Abstract
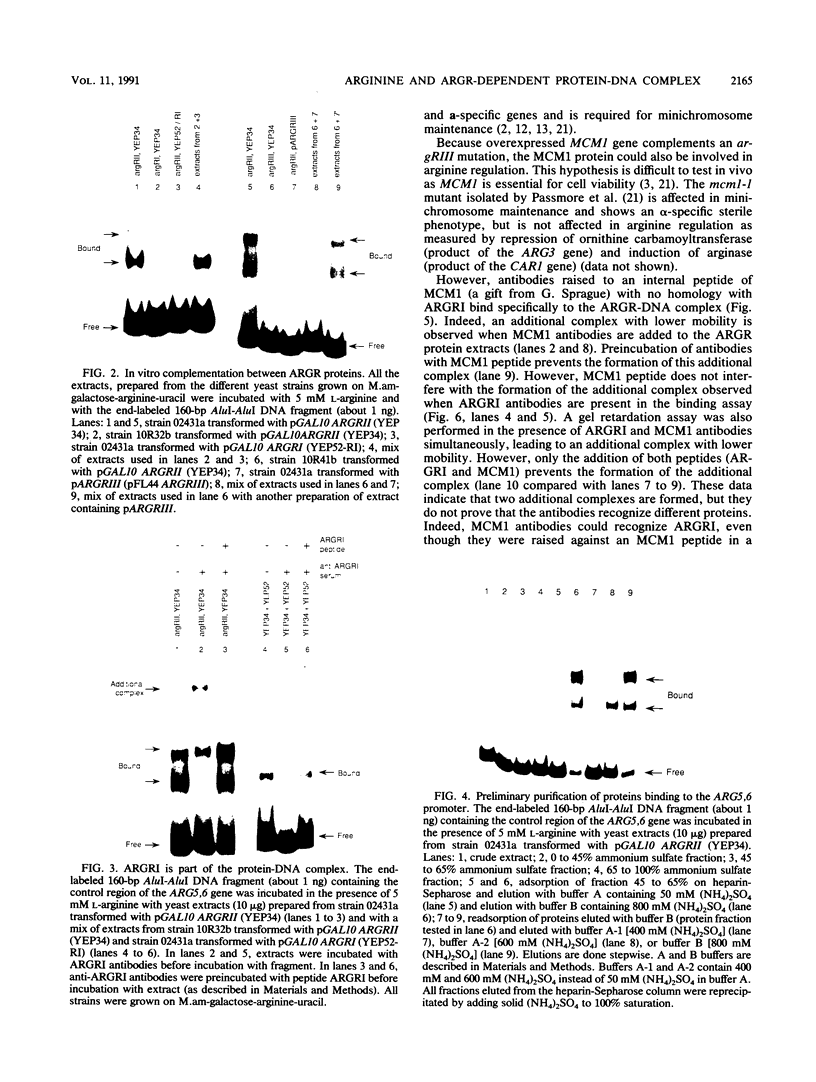
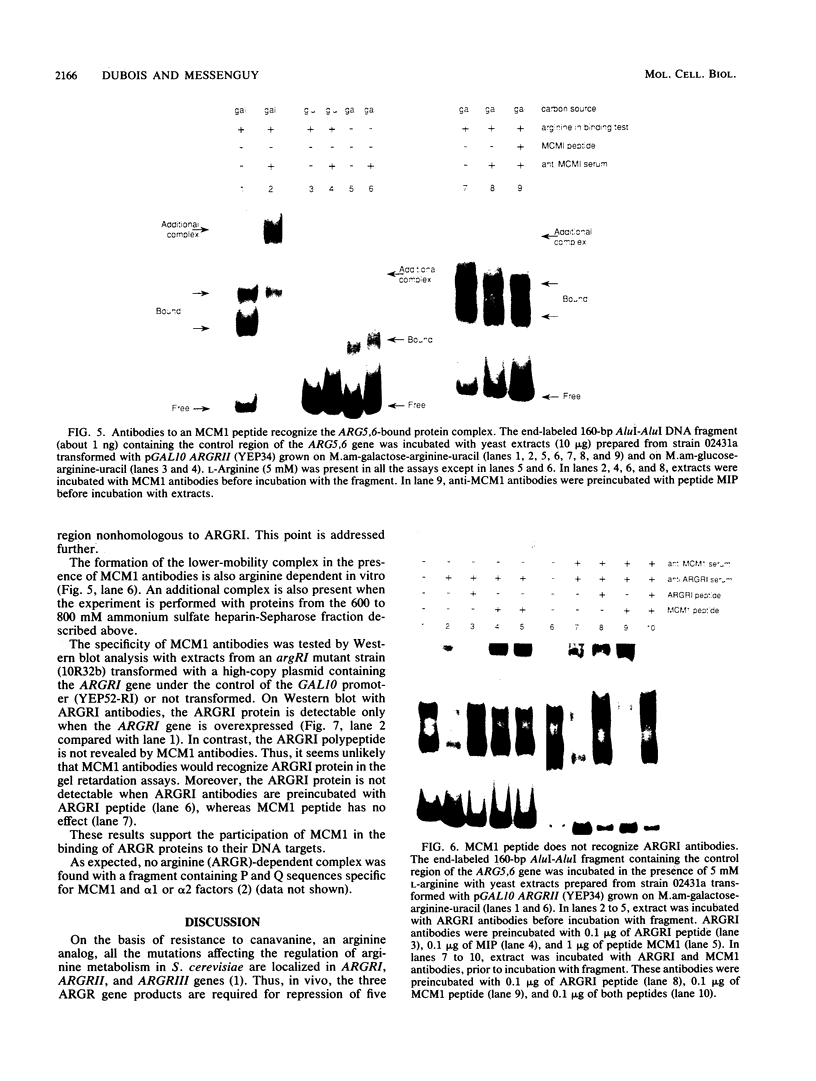
ARGRI, ARGRII, and ARGRIII regulatory proteins control the expression of arginine anabolic and catabolic genes in Saccharomyces cerevisiae. We show here that they are also required in vitro to observe a protein-DNA complex with the promoter of the ARG5,6 gene. The specific binding of ARGR proteins in vitro is stimulated by arginine. Antibodies raised against a synthetic MCM1 polypeptide retard the migration of ARGR-DNA complex on gel mobility shift assays. This result suggests that MCM1 could be an additional regulatory element of arginine metabolism.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bechet J., Greenson M., Wiame J. M. Mutations affecting the repressibility of arginine biosynthetic enzymes in Saccharomyces cerevisiae. Eur J Biochem. 1970 Jan;12(1):31–39. doi: 10.1111/j.1432-1033.1970.tb00817.x. [DOI] [PubMed] [Google Scholar]
- Bender A., Sprague G. F., Jr MAT alpha 1 protein, a yeast transcription activator, binds synergistically with a second protein to a set of cell-type-specific genes. Cell. 1987 Aug 28;50(5):681–691. doi: 10.1016/0092-8674(87)90326-6. [DOI] [PubMed] [Google Scholar]
- Bercy J., Dubois E., Messenguy F. Regulation of arginine metabolism in Saccharomyces cerevisiae: expression of the three ARGR regulatory genes and cellular localization of their products. Gene. 1987;55(2-3):277–285. doi: 10.1016/0378-1119(87)90287-3. [DOI] [PubMed] [Google Scholar]
- Crabeel M., Huygen R., Verschueren K., Messenguy F., Tinel K., Cunin R., Glansdorff N. General amino acid control and specific arginine repression in Saccharomyces cerevisiae: physical study of the bifunctional regulatory region of the ARG3 gene. Mol Cell Biol. 1985 Nov;5(11):3139–3148. doi: 10.1128/mcb.5.11.3139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deschamps J., Dubois E., Wiame J. M. L-Ornithine transaminase synthesis in Saccharomyces cerevisiae: regulation by inducer exclusion. Mol Gen Genet. 1979 Jul 24;174(3):225–232. doi: 10.1007/BF00267794. [DOI] [PubMed] [Google Scholar]
