Abstract
rad5 (rev2) mutants of Saccharomyces cerevisiae are sensitive to UV light and other DNA-damaging agents, and RAD5 is in the RAD6 epistasis group of DNA repair genes. To unambiguously define the function of RAD5, we have cloned the RAD5 gene, determined the effects of the rad5 deletion mutation on DNA repair, DNA damage-induced mutagenesis, and other cellular processes, and analyzed the sequence of RAD5-encoded protein. Our genetic studies indicate that RAD5 functions primarily with RAD18 in error-free postreplication repair. We also show that RAD5 affects the rate of instability of poly(GT) repeat sequences. Genomic poly(GT) sequences normally change length at a rate of about 10(-4); this rate is approximately 10-fold lower in the rad5 deletion mutant than in the corresponding isogenic wild-type strain. RAD5 encodes a protein of 1,169 amino acids of M(r) 134,000, and it contains several interesting sequence motifs. All seven conserved domains found associated with DNA helicases are present in RAD5. RAD5 also contains a cysteine-rich sequence motif that resembles the corresponding sequences found in 11 other proteins, including those encoded by the DNA repair gene RAD18 and the RAG1 gene required for immunoglobin gene arrangement. A leucine zipper motif preceded by a basic region is also present in RAD5. The cysteine-rich region may coordinate the binding of zinc; this