Abstract
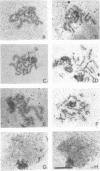
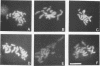
Although the processes of mitosis and meiosis are similar, there is evidence for fundamental regulatory differences between the two. To examine these differences, we have compared the meiotic phenotype of DNA topoisomerase II mutants with their previously described mitotic phenotype (C. Holm, T. Goto, J. Wang, and D. Botstein, Cell 41:553-563, 1985). top2 mutants in meiosis show no defects in the latest detectable stages of recombination, yet they arrest prior to spindle establishment at meiosis I. Fluorescence and electron microscopy reveal that top2 mutants exhibit wild-type levels of meiotic chromosome condensation and form morphologically normal synaptonemal complex but are delayed in the exit from pachytene. Arrested cells retain viability and form colonies if transferred to mitotic medium. Our results suggest that the top2 meiotic arrest is regulatory in nature. This arrest may have evolved to ensure the resolution of fortuitous tangles between nonhomologous chromosomes.
Full text
PDF










Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baldi M. I., Benedetti P., Mattoccia E., Tocchini-Valentini G. P. In vitro catenation and decatenation of DNA and a novel eucaryotic ATP-dependent topoisomerase. Cell. 1980 Jun;20(2):461–467. doi: 10.1016/0092-8674(80)90632-7. [DOI] [PubMed] [Google Scholar]
- Beach D., Durkacz B., Nurse P. Functionally homologous cell cycle control genes in budding and fission yeast. Nature. 1982 Dec 23;300(5894):706–709. doi: 10.1038/300706a0. [DOI] [PubMed] [Google Scholar]
- Bishop D. K., Park D., Xu L., Kleckner N. DMC1: a meiosis-specific yeast homolog of E. coli recA required for recombination, synaptonemal complex formation, and cell cycle progression. Cell. 1992 May 1;69(3):439–456. doi: 10.1016/0092-8674(92)90446-j. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Borts R. H., Lichten M., Hearn M., Davidow L. S., Haber J. E. Physical monitoring of meiotic recombination in Saccharomyces cerevisiae. Cold Spring Harb Symp Quant Biol. 1984;49:67–76. doi: 10.1101/sqb.1984.049.01.010. [DOI] [PubMed] [Google Scholar]
- Brown P. O., Cozzarelli N. R. A sign inversion mechanism for enzymatic supercoiling of DNA. Science. 1979 Nov 30;206(4422):1081–1083. doi: 10.1126/science.227059. [DOI] [PubMed] [Google Scholar]
- Davis R. W., Thomas M., Cameron J., St John T. P., Scherer S., Padgett R. A. Rapid DNA isolations for enzymatic and hybridization analysis. Methods Enzymol. 1980;65(1):404–411. doi: 10.1016/s0076-6879(80)65051-4. [DOI] [PubMed] [Google Scholar]
- Dresser M. E., Giroux C. N. Meiotic chromosome behavior in spread preparations of yeast. J Cell Biol. 1988 Mar;106(3):567–573. doi: 10.1083/jcb.106.3.567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunphy W. G., Brizuela L., Beach D., Newport J. The Xenopus cdc2 protein is a component of MPF, a cytoplasmic regulator of mitosis. Cell. 1988 Jul 29;54(3):423–431. doi: 10.1016/0092-8674(88)90205-x. [DOI] [PubMed] [Google Scholar]
- Earnshaw W. C., Halligan B., Cooke C. A., Heck M. M., Liu L. F. Topoisomerase II is a structural component of mitotic chromosome scaffolds. J Cell Biol. 1985 May;100(5):1706–1715. doi: 10.1083/jcb.100.5.1706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engebrecht J., Hirsch J., Roeder G. S. Meiotic gene conversion and crossing over: their relationship to each other and to chromosome synapsis and segregation. Cell. 1990 Sep 7;62(5):927–937. doi: 10.1016/0092-8674(90)90267-i. [DOI] [PubMed] [Google Scholar]