- Dubois E., Bercy J., Descamps F., Messenguy F. Characterization of two new genes essential for vegetative growth in Saccharomyces cerevisiae: nucleotide sequence determination and chromosome mapping. Gene. 1987;55(2-3):265–275. doi: 10.1016/0378-1119(87)90286-1. [DOI] [PubMed] [Google Scholar]
- Dubois E., Bercy J., Messenguy F. Characterization of two genes, ARGRI and ARGRIII required for specific regulation of arginine metabolism in yeast. Mol Gen Genet. 1987 Apr;207(1):142–148. doi: 10.1007/BF00331501. [DOI] [PubMed] [Google Scholar]
- Dubois E., Hiernaux D., Grennon M., Wiame J. M. Specific induction of catabolism and its relation to repression of biosynthesis in arginine metabolism of Saccharomyces cerevisiae. J Mol Biol. 1978 Jul 15;122(4):383–406. doi: 10.1016/0022-2836(78)90417-5. [DOI] [PubMed] [Google Scholar]
- Dubois E., Messenguy F. Isolation and characterization of the yeast ARGRII gene involved in regulating both anabolism and catabolism of arginine. Mol Gen Genet. 1985;198(2):283–289. doi: 10.1007/BF00383008. [DOI] [PubMed] [Google Scholar]
- Hinnen A., Hicks J. B., Fink G. R. Transformation of yeast. Proc Natl Acad Sci U S A. 1978 Apr;75(4):1929–1933. doi: 10.1073/pnas.75.4.1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jarvis E. E., Clark K. L., Sprague G. F., Jr The yeast transcription activator PRTF, a homolog of the mammalian serum response factor, is encoded by the MCM1 gene. Genes Dev. 1989 Jul;3(7):936–945. doi: 10.1101/gad.3.7.936. [DOI] [PubMed] [Google Scholar]
- Keleher C. A., Goutte C., Johnson A. D. The yeast cell-type-specific repressor alpha 2 acts cooperatively with a non-cell-type-specific protein. Cell. 1988 Jun 17;53(6):927–936. doi: 10.1016/s0092-8674(88)90449-7. [DOI] [PubMed] [Google Scholar]
- Lee A. C., Powell J. E., Tregear G. W., Niall H. D., Stevens V. C. A method for preparing beta-hCG COOH peptide-carrier conjugates of predictable composition. Mol Immunol. 1980 Jun;17(6):749–756. doi: 10.1016/0161-5890(80)90145-5. [DOI] [PubMed] [Google Scholar]
- Messenguy F., Dubois E. Participation of transcriptional and post-transcriptional regulatory mechanisms in the control of arginine metabolism in yeast. Mol Gen Genet. 1983;189(1):148–156. doi: 10.1007/BF00326068. [DOI] [PubMed] [Google Scholar]