region and the basic segment might constitute distinct DNA-binding domains in RAD5. Possible roles of RAD5 putative ATPase/DNA helicase activity in DNA repair and in the maintenance of wild-type rates of instability of simple repetitive sequences are discussed.
Full text
PDF

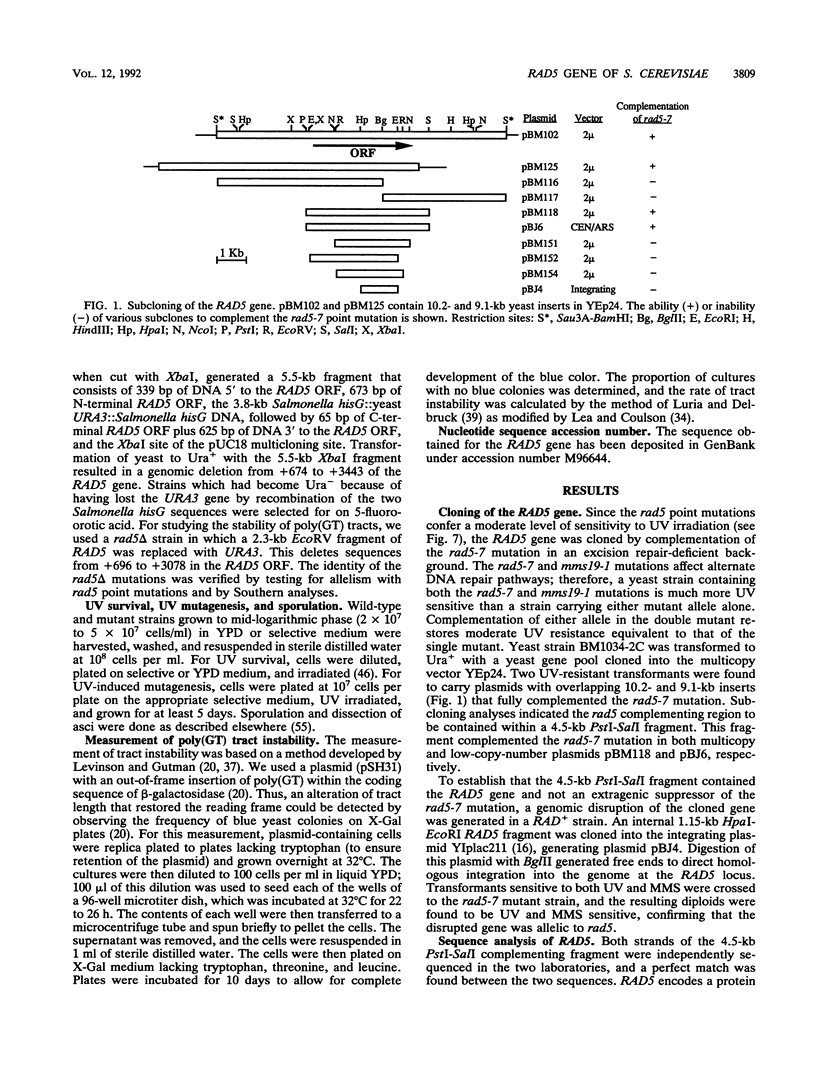



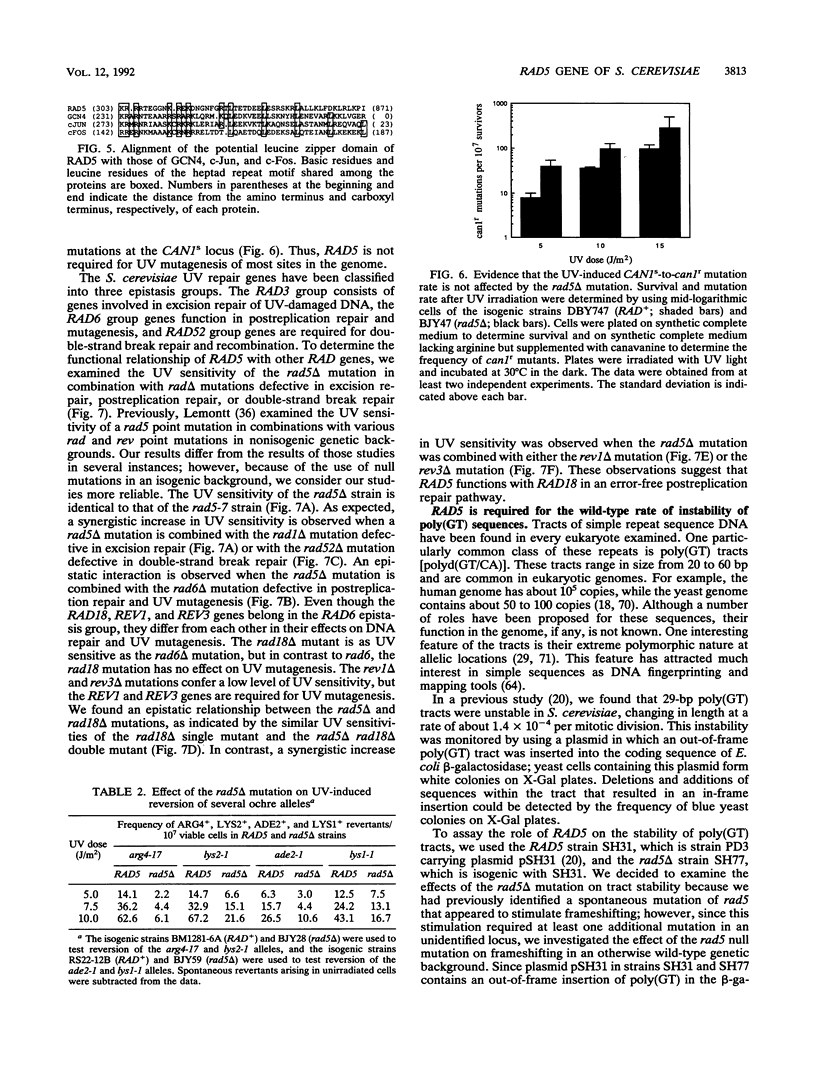
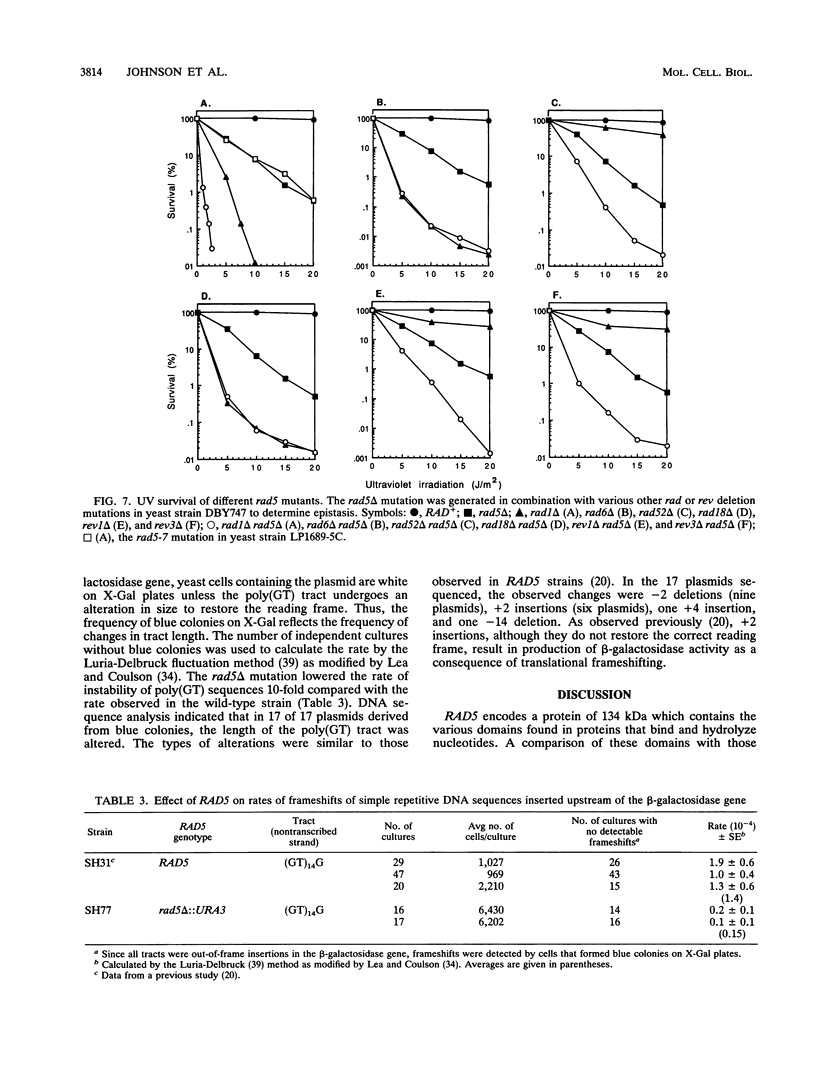
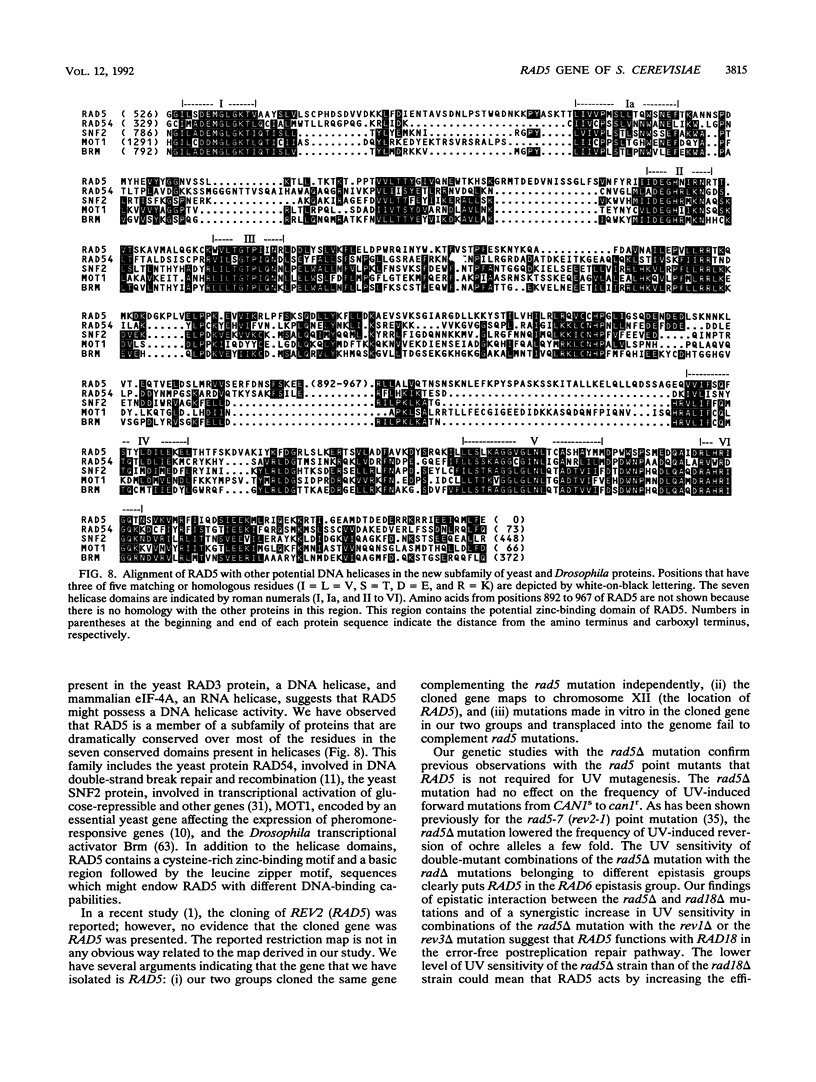



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ahne F., Wendel S., Eckardt-Schupp F. Molecular analysis of the REV2 gene of Saccharomyces cerevisiae--a review. Radiat Environ Biophys. 1990;29(4):293–301. doi: 10.1007/BF01210409. [DOI] [PubMed] [Google Scholar]
- Alani E., Cao L., Kleckner N. A method for gene disruption that allows repeated use of URA3 selection in the construction of multiply disrupted yeast strains. Genetics. 1987 Aug;116(4):541–545. doi: 10.1534/genetics.112.541.test. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrews B. J. Gene expression. Dialogue with the cell cycle. Nature. 1992 Jan 30;355(6359):393–394. doi: 10.1038/355393a0. [DOI] [PubMed] [Google Scholar]
- Bailly V., Sung P., Prakash L., Prakash S. DNA.RNA helicase activity of RAD3 protein of Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9712–9716. doi: 10.1073/pnas.88.21.9712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Botstein D., Falco