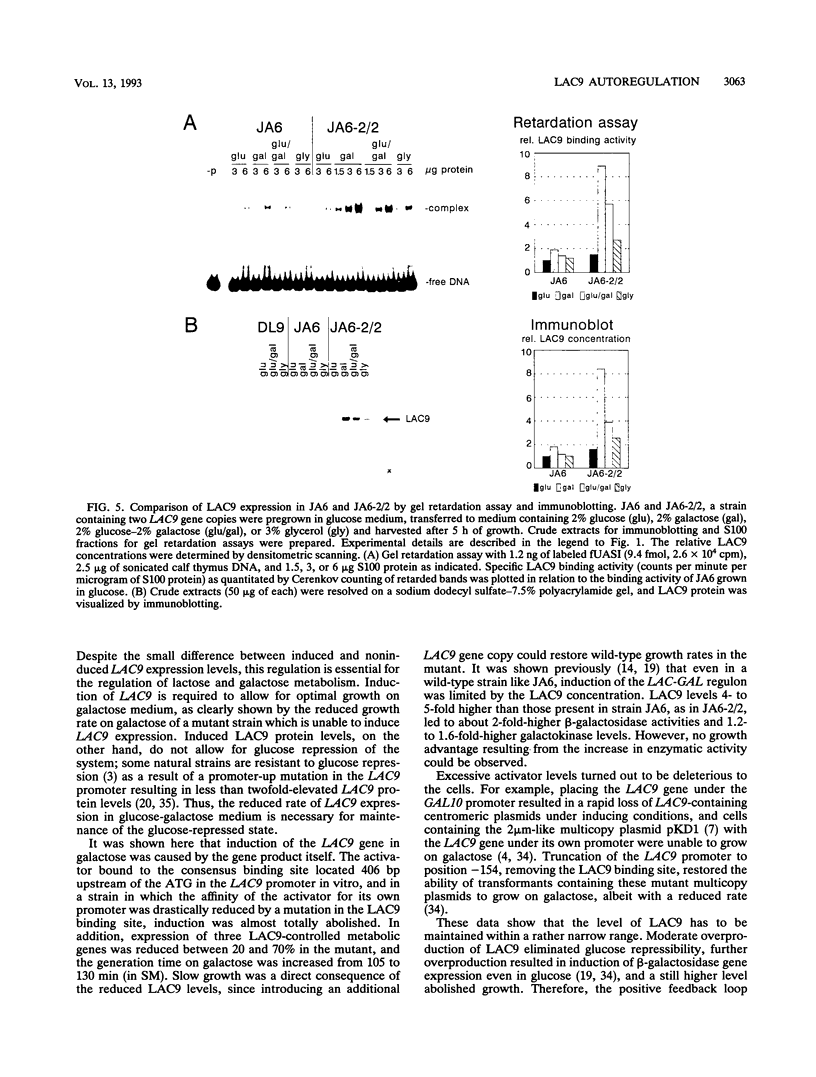
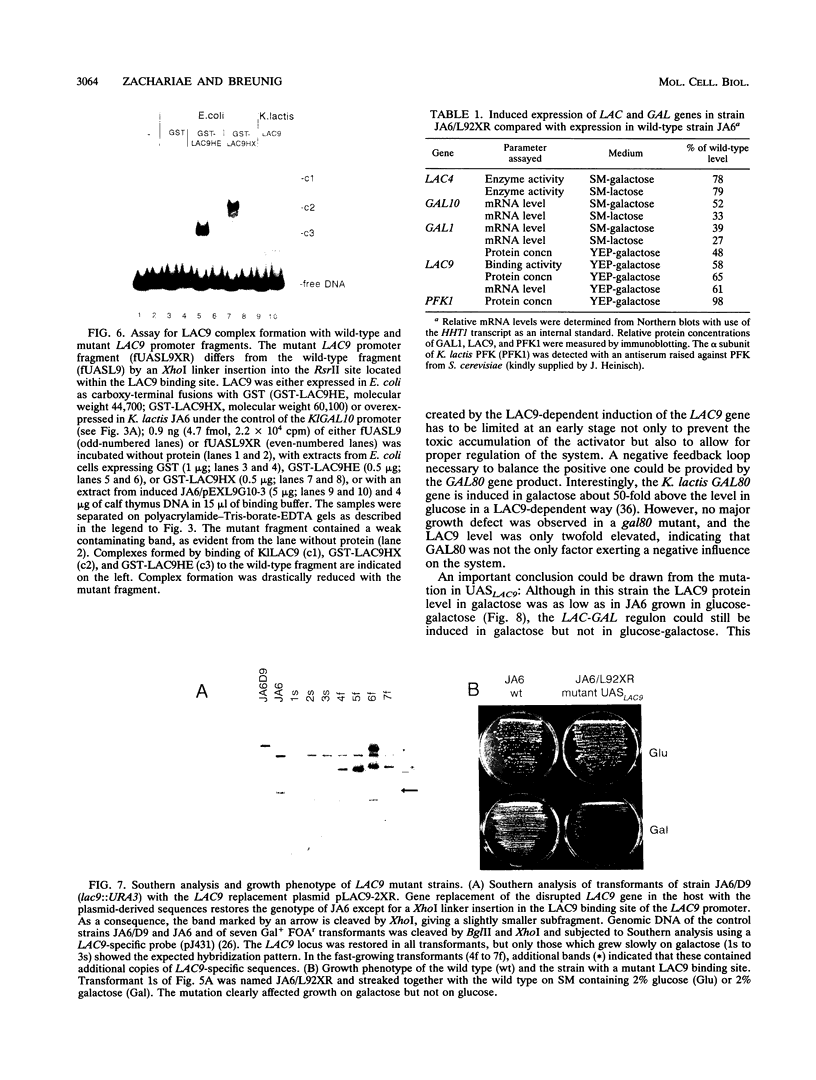
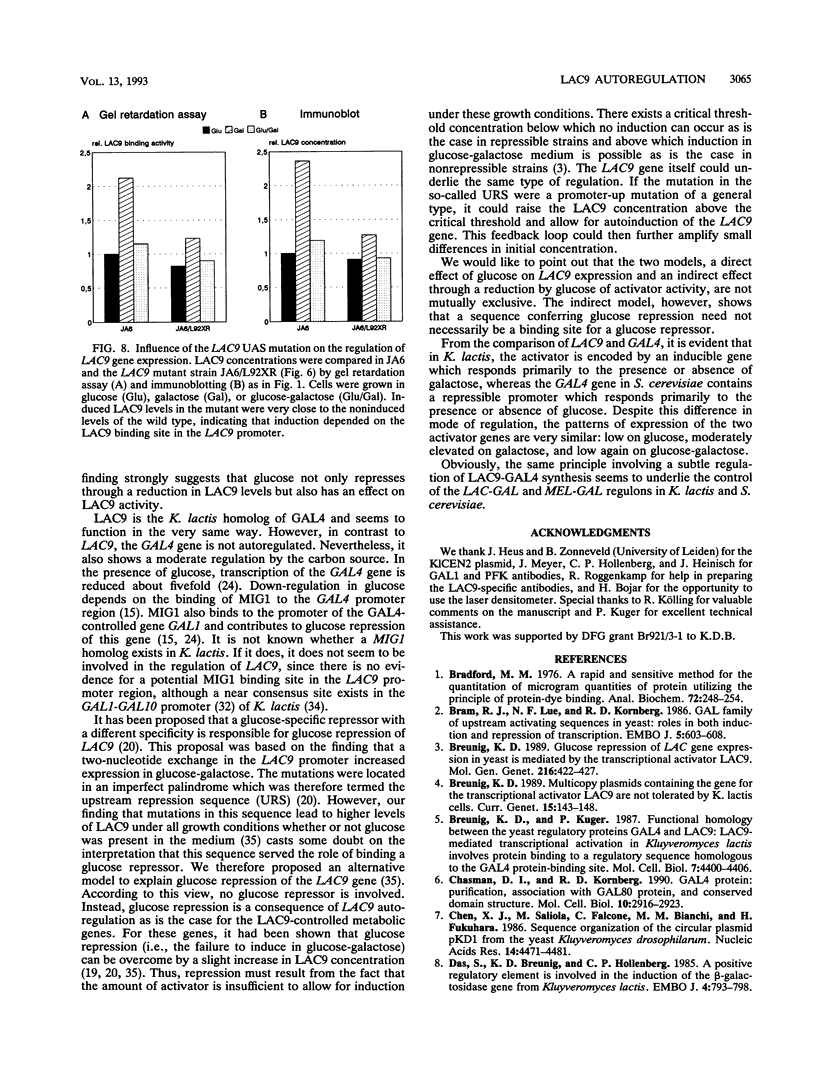

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Bram R. J., Lue N. F., Kornberg R. D. A GAL family of upstream activating sequences in yeast: roles in both induction and repression of transcription. EMBO J. 1986 Mar;5(3):603–608. doi: 10.1002/j.1460-2075.1986.tb04253.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breunig K. D. Glucose repression of LAC gene expression in yeast is mediated by the transcriptional activator LAC9. Mol Gen Genet. 1989 Apr;216(2-3):422–427. doi: 10.1007/BF00334386. [DOI] [PubMed] [Google Scholar]
- Breunig K. D., Kuger P. Functional homology between the yeast regulatory proteins GAL4 and LAC9: LAC9-mediated transcriptional activation in Kluyveromyces lactis involves protein binding to a regulatory sequence homologous to the GAL4 protein-binding site. Mol Cell Biol. 1987 Dec;7(12):4400–4406. doi: 10.1128/mcb.7.12.4400. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breunig K. D. Multicopy plasmids containing the gene for the transcriptional activator LAC9 are not tolerated by K. lactis cells. Curr Genet. 1989 Feb;15(2):143–148. doi: 10.1007/BF00435461. [DOI] [PubMed] [Google Scholar]
- Chasman D. I., Kornberg R. D. GAL4 protein: purification, association with GAL80 protein, and conserved domain structure. Mol Cell Biol. 1990 Jun;10(6):2916–2923. doi: 10.1128/mcb.10.6.2916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen X. J., Saliola M., Falcone C., Bianchi M. M., Fukuhara H. Sequence organization of the circular plasmid pKD1 from the yeast Kluyveromyces drosophilarum. Nucleic Acids Res. 1986 Jun 11;14(11):4471–4481. doi: 10.1093/nar/14.11.4471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das S., Breunig K. D., Hollenberg C. P. A positive regulatory element is involved in the induction of the beta-galactosidase gene from Kluyveromyces lactis. EMBO J. 1985 Mar;4(3):793–798. doi: 10.1002/j.1460-2075.1985.tb03699.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denis C. L., Ciriacy M., Young E. T. A positive regulatory gene is required for accumulation of the functional messenger RNA for the glucose-repressible alcohol dehydrogenase from Saccharomyces cerevisiae. J Mol Biol. 1981 Jun 5;148(4):355–368. doi: 10.1016/0022-2836(81)90181-9. [DOI] [PubMed] [Google Scholar]
