Abstract
The site-specific recombinases Flp and R from Saccharomyces cerevisiae and Zygosaccharomyces rouxii, respectively, are related proteins that belong to the yeast family of site-specific recombinases. They share approximately 30% amino acid matches and exhibit a common reaction mechanism that appears to be conserved within the larger integrase family of site-specific recombinases. Two regions of the proteins, designated box I and box II, also harbor a significantly high degree of homology at the nucleotide sequence level. We have analyzed the properties of Flp and R variants carrying point mutations within the box I segment in substrate-binding, DNA cleavage, and full-site and half-site strand transfer reactions. All mutations abolish or seriously diminish recombinase function either at the substrate-binding step or at the catalytic steps of strand cleavage or strand transfer. Of particular interest are mutations of Arg-191 of Flp and R, residues which correspond to one of the two invariant arginine residues of the integrase family. These variant proteins bind substrate with affinities comparable to those of the corresponding wild-type recombinases. Among the binding-competent variants, only Flp(R191K) is capable of efficient substrate cleavage in a full recombination target. However, this protein does not cleave a half recombination site and fails to complete strand exchange in a full site. Strikingly, the Arg-191 mutants of Flp and R can be rescued in half-site strand transfer reactions by a second point mutant of the corresponding recombinase that lacks its active-site tyrosine (Tyr-343). Similarly, Flp and R variants of Cys-189 and Flp variants at Asp-194 and Asp-199 can also be complemented by the corresponding Tyr-343-to-phenylalanine recombinase mutant.
Full text
PDF



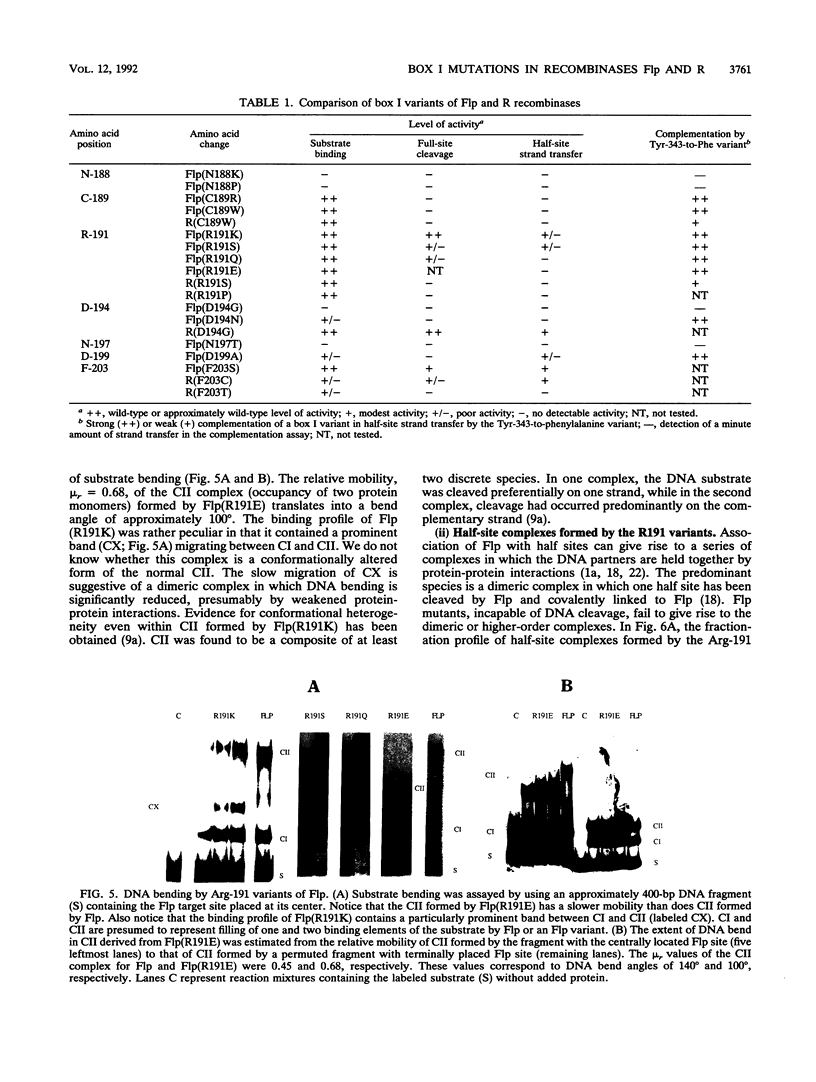




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abremski K. E., Hoess R. H. Evidence for a second conserved arginine residue in the integrase family of recombination proteins. Protein Eng. 1992 Jan;5(1):87–91. doi: 10.1093/protein/5.1.87. [DOI] [PubMed] [Google Scholar]
- Amin A., Roca H., Luetke K., Sadowski P. D. Synapsis, strand scission, and strand exchange induced by the FLP recombinase: analysis with half-FRT sites. Mol Cell Biol. 1991 Sep;11(9):4497–4508. doi: 10.1128/mcb.11.9.4497. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Araki H., Nakanishi N., Evans B. R., Matsuzaki H., Jayaram M., Oshima Y. Site-specific recombinase, R, encoded by yeast plasmid pSR1. J Mol Biol. 1992 May 5;225(1):25–37. doi: 10.1016/0022-2836(92)91023-i. [DOI] [PubMed] [Google Scholar]
- Argos P., Landy A., Abremski K., Egan J. B., Haggard-Ljungquist E., Hoess R. H., Kahn M. L., Kalionis B., Narayana S. V., Pierson L. S., 3rd The integrase family of site-specific recombinases: regional similarities and global diversity. EMBO J. 1986 Feb;5(2):433–440. doi: 10.1002/j.1460-2075.1986.tb04229.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J. W., Evans B. R., Yang S. H., Teplow D. B., Jayaram M. Domain of a yeast site-specific recombinase (Flp) that recognizes its target site. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):5944–5948. doi: 10.1073/pnas.88.14.5944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J. W., Evans B. R., Zheng L., Jayaram M. Tyr60 variants of Flp recombinase generate conformationally altered protein-DNA complexes. Differential activity in full-site and half-site recombinations. J Mol Biol. 1991 Mar 5;218(1):107–118. doi: 10.1016/0022-2836(91)90877-9. [DOI] [PubMed] [Google Scholar]
- Chen J. W., Lee J., Jayaram M. DNA cleavage in trans by the active site tyrosine during Flp recombination: switching protein partners before exchanging strands. Cell. 1992 May 15;69(4):647–658. doi: 10.1016/0092-8674(92)90228-5. [DOI] [PubMed] [Google Scholar]
