Abstract
While inverted DNA repeats are generally acknowledged to be an important source of genetic instability in prokaryotes, relatively little is known about their effects in eukaryotes. Using bacterial transposon Tn5 and its derivatives, we demonstrate that long inverted repeats also cause genetic instability leading to deletion in the yeast Saccharomyces cerevisiae. Furthermore, they induce homologous recombination. Replication plays a major role in the deletion formation. Deletions are stimulated by a mutation in the DNA polymerase delta gene (pol3). The majority of deletions result from imprecise excision between small (4- to 6-bp) repeats in a polar fashion, and they often generate quasipalindrome structures that subsequently may be highly unstable. Breakpoints are clustered near the ends of the long inverted repeats (< 150 bp). The repeats have both intra- and interchromosomal effects in that they also create hot spots for mitotic interchromosomal recombination. Intragenic recombination is 4 to 18 times more frequent for heteroalleles in which one of the two mutations is due to the insertion of a long inverted repeat, compared with other pairs of heteroalleles in which neither mutation has a long repeat. We propose that both deletion and recombination are the result of altered replication at the basal part of the stem formed by the inverted repeats.
Full text
PDF
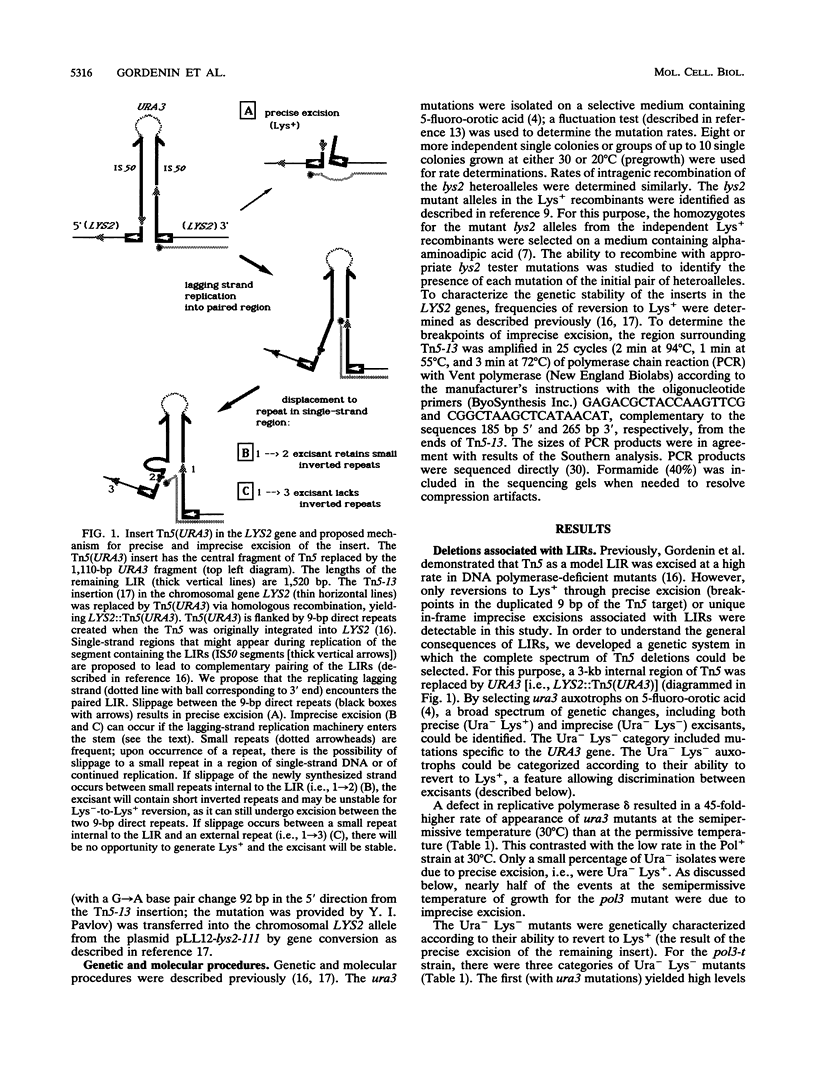





Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aguilera A., Klein H. L. Genetic control of intrachromosomal recombination in Saccharomyces cerevisiae. I. Isolation and genetic characterization of hyper-recombination mutations. Genetics. 1988 Aug;119(4):779–790. doi: 10.1093/genetics/119.4.779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Auerswald E. A., Ludwig G., Schaller H. Structural analysis of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):107–113. doi: 10.1101/sqb.1981.045.01.019. [DOI] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Bonneaud N., Ozier-Kalogeropoulos O., Li G. Y., Labouesse M., Minvielle-Sebastia L., Lacroute F. A family of low and high copy replicative, integrative and single-stranded S. cerevisiae/E. coli shuttle vectors. Yeast. 1991 Aug-Sep;7(6):609–615. doi: 10.1002/yea.320070609. [DOI] [PubMed] [Google Scholar]
- Burgers P. M., Bambara R. A., Campbell J. L., Chang L. M., Downey K. M., Hübscher U., Lee M. Y., Linn S. M., So A. G., Spadari S. Revised nomenclature for eukaryotic DNA polymerases. Eur J Biochem. 1990 Aug 17;191(3):617–618. doi: 10.1111/j.1432-1033.1990.tb19165.x. [DOI] [PubMed] [Google Scholar]