S. C., Stewart S. E., Brennan M., Scherer S., Stinchcomb D. T., Struhl K., Davis R. W. Sterile host yeasts (SHY): a eukaryotic system of biological containment for recombinant DNA experiments. Gene. 1979 Dec;8(1):17–24. doi: 10.1016/0378-1119(79)90004-0. [DOI] [PubMed] [Google Scholar]
- Carlson M., Botstein D. Two differentially regulated mRNAs with different 5' ends encode secreted with intracellular forms of yeast invertase. Cell. 1982 Jan;28(1):145–154. doi: 10.1016/0092-8674(82)90384-1. [DOI] [PubMed] [Google Scholar]
- Davis J. L., Kunisawa R., Thorner J. A presumptive helicase (MOT1 gene product) affects gene expression and is required for viability in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1992 Apr;12(4):1879–1892. doi: 10.1128/mcb.12.4.1879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davison A. J., Scott J. E. The complete DNA sequence of varicella-zoster virus. J Gen Virol. 1986 Sep;67(Pt 9):1759–1816. doi: 10.1099/0022-1317-67-9-1759. [DOI] [PubMed] [Google Scholar]
- Emery H. S., Schild D., Kellogg D. E., Mortimer R. K. Sequence of RAD54, a Saccharomyces cerevisiae gene involved in recombination and repair. Gene. 1991 Jul 31;104(1):103–106. doi: 10.1016/0378-1119(91)90473-o. [DOI] [PubMed] [Google Scholar]
- Freemont P. S., Hanson I. M., Trowsdale J. A novel cysteine-rich sequence motif. Cell. 1991 Feb 8;64(3):483–484. doi: 10.1016/0092-8674(91)90229-r. [DOI] [PubMed] [Google Scholar]
