Abstract
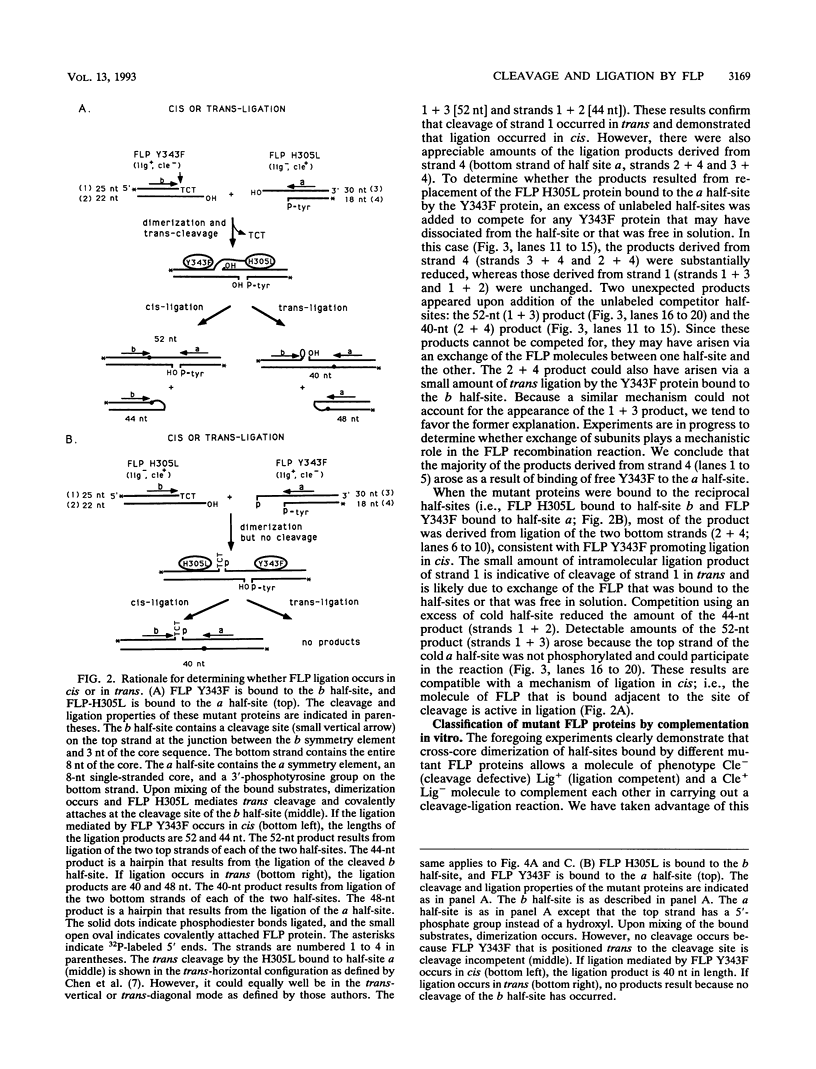
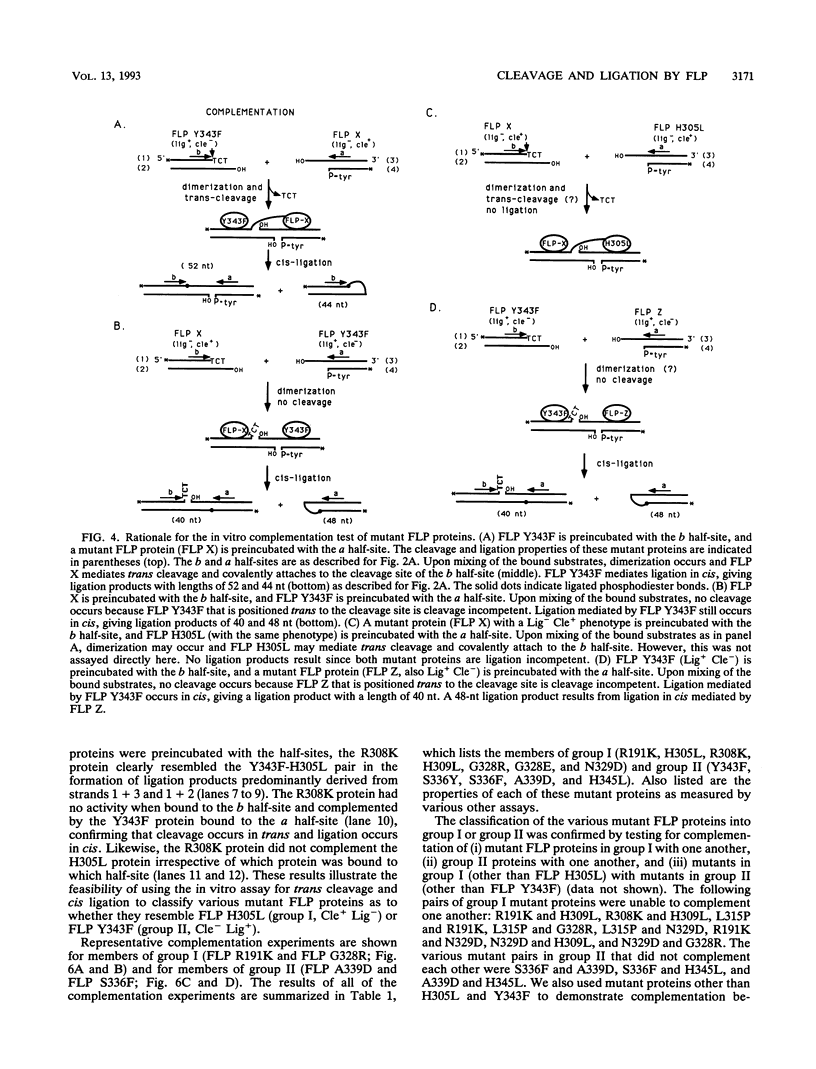
The FLP recombinase of the 2 microns plasmid of Saccharomyces cerevisiae is a member of the integrase family of site-specific recombinases. Recombination catalyzed by members of this family proceeds via the ordered cleavage and religation of four strands of DNA. Although the amino acid sequences of integrase family members are quite different, each recombinase maintains an absolutely conserved tetrad of amino acids (R-191, H-305, R-308, Y-343; numbers are those of the FLP protein). This tetrad is presumed to reflect a common chemical mechanism for cleavage and ligation that has evolved among all family members. The tyrosine is the nucleophile that causes phosphodiester bond cleavage and covalently attaches to the 3'-PO4 terminus, whereas the other three residues have been implicated in ligation of strands. It has recently been shown that cleavage by FLP takes place in trans; that is, a FLP molecule binds adjacent to the site of cleavage but receives the nucleophilic tyrosine from a molecule of FLP that is bound to another FLP-binding element (J.-W. Chen, J. Lee, and M. Jayaram, Cell 69:647-658, 1992). These studies led us to examine whether the ligation step of the FLP reaction is performed by the FLP molecule bound adjacent to the cleavage site (ligation in cis). We have found that FLP promotes ligation in cis. Furthermore, using in vitro complementation analysis, we have classified several mutant FLP proteins into one of two groups: those proteins that are cleavage competent but ligation deficient (group I) and those that are ligation competent but cleavage defective (group II). This observation suggests that the active site of FLP is composed of several amino acid residues from each of two FLP molecules.