- Dickson R. C., Gerardot C. J., Martin A. K. Genetic evidence for similar negative regulatory domains in the yeast transcription activators GAL4 and LAC9. Nucleic Acids Res. 1990 Sep 11;18(17):5213–5217. doi: 10.1093/nar/18.17.5213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dohmen R. J., Strasser A. W., Höner C. B., Hollenberg C. P. An efficient transformation procedure enabling long-term storage of competent cells of various yeast genera. Yeast. 1991 Oct;7(7):691–692. doi: 10.1002/yea.320070704. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Flick J. S., Johnston M. Two systems of glucose repression of the GAL1 promoter in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Sep;10(9):4757–4769. doi: 10.1128/mcb.10.9.4757. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Griggs D. W., Johnston M. Regulated expression of the GAL4 activator gene in yeast provides a sensitive genetic switch for glucose repression. Proc Natl Acad Sci U S A. 1991 Oct 1;88(19):8597–8601. doi: 10.1073/pnas.88.19.8597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gödecke A., Zachariae W., Arvanitidis A., Breunig K. D. Coregulation of the Kluyveromyces lactis lactose permease and beta-galactosidase genes is achieved by interaction of multiple LAC9 binding sites in a 2.6 kbp divergent promoter. Nucleic Acids Res. 1991 Oct 11;19(19):5351–5358. doi: 10.1093/nar/19.19.5351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heus J. J., Zonneveld B. J., Steensma H. Y., Van den Berg J. A. Centromeric DNA of Kluyveromyces lactis. Curr Genet. 1990 Dec;18(6):517–522. doi: 10.1007/BF00327022. [DOI] [PubMed] [Google Scholar]
- Johnston S. A., Salmeron J. M., Jr, Dincher S. S. Interaction of positive and negative regulatory proteins in the galactose regulon of yeast. Cell. 1987 Jul 3;50(1):143–146. doi: 10.1016/0092-8674(87)90671-4. [DOI] [PubMed] [Google Scholar]