- Chattoo B. B., Sherman F., Azubalis D. A., Fjellstedt T. A., Mehnert D., Ogur M. Selection of lys2 Mutants of the Yeast SACCHAROMYCES CEREVISIAE by the Utilization of alpha-AMINOADIPATE. Genetics. 1979 Sep;93(1):51–65. doi: 10.1093/genetics/93.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chernov Iu O., Gordenin D. A. Fenotipicheskoe proiavlenie insertsii drozhzhevogo transpozona v gene LYS2 i deletsii, voznikaiushchikh pri netochnom vyrezanii transpozona. Genetika. 1987 Jan;23(1):30–40. [PubMed] [Google Scholar]
- Collins J. Instability of palindromic DNA in Escherichia coli. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):409–416. doi: 10.1101/sqb.1981.045.01.055. [DOI] [PubMed] [Google Scholar]
- Collins J., Volckaert G., Nevers P. Precise and nearly-precise excision of the symmetrical inverted repeats of Tn5; common features of recA-independent deletion events in Escherichia coli. Gene. 1982 Jul-Aug;19(1):139–146. doi: 10.1016/0378-1119(82)90198-6. [DOI] [PubMed] [Google Scholar]
- Drake J. W. A constant rate of spontaneous mutation in DNA-based microbes. Proc Natl Acad Sci U S A. 1991 Aug 15;88(16):7160–7164. doi: 10.1073/pnas.88.16.7160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Formosa T., Alberts B. M. DNA synthesis dependent on genetic recombination: characterization of a reaction catalyzed by purified bacteriophage T4 proteins. Cell. 1986 Dec 5;47(5):793–806. doi: 10.1016/0092-8674(86)90522-2. [DOI] [PubMed] [Google Scholar]
- Gordenin D. A., Malkova A. L., Peterzen A., Kulikov V. N., Pavlov Y. I., Perkins E., Resnick M. A. Transposon Tn5 excision in yeast: influence of DNA polymerases alpha, delta, and epsilon and repair genes. Proc Natl Acad Sci U S A. 1992 May 1;89(9):3785–3789. doi: 10.1073/pnas.89.9.3785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordenin D. A., Proscyavichus Y. Y., Malkova A. L., Trofimova M. V., Peterzen A. Yeast mutants with increased bacterial transposon Tn5 excision. Yeast. 1991 Jan;7(1):37–50. doi: 10.1002/yea.320070105. [DOI] [PubMed] [Google Scholar]
- Gordenin D. A., Trofimova M. V., Shaburova O. N., Pavlov Y. I., Chernoff Y. O., Chekuolene Y. V., Proscyavichus Y. Y., Sasnauskas K. V., Janulaitis A. A. Precise excision of bacterial transposon Tn5 in yeast. Mol Gen Genet. 1988 Aug;213(2-3):388–393. doi: 10.1007/BF00339607. [DOI] [PubMed] [Google Scholar]
- Henderson S. T., Petes T. D. Instability of a plasmid-borne inverted repeat in Saccharomyces cerevisiae. Genetics. 1993 May;134(1):57–62. doi: 10.1093/genetics/134.1.57. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris M. E., Jinks-Robertson S. Nucleotide sequence of the LYS2 gene of Saccharomyces cerevisiae: homology to Bacillus brevis tyrocidine synthetase 1. Gene. 1991 Feb 1;98(1):141–145. doi: 10.1016/0378-1119(91)90117-t. [DOI] [PubMed] [Google Scholar]