Full text
PDF




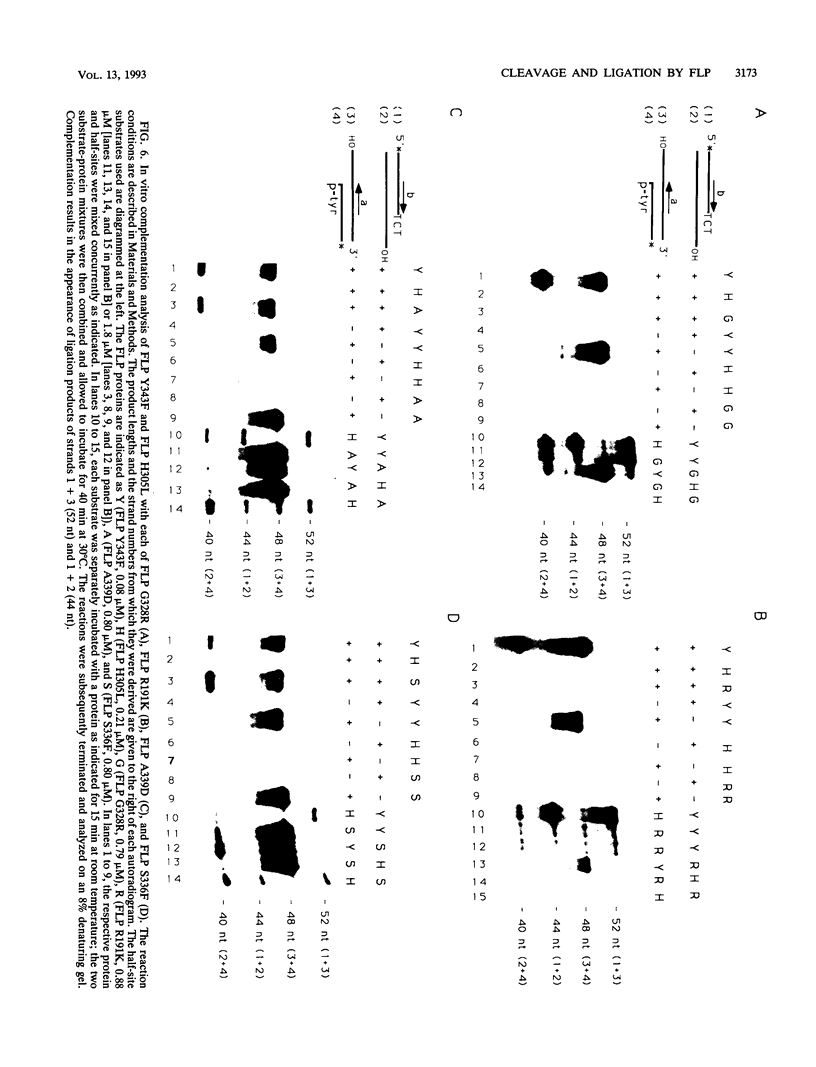
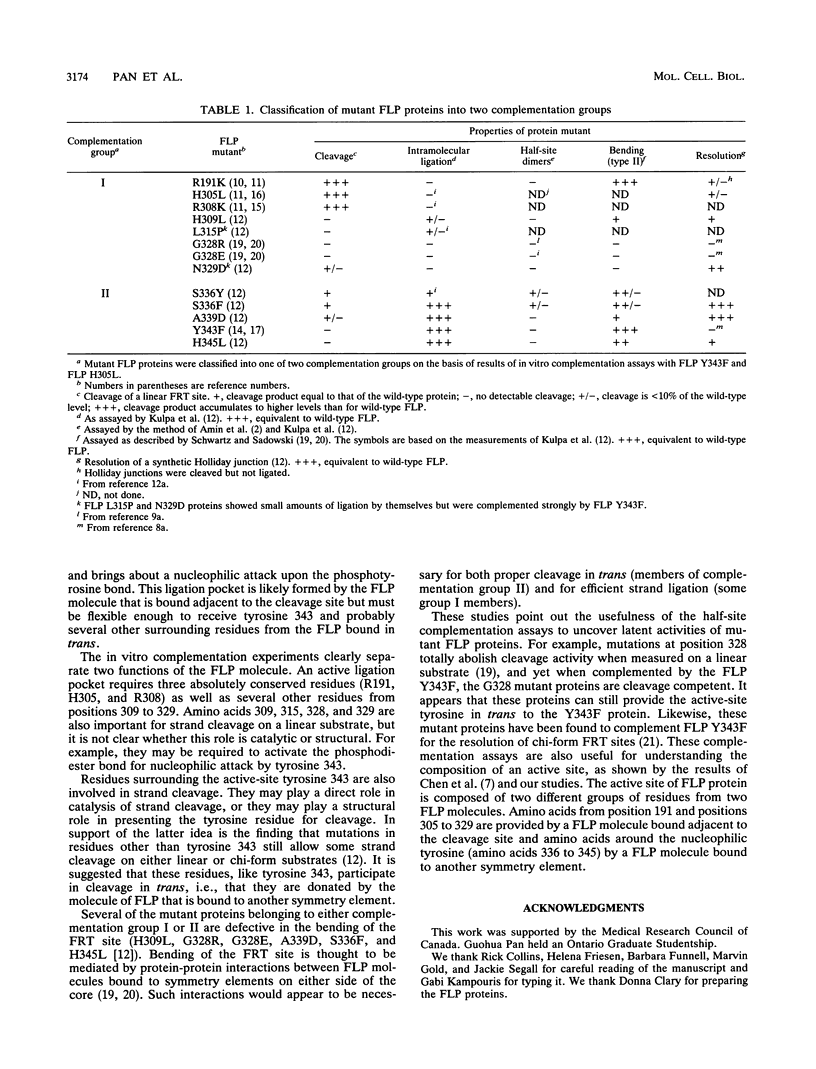

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abremski K. E., Hoess R. H. Evidence for a second conserved arginine residue in the integrase family of recombination proteins. Protein Eng. 1992 Jan;5(1):87–91. doi: 10.1093/protein/5.1.87. [DOI] [PubMed] [Google Scholar]
- Amin A., Roca H., Luetke K., Sadowski P. D. Synapsis, strand scission, and strand exchange induced by the FLP recombinase: analysis with half-FRT sites. Mol Cell Biol. 1991 Sep;11(9):4497–4508. doi: 10.1128/mcb.11.9.4497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrews B. J., Beatty L. G., Sadowski P. D. Isolation of intermediates in the binding of the FLP recombinase of the yeast plasmid 2-micron circle to its target sequence. J Mol Biol. 1987 Jan 20;193(2):345–358. doi: 10.1016/0022-2836(87)90223-3. [DOI] [PubMed] [Google Scholar]
- Andrews B. J., Proteau G. A., Beatty L. G., Sadowski P. D. The FLP recombinase of the 2 micron circle DNA of yeast: interaction with its target sequences. Cell. 1985 Apr;40(4):795–803. doi: 10.1016/0092-8674(85)90339-3. [DOI] [PubMed] [Google Scholar]
- Argos P., Landy A., Abremski K., Egan J. B., Haggard-Ljungquist E., Hoess R. H., Kahn M. L., Kalionis B., Narayana S. V., Pierson L. S., 3rd The integrase family of site-specific recombinases: regional similarities and global diversity. EMBO J. 1986 Feb;5(2):433–440. doi: 10.1002/j.1460-2075.1986.tb04229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruckner R. C., Cox M. M. Specific contacts between the FLP protein of the yeast 2-micron plasmid and its recombination site. J Biol Chem. 1986 Sep 5;261(25):11798–11807. [PubMed] [Google Scholar]