- Evans B. R., Chen J. W., Parsons R. L., Bauer T. K., Teplow D. B., Jayaram M. Identification of the active site tyrosine of Flp recombinase. Possible relevance of its location to the mechanism of recombination. J Biol Chem. 1990 Oct 25;265(30):18504–18510. [PubMed] [Google Scholar]
- Friesen H., Sadowski P. D. Mutagenesis of a conserved region of the gene encoding the FLP recombinase of Saccharomyces cerevisiae. A role for arginine 191 in binding and ligation. J Mol Biol. 1992 May 20;225(2):313–326. doi: 10.1016/0022-2836(92)90924-9. [DOI] [PubMed] [Google Scholar]
- Kramer W., Drutsa V., Jansen H. W., Kramer B., Pflugfelder M., Fritz H. J. The gapped duplex DNA approach to oligonucleotide-directed mutation construction. Nucleic Acids Res. 1984 Dec 21;12(24):9441–9456. doi: 10.1093/nar/12.24.9441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsunami N., Hamaguchi Y., Yamamoto Y., Kuze K., Kangawa K., Matsuo H., Kawaichi M., Honjo T. A protein binding to the J kappa recombination sequence of immunoglobulin genes contains a sequence related to the integrase motif. Nature. 1989 Dec 21;342(6252):934–937. doi: 10.1038/342934a0. [DOI] [PubMed] [Google Scholar]
- Messing J., Crea R., Seeburg P. H. A system for shotgun DNA sequencing. Nucleic Acids Res. 1981 Jan 24;9(2):309–321. doi: 10.1093/nar/9.2.309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pan H., Clary D., Sadowski P. D. Identification of the DNA-binding domain of the FLP recombinase. J Biol Chem. 1991 Jun 15;266(17):11347–11354. [PubMed] [Google Scholar]
- Parsons R. L., Evans B. R., Zheng L., Jayaram M. Functional analysis of Arg-308 mutants of Flp recombinase. Possible role of Arg-308 in coupling substrate binding to catalysis. J Biol Chem. 1990 Mar 15;265(8):4527–4533. [PubMed] [Google Scholar]
- Parsons R. L., Prasad P. V., Harshey R. M., Jayaram M. Step-arrest mutants of FLP recombinase: implications for the catalytic mechanism of DNA recombination. Mol Cell Biol. 1988 Aug;8(8):3303–3310. doi: 10.1128/mcb.8.8.3303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prasad P. V., Young L. J., Jayaram M. Mutations in the 2-microns circle site-specific recombinase that abolish recombination without affecting substrate recognition. Proc Natl Acad Sci U S A. 1987 Apr;84(8):2189–2193. doi: 10.1073/pnas.84.8.2189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qian X. H., Inman R. B., Cox M. M. Protein-based asymmetry and protein-protein interactions in FLP recombinase-mediated site-specific recombination. J Biol Chem. 1990 Dec 15;265(35):21779–21788. [PubMed] [Google Scholar]
- Schwartz C. J., Sadowski P. D. FLP protein of 2 mu circle plasmid of yeast induces multiple bends in the FLP recognition target site. J Mol Biol. 1990 Nov 20;216(2):289–298. doi: 10.1016/s0022-2836(05)80320-1. [DOI] [PubMed] [Google Scholar]
- Senecoff J. F., Bruckner R. C., Cox M. M. The FLP recombinase of the yeast 2-micron plasmid: characterization of its recombination site. Proc Natl Acad Sci U S A. 1985 Nov;82(21):7270–7274. doi: 10.1073/pnas.82.21.7270. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson J. F., Landy A. Empirical estimation of protein-induced DNA bending angles: applications to lambda site-specific recombination complexes. Nucleic Acids Res. 1988 Oct 25;16(20):9687–9705. doi: 10.1093/nar/16.20.9687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Utatsu I., Sakamoto S., Imura T., Toh-e A. Yeast plasmids resembling 2 micron DNA: regional similarities and diversities at the molecular level. J Bacteriol. 1987 Dec;169(12):5537–5545. doi: 10.1128/jb.169.12.5537-5545.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Volkert F. C., Wilson D. W., Broach J. R. Deoxyribonucleic acid plasmids in yeasts. Microbiol Rev. 1989 Sep;53(3):299–317. doi: 10.1128/mr.53.3.299-317.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]