- Morrison A., Araki H., Clark A. B., Hamatake R. K., Sugino A. A third essential DNA polymerase in S. cerevisiae. Cell. 1990 Sep 21;62(6):1143–1151. doi: 10.1016/0092-8674(90)90391-q. [DOI] [PubMed] [Google Scholar]
- Neil D. L., Villasante A., Fisher R. B., Vetrie D., Cox B., Tyler-Smith C. Structural instability of human tandemly repeated DNA sequences cloned in yeast artificial chromosome vectors. Nucleic Acids Res. 1990 Mar 25;18(6):1421–1428. doi: 10.1093/nar/18.6.1421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Noskov V. N., Tarutina M. G., Pavlov Iu I., Kulikov V. N., Trofimova M. V., Gorbacheva A. V., Chernov Iu O., Sasnauskas K. V., Neistat M. A., Tolstorukov I. I. Razrabotka sistemy vnutrigennogo kartirovaniia dlia molekuliarno- geneticheskogo analiza mutatsii v gene LYS2 u drozhzhei sakharomitsetov. Genetika. 1990 Jul;26(7):1161–1168. [PubMed] [Google Scholar]
- Ohshima A., Inouye S., Inouye M. In vivo duplication of genetic elements by the formation of stem-loop DNA without an RNA intermediate. Proc Natl Acad Sci U S A. 1992 Feb 1;89(3):1016–1020. doi: 10.1073/pnas.89.3.1016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruskin B., Fink G. R. Mutations in POL1 increase the mitotic instability of tandem inverted repeats in Saccharomyces cerevisiae. Genetics. 1993 May;134(1):43–56. doi: 10.1093/genetics/134.1.43. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tindall K. R., Stankowski L. F., Jr Molecular analysis of spontaneous mutations at the gpt locus in Chinese hamster ovary (AS52) cells. Mutat Res. 1989 Mar-May;220(2-3):241–253. doi: 10.1016/0165-1110(89)90028-6. [DOI] [PubMed] [Google Scholar]
- Trinh T. Q., Sinden R. R. Preferential DNA secondary structure mutagenesis in the lagging strand of replication in E. coli. Nature. 1991 Aug 8;352(6335):544–547. doi: 10.1038/352544a0. [DOI] [PubMed] [Google Scholar]
- Vincent A., Petes T. D. Mitotic and meiotic gene conversion of Ty elements and other insertions in Saccharomyces cerevisiae. Genetics. 1989 Aug;122(4):759–772. doi: 10.1093/genetics/122.4.759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Voelkel-Meiman K., Roeder G. S. A chromosome containing HOT1 preferentially receives information during mitotic interchromosomal gene conversion. Genetics. 1990 Mar;124(3):561–572. doi: 10.1093/genetics/124.3.561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang T. S. Eukaryotic DNA polymerases. Annu Rev Biochem. 1991;60:513–552. doi: 10.1146/annurev.bi.60.070191.002501. [DOI] [PubMed] [Google Scholar]