- Chen J. W., Lee J., Jayaram M. DNA cleavage in trans by the active site tyrosine during Flp recombination: switching protein partners before exchanging strands. Cell. 1992 May 15;69(4):647–658. doi: 10.1016/0092-8674(92)90228-5. [DOI] [PubMed] [Google Scholar]
- Evans B. R., Chen J. W., Parsons R. L., Bauer T. K., Teplow D. B., Jayaram M. Identification of the active site tyrosine of Flp recombinase. Possible relevance of its location to the mechanism of recombination. J Biol Chem. 1990 Oct 25;265(30):18504–18510. [PubMed] [Google Scholar]
- Friesen H., Sadowski P. D. Mutagenesis of a conserved region of the gene encoding the FLP recombinase of Saccharomyces cerevisiae. A role for arginine 191 in binding and ligation. J Mol Biol. 1992 May 20;225(2):313–326. doi: 10.1016/0022-2836(92)90924-9. [DOI] [PubMed] [Google Scholar]
- Jayaram M., Crain K. L., Parsons R. L., Harshey R. M. Holliday junctions in FLP recombination: resolution by step-arrest mutants of FLP protein. Proc Natl Acad Sci U S A. 1988 Nov;85(21):7902–7906. doi: 10.1073/pnas.85.21.7902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kulpa J., Dixon J. E., Pan G., Sadowski P. D. Mutations of the FLP recombinase gene that cause a deficiency in DNA bending and strand cleavage. J Biol Chem. 1993 Jan 15;268(2):1101–1108. [PubMed] [Google Scholar]
- Pan G., Luetke K., Juby C. D., Brousseau R., Sadowski P. Ligation of synthetic activated DNA substrates by site-specific recombinases and topoisomerase I. J Biol Chem. 1993 Feb 15;268(5):3683–3689. [PubMed] [Google Scholar]
