Abstract
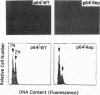
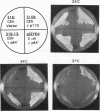
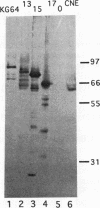
Yeast centromere DNA (CEN) affinity column chromatography has been used to purify several putative centromere and kinetochore proteins from yeast chromatin extracts. The single yeast gene (CBF5) specifying one of the major low-affinity centromere-binding proteins (p64'/CBF5p) has been cloned and shown to be essential for viability of Saccharomyces cerevisiae. CBF5 specifies a 55-kDa highly charged protein that contains a repeating KKD/E sequence domain near the C terminus, similar to known microtubule-binding domains in microtubule-associated proteins 1A and 1B, CBF5p, obtained by overexpression in bacterial cells, binds microtubules in vitro, whereas C-terminal deleted proteins lacking the (KKD/E)n domain do not. Dividing yeast cells containing a C-terminal truncated CBF5 gene, producing CBF5p containing only three copies of the KKD/E repeat, delay with replicated genomes at the G2/M phase of the cell cycle, while depletion of CBF5p arrests most cells in G1/S. Overproduction of CBF5p in S. cerevisiae complements a temperature sensitivity mutation in the gene (CBF2) specifying the 110-kDa subunit of the high-affinity CEN DNA-binding factor CBF3, suggesting in vivo interaction of CBF5p and CBF3. A second low-affinity centromere-binding factor has been identified as topoisomerase II.
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allen R. D., Weiss D. G., Hayden J. H., Brown D. T., Fujiwake H., Simpson M. Gliding movement of and bidirectional transport along single native microtubules from squid axoplasm: evidence for an active role of microtubules in cytoplasmic transport. J Cell Biol. 1985 May;100(5):1736–1752. doi: 10.1083/jcb.100.5.1736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker R. E., Fitzgerald-Hayes M., O'Brien T. C. Purification of the yeast centromere binding protein CP1 and a mutational analysis of its binding site. J Biol Chem. 1989 Jun 25;264(18):10843–10850. [PubMed] [Google Scholar]
- Baker R. E., Masison D. C. Isolation of the gene encoding the Saccharomyces cerevisiae centromere-binding protein CP1. Mol Cell Biol. 1990 Jun;10(6):2458–2467. doi: 10.1128/mcb.10.6.2458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M. J., Davis R. W. Purification of a yeast centromere-binding protein that is able to distinguish single base-pair mutations in its recognition site. Mol Cell Biol. 1989 Jun;9(6):2544–2550. doi: 10.1128/mcb.9.6.2544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cai M., Davis R. W. Yeast centromere binding protein CBF1, of the helix-loop-helix protein family, is required for chromosome stability and methionine prototrophy. Cell. 1990 May 4;61(3):437–446. doi: 10.1016/0092-8674(90)90525-j. [DOI] [PubMed] [Google Scholar]
- Carle G. F., Olson M. V. An electrophoretic karyotype for yeast. Proc Natl Acad Sci U S A. 1985 Jun;82(11):3756–3760. doi: 10.1073/pnas.82.11.3756. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarke L., Carbon J. Isolation of a yeast centromere and construction of functional small circular chromosomes. Nature. 1980 Oct 9;287(5782):504–509. doi: 10.1038/287504a0. [DOI] [PubMed] [Google Scholar]
- Clarke L. Centromeres of budding and fission yeasts. Trends Genet. 1990 May;6(5):150–154. doi: 10.1016/0168-9525(90)90149-z. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giaever G., Lynn R., Goto T., Wang J. C. The complete nucleotide sequence of the structural gene TOP2 of yeast DNA topoisomerase II. J Biol Chem. 1986 Sep 25;261(27):12448–12454. [PubMed] [Google Scholar]
- Goh P. Y., Kilmartin J. V. NDC10: a gene involved in chromosome segregation in Saccharomyces cerevisiae. J Cell Biol. 1993 May;121(3):503–512. doi: 10.1083/jcb.121.3.503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan K. L., Dixon J. E. Eukaryotic proteins expressed in Escherichia coli: an improved thrombin cleavage and purification procedure of fusion proteins with glutathione S-transferase. Anal Biochem. 1991 Feb 1;192(2):262–267. doi: 10.1016/0003-2697(91)90534-z. [DOI] [PubMed] [Google Scholar]
- Holm C., Goto T., Wang J. C., Botstein D. DNA topoisomerase II is required at the time of mitosis in yeast. Cell. 1985 Jun;41(2):553–563. doi: 10.1016/s0092-8674(85)80028-3. [DOI] [PubMed] [Google Scholar]