- Engebrecht J., Roeder G. S. Yeast mer1 mutants display reduced levels of meiotic recombination. Genetics. 1989 Feb;121(2):237–247. doi: 10.1093/genetics/121.2.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esposito M. S., Esposito R. E. The genetic control of sporulation in Saccharomyces. I. The isolation of temperature-sensitive sporulation-deficient mutants. Genetics. 1969 Jan;61(1):79–89. doi: 10.1093/genetics/61.1.79. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Esposito R. E., Esposito M. S. Genetic recombination and commitment to meiosis in Saccharomyces. Proc Natl Acad Sci U S A. 1974 Aug;71(8):3172–3176. doi: 10.1073/pnas.71.8.3172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gautier J., Norbury C., Lohka M., Nurse P., Maller J. Purified maturation-promoting factor contains the product of a Xenopus homolog of the fission yeast cell cycle control gene cdc2+. Cell. 1988 Jul 29;54(3):433–439. doi: 10.1016/0092-8674(88)90206-1. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H., Culotti J., Pringle J. R., Reid B. J. Genetic control of the cell division cycle in yeast. Science. 1974 Jan 11;183(4120):46–51. doi: 10.1126/science.183.4120.46. [DOI] [PubMed] [Google Scholar]
- Hartwell L. H., Weinert T. A. Checkpoints: controls that ensure the order of cell cycle events. Science. 1989 Nov 3;246(4930):629–634. doi: 10.1126/science.2683079. [DOI] [PubMed] [Google Scholar]
- Hirano T., Mitchison T. J. Topoisomerase II does not play a scaffolding role in the organization of mitotic chromosomes assembled in Xenopus egg extracts. J Cell Biol. 1993 Feb;120(3):601–612. doi: 10.1083/jcb.120.3.601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollingsworth N. M., Byers B. HOP1: a yeast meiotic pairing gene. Genetics. 1989 Mar;121(3):445–462. doi: 10.1093/genetics/121.3.445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holm C., Goto T., Wang J. C., Botstein D. DNA topoisomerase II is required at the time of mitosis in yeast. Cell. 1985 Jun;41(2):553–563. doi: 10.1016/s0092-8674(85)80028-3. [DOI] [PubMed] [Google Scholar]
- Holm C., Stearns T., Botstein D. DNA topoisomerase II must act at mitosis to prevent nondisjunction and chromosome breakage. Mol Cell Biol. 1989 Jan;9(1):159–168. doi: 10.1128/mcb.9.1.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horesh O., Simchen G., Friedmann A. Morphogenesis of the synapton during yeast meiosis. Chromosoma. 1979 Oct 2;75(1):101–115. doi: 10.1007/BF00330628. [DOI] [PubMed] [Google Scholar]
- Howell W. M., Black D. A. Controlled silver-staining of nucleolus organizer regions with a protective colloidal developer: a 1-step method. Experientia. 1980 Aug 15;36(8):1014–1015. doi: 10.1007/BF01953855. [DOI] [PubMed] [Google Scholar]
- Hsieh T., Brutlag D. ATP-dependent DNA topoisonmerase from D. melanogaster reversibly catenates duplex DNA rings. Cell. 1980 Aug;21(1):115–125. doi: 10.1016/0092-8674(80)90119-1. [DOI] [PubMed] [Google Scholar]
- Hsieh T. Knotting of the circular duplex DNA by type II DNA topoisomerase from Drosophila melanogaster. J Biol Chem. 1983 Jul 10;258(13):8413–8420. [PubMed] [Google Scholar]
- Kleckner N., Padmore R., Bishop D. K. Meiotic chromosome metabolism: one view. Cold Spring Harb Symp Quant Biol. 1991;56:729–743. doi: 10.1101/sqb.1991.056.01.082. [DOI] [PubMed] [Google Scholar]
- Klein F., Laroche T., Cardenas M. E., Hofmann J. F., Schweizer D., Gasser S. M. Localization of RAP1 and topoisomerase II in nuclei and meiotic chromosomes of yeast. J Cell Biol. 1992 Jun;117(5):935–948. doi: 10.1083/jcb.117.5.935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreuzer K. N., Cozzarelli N. R. Formation and resolution of DNA catenanes by DNA gyrase. Cell. 1980 May;20(1):245–254. doi: 10.1016/0092-8674(80)90252-4. [DOI] [PubMed] [Google Scholar]