- Klebe R. J., Harriss J. V., Sharp Z. D., Douglas M. G. A general method for polyethylene-glycol-induced genetic transformation of bacteria and yeast. Gene. 1983 Nov;25(2-3):333–341. doi: 10.1016/0378-1119(83)90238-x. [DOI] [PubMed] [Google Scholar]
- Kuger P., Gödecke A., Breunig K. D. A mutation in the Zn-finger of the GAL4 homolog LAC9 results in glucose repression of its target genes. Nucleic Acids Res. 1990 Feb 25;18(4):745–751. doi: 10.1093/nar/18.4.745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuzhandaivelu N., Jones W. K., Martin A. K., Dickson R. C. The signal for glucose repression of the lactose-galactose regulon is amplified through subtle modulation of transcription of the Kluyveromyces lactis Kl-GAL4 activator gene. Mol Cell Biol. 1992 May;12(5):1924–1931. doi: 10.1128/mcb.12.5.1924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lamphier M. S., Ptashne M. Multiple mechanisms mediate glucose repression of the yeast GAL1 gene. Proc Natl Acad Sci U S A. 1992 Jul 1;89(13):5922–5926. doi: 10.1073/pnas.89.13.5922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma J., Ptashne M. The carboxy-terminal 30 amino acids of GAL4 are recognized by GAL80. Cell. 1987 Jul 3;50(1):137–142. doi: 10.1016/0092-8674(87)90670-2. [DOI] [PubMed] [Google Scholar]
- Nehlin J. O., Carlberg M., Ronne H. Control of yeast GAL genes by MIG1 repressor: a transcriptional cascade in the glucose response. EMBO J. 1991 Nov;10(11):3373–3377. doi: 10.1002/j.1460-2075.1991.tb04901.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruzzi M., Breunig K. D., Ficca A. G., Hollenberg C. P. Positive regulation of the beta-galactosidase gene from Kluyveromyces lactis is mediated by an upstream activation site that shows homology to the GAL upstream activation site of Saccharomyces cerevisiae. Mol Cell Biol. 1987 Mar;7(3):991–997. doi: 10.1128/mcb.7.3.991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salmeron J. M., Jr, Johnston S. A. Analysis of the Kluyveromyces lactis positive regulatory gene LAC9 reveals functional homology to, but sequence divergence from, the