- Pan G., Sadowski P. D. Ligation activity of FLP recombinase. The strand ligation activity of a site-specific recombinase using an activated DNA substrate. J Biol Chem. 1992 Jun 25;267(18):12397–12399. [PubMed] [Google Scholar]
- Parsons R. L., Evans B. R., Zheng L., Jayaram M. Functional analysis of Arg-308 mutants of Flp recombinase. Possible role of Arg-308 in coupling substrate binding to catalysis. J Biol Chem. 1990 Mar 15;265(8):4527–4533. [PubMed] [Google Scholar]
- Parsons R. L., Prasad P. V., Harshey R. M., Jayaram M. Step-arrest mutants of FLP recombinase: implications for the catalytic mechanism of DNA recombination. Mol Cell Biol. 1988 Aug;8(8):3303–3310. doi: 10.1128/mcb.8.8.3303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qian X. H., Inman R. B., Cox M. M. Protein-based asymmetry and protein-protein interactions in FLP recombinase-mediated site-specific recombination. J Biol Chem. 1990 Dec 15;265(35):21779–21788. [PubMed] [Google Scholar]
- Schwartz C. J., Sadowski P. D. FLP protein of 2 mu circle plasmid of yeast induces multiple bends in the FLP recognition target site. J Mol Biol. 1990 Nov 20;216(2):289–298. doi: 10.1016/s0022-2836(05)80320-1. [DOI] [PubMed] [Google Scholar]
- Schwartz C. J., Sadowski P. D. FLP recombinase of the 2 microns circle plasmid of Saccharomyces cerevisiae bends its DNA target. Isolation of FLP mutants defective in DNA bending. J Mol Biol. 1989 Feb 20;205(4):647–658. doi: 10.1016/0022-2836(89)90310-0. [DOI] [PubMed] [Google Scholar]
- Utatsu I., Sakamoto S., Imura T., Toh-e A. Yeast plasmids resembling 2 micron DNA: regional similarities and diversities at the molecular level. J Bacteriol. 1987 Dec;169(12):5537–5545. doi: 10.1128/jb.169.12.5537-5545.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]