- Liu L. F., Davis J. L., Calendar R. Novel topologically knotted DNA from bacteriophage P4 capsids: studies with DNA topoisomerases. Nucleic Acids Res. 1981 Aug 25;9(16):3979–3989. doi: 10.1093/nar/9.16.3979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu L. F., Liu C. C., Alberts B. M. Type II DNA topoisomerases: enzymes that can unknot a topologically knotted DNA molecule via a reversible double-strand break. Cell. 1980 Mar;19(3):697–707. doi: 10.1016/s0092-8674(80)80046-8. [DOI] [PubMed] [Google Scholar]
- Malone R. E., Esposito R. E. Recombinationless meiosis in Saccharomyces cerevisiae. Mol Cell Biol. 1981 Oct;1(10):891–901. doi: 10.1128/mcb.1.10.891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moens P. B., Earnshaw W. C. Anti-topoisomerase II recognizes meiotic chromosome cores. Chromosoma. 1989 Nov;98(5):317–322. doi: 10.1007/BF00292383. [DOI] [PubMed] [Google Scholar]
- Murray A. W., Szostak J. W. Chromosome segregation in mitosis and meiosis. Annu Rev Cell Biol. 1985;1:289–315. doi: 10.1146/annurev.cb.01.110185.001445. [DOI] [PubMed] [Google Scholar]
- Nicklas R. B. The forces that move chromosomes in mitosis. Annu Rev Biophys Biophys Chem. 1988;17:431–449. doi: 10.1146/annurev.bb.17.060188.002243. [DOI] [PubMed] [Google Scholar]
- Padmore R., Cao L., Kleckner N. Temporal comparison of recombination and synaptonemal complex formation during meiosis in S. cerevisiae. Cell. 1991 Sep 20;66(6):1239–1256. doi: 10.1016/0092-8674(91)90046-2. [DOI] [PubMed] [Google Scholar]
- Purrello M., Balazs I. Direct hybridization of labeled DNA to DNA in agarose gels. Anal Biochem. 1983 Feb 1;128(2):393–397. doi: 10.1016/0003-2697(83)90391-3. [DOI] [PubMed] [Google Scholar]
- Rose D., Thomas W., Holm C. Segregation of recombined chromosomes in meiosis I requires DNA topoisomerase II. Cell. 1990 Mar 23;60(6):1009–1017. doi: 10.1016/0092-8674(90)90349-j. [DOI] [PubMed] [Google Scholar]
- Shuster E. O., Byers B. Pachytene arrest and other meiotic effects of the start mutations in Saccharomyces cerevisiae. Genetics. 1989 Sep;123(1):29–43. doi: 10.1093/genetics/123.1.29. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simchen G. Are mitotic functions required in meiosis? Genetics. 1974 Apr;76(4):745–753. doi: 10.1093/genetics/76.4.745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Uemura T., Ohkura H., Adachi Y., Morino K., Shiozaki K., Yanagida M. DNA topoisomerase II is required for condensation and separation of mitotic chromosomes in S. pombe. Cell. 1987 Sep 11;50(6):917–925. doi: 10.1016/0092-8674(87)90518-6. [DOI] [PubMed] [Google Scholar]
- Weber L., Byers B. A RAD9-dependent checkpoint blocks meiosis of cdc13 yeast cells. Genetics. 1992 May;131(1):55–63. doi: 10.1093/genetics/131.1.55. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinert T. A., Hartwell L. H. The RAD9 gene controls the cell cycle response to DNA damage in Saccharomyces cerevisiae. Science. 1988 Jul 15;241(4863):317–322. doi: 10.1126/science.3291120. [DOI] [PubMed] [Google Scholar]
- Zechiedrich E. L., Osheroff N. Eukaryotic topoisomerases recognize nucleic acid topology by preferentially interacting with DNA crossovers. EMBO J. 1990 Dec;9(13):4555–4562. doi: 10.1002/j.1460-2075.1990.tb07908.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wettstein D., Rasmussen S. W., Holm P. B. The synaptonemal complex in genetic segregation. Annu Rev Genet. 1984;18:331–413. doi: 10.1146/annurev.ge.18.120184.001555. [DOI] [PubMed] [Google Scholar]