- Messenguy F., Dubois E. The yeast ARGRII regulatory protein has homology with various RNases and DNA binding proteins. Mol Gen Genet. 1988 Jan;211(1):102–105. doi: 10.1007/BF00338399. [DOI] [PubMed] [Google Scholar]
- Messenguy F. Regulation of arginine biosynthesis in Saccharomyces cerevisiae: isolation of a cis-dominant, constitutive mutant for ornithine carbamoyltransferase synthesis. J Bacteriol. 1976 Oct;128(1):49–55. doi: 10.1128/jb.128.1.49-55.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norman C., Runswick M., Pollock R., Treisman R. Isolation and properties of cDNA clones encoding SRF, a transcription factor that binds to the c-fos serum response element. Cell. 1988 Dec 23;55(6):989–1003. doi: 10.1016/0092-8674(88)90244-9. [DOI] [PubMed] [Google Scholar]
- Passmore S., Maine G. T., Elble R., Christ C., Tye B. K. Saccharomyces cerevisiae protein involved in plasmid maintenance is necessary for mating of MAT alpha cells. J Mol Biol. 1988 Dec 5;204(3):593–606. doi: 10.1016/0022-2836(88)90358-0. [DOI] [PubMed] [Google Scholar]
- Pfeifer K., Arcangioli B., Guarente L. Yeast HAP1 activator competes with the factor RC2 for binding to the upstream activation site UAS1 of the CYC1 gene. Cell. 1987 Apr 10;49(1):9–18. doi: 10.1016/0092-8674(87)90750-1. [DOI] [PubMed] [Google Scholar]
- Portetelle D., Dandoy C., Burny A., Zavada J., Siakkou H., Gras-Masse H., Drobecq H., Tartar A. Synthetic peptides approach to identification of epitopes on bovine leukemia virus envelope glycoprotein gp51. Virology. 1989 Mar;169(1):34–41. doi: 10.1016/0042-6822(89)90038-x. [DOI] [PubMed] [Google Scholar]
- Qiu H. F., Dubois E., Broën P., Messenguy F. Functional analysis of ARGRI and ARGRIII regulatory proteins involved in the regulation of arginine metabolism in Saccharomyces cerevisiae. Mol Gen Genet. 1990 Jul;222(2-3):192–200. doi: 10.1007/BF00633817. [DOI] [PubMed] [Google Scholar]
- Qui H. F., Dubois E., Messenguy F. Dissection of the bifunctional ARGRII protein involved in the regulation of arginine anabolic and catabolic pathways. Mol Cell Biol. 1991 Apr;11(4):2169–2179. doi: 10.1128/mcb.11.4.2169. [DOI] [PMC free article] [PubMed] [Google Scholar]