- Hyman A. A., Middleton K., Centola M., Mitchison T. J., Carbon J. Microtubule-motor activity of a yeast centromere-binding protein complex. Nature. 1992 Oct 8;359(6395):533–536. doi: 10.1038/359533a0. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang W. D., Philippsen P. Purification of a protein binding to the CDEI subregion of Saccharomyces cerevisiae centromere DNA. Mol Cell Biol. 1989 Dec;9(12):5585–5593. doi: 10.1128/mcb.9.12.5585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jiang W., Lechner J., Carbon J. Isolation and characterization of a gene (CBF2) specifying a protein component of the budding yeast kinetochore. J Cell Biol. 1993 May;121(3):513–519. doi: 10.1083/jcb.121.3.513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Langkopf A., Hammarback J. A., Müller R., Vallee R. B., Garner C. C. Microtubule-associated proteins 1A and LC2. Two proteins encoded in one messenger RNA. J Biol Chem. 1992 Aug 15;267(23):16561–16566. [PubMed] [Google Scholar]
- Lechner J., Carbon J. A 240 kd multisubunit protein complex, CBF3, is a major component of the budding yeast centromere. Cell. 1991 Feb 22;64(4):717–725. doi: 10.1016/0092-8674(91)90501-o. [DOI] [PubMed] [Google Scholar]
- Masumoto H., Masukata H., Muro Y., Nozaki N., Okazaki T. A human centromere antigen (CENP-B) interacts with a short specific sequence in alphoid DNA, a human centromeric satellite. J Cell Biol. 1989 Nov;109(5):1963–1973. doi: 10.1083/jcb.109.5.1963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mellor J., Jiang W., Funk M., Rathjen J., Barnes C. A., Hinz T., Hegemann J. H., Philippsen P. CPF1, a yeast protein which functions in centromeres and promoters. EMBO J. 1990 Dec;9(12):4017–4026. doi: 10.1002/j.1460-2075.1990.tb07623.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Neff M. W., Burke D. J. A delay in the Saccharomyces cerevisiae cell cycle that is induced by a dicentric chromosome and dependent upon mitotic checkpoints. Mol Cell Biol. 1992 Sep;12(9):3857–3864. doi: 10.1128/mcb.12.9.3857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ng R., Carbon J. Mutational and in vitro protein-binding studies on centromere DNA from Saccharomyces cerevisiae. Mol Cell Biol. 1987 Dec;7(12):4522–4534. doi: 10.1128/mcb.7.12.4522. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ngsee J. K., Scheller R. H. Isolation and characterization of two homologous cDNA clones from Torpedo electromotor neurons. DNA. 1989 Oct;8(8):555–561. doi: 10.1089/dna.1989.8.555. [DOI] [PubMed] [Google Scholar]
- Noble M., Lewis S. A., Cowan N. J. The microtubule binding domain of microtubule-associated protein MAP1B contains a repeated sequence motif unrelated to that of MAP2 and tau. J Cell Biol. 1989 Dec;109(6 Pt 2):3367–3376. doi: 10.1083/jcb.109.6.3367. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pluta A. F., Saitoh N., Goldberg I., Earnshaw W. C. Identification of a subdomain of CENP-B that is necessary and sufficient for localization to the human centromere. J Cell Biol. 1992 Mar;116(5):1081–1093. doi: 10.1083/jcb.116.5.1081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Périer F., Carbon J. A colony color assay for Saccharomyces cerevisiae mutants defective in kinetochore structure and function. Genetics. 1992 Sep;132(1):39–51. doi: 10.1093/genetics/132.1.39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose D., Thomas W., Holm C. Segregation of recombined chromosomes in meiosis I requires DNA topoisomerase II. Cell. 1990 Mar 23;60(6):1009–1017. doi: 10.1016/0092-8674(90)90349-j. [DOI] [PubMed] [Google Scholar]
- Sawin K. E., Scholey J. M. Motor proteins in cell division. Trends Cell Biol. 1991 Nov;1(5):122–129. doi: 10.1016/0962-8924(91)90117-r. [DOI] [PubMed] [Google Scholar]
- Sikorski R. S., Hieter P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics. 1989 May;122(1):19–27. doi: 10.1093/genetics/122.1.19. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spencer F., Hieter P. Centromere DNA mutations induce a mitotic delay in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1992 Oct 1;89(19):8908–8912. doi: 10.1073/pnas.89.19.8908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Triglia T., Stahl H. D., Crewther P. E., Silva A., Anders R. F., Kemp D. J. Structure of a Plasmodium falciparum gene that encodes a glutamic acid-rich protein (GARP). Mol Biochem Parasitol. 1988 Nov;31(2):199–201. doi: 10.1016/0166-6851(88)90170-3. [DOI] [PubMed] [Google Scholar]
- Yen T. J., Li G., Schaar B. T., Szilak I., Cleveland D. W. CENP-E is a putative kinetochore motor that accumulates just before mitosis. Nature. 1992 Oct 8;359(6395):536–539. doi: 10.1038/359536a0. [DOI] [PubMed] [Google Scholar]