Saccharomyces cerevisiae GAL4 gene. Nucleic Acids Res. 1986 Oct 10;14(19):7767–7781. doi: 10.1093/nar/14.19.7767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salmeron J. M., Jr, Langdon S. D., Johnston S. A. Interaction between transcriptional activator protein LAC9 and negative regulatory protein GAL80. Mol Cell Biol. 1989 Jul;9(7):2950–2956. doi: 10.1128/mcb.9.7.2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith D. B., Johnson K. S. Single-step purification of polypeptides expressed in Escherichia coli as fusions with glutathione S-transferase. Gene. 1988 Jul 15;67(1):31–40. doi: 10.1016/0378-1119(88)90005-4. [DOI] [PubMed] [Google Scholar]
- Stark M. J., Milner J. S. Cloning and analysis of the Kluyveromyces lactis TRP1 gene: a chromosomal locus flanked by genes encoding inorganic pyrophosphatase and histone H3. Yeast. 1989 Jan-Feb;5(1):35–50. doi: 10.1002/yea.320050106. [DOI] [PubMed] [Google Scholar]
- Webster T. D., Dickson R. C. Nucleotide sequence of the galactose gene cluster of Kluyveromyces lactis. Nucleic Acids Res. 1988 Aug 25;16(16):8192–8194. doi: 10.1093/nar/16.16.8192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Webster T. D., Dickson R. C. The organization and transcription of the galactose gene cluster of Kluyveromyces lactis. Nucleic Acids Res. 1988 Aug 25;16(16):8011–8028. doi: 10.1093/nar/16.16.8011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wray L. V., Jr, Witte M. M., Dickson R. C., Riley M. I. Characterization of a positive regulatory gene, LAC9, that controls induction of the lactose-galactose regulon of Kluyveromyces lactis: structural and functional relationships to GAL4 of Saccharomyces cerevisiae. Mol Cell Biol. 1987 Mar;7(3):1111–1121. doi: 10.1128/mcb.7.3.1111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zachariae W., Kuger P., Breunig K. D. Glucose repression of lactose/galactose metabolism in Kluyveromyces lactis is determined by the concentration of the transcriptional activator LAC9 (K1GAL4) [corrected]. Nucleic Acids Res. 1993 Jan 11;21(1):69–77. doi: 10.1093/nar/21.1.69. [DOI] [PMC free article] [PubMed] [Google Scholar]