- Fry D. C., Kuby S. A., Mildvan A. S. ATP-binding site of adenylate kinase: mechanistic implications of its homology with ras-encoded p21, F1-ATPase, and other nucleotide-binding proteins. Proc Natl Acad Sci U S A. 1986 Feb;83(4):907–911. doi: 10.1073/pnas.83.4.907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gentz R., Rauscher F. J., 3rd, Abate C., Curran T. Parallel association of Fos and Jun leucine zippers juxtaposes DNA binding domains. Science. 1989 Mar 31;243(4899):1695–1699. doi: 10.1126/science.2494702. [DOI] [PubMed] [Google Scholar]
- Gietz R. D., Sugino A. New yeast-Escherichia coli shuttle vectors constructed with in vitro mutagenized yeast genes lacking six-base pair restriction sites. Gene. 1988 Dec 30;74(2):527–534. doi: 10.1016/0378-1119(88)90185-0. [DOI] [PubMed] [Google Scholar]
- Gorbalenya A. E., Koonin E. V., Donchenko A. P., Blinov V. M. Two related superfamilies of putative helicases involved in replication, recombination, repair and expression of DNA and RNA genomes. Nucleic Acids Res. 1989 Jun 26;17(12):4713–4730. doi: 10.1093/nar/17.12.4713. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hamada H., Petrino M. G., Kakunaga T. A novel repeated element with Z-DNA-forming potential is widely found in evolutionarily diverse eukaryotic genomes. Proc Natl Acad Sci U S A. 1982 Nov;79(21):6465–6469. doi: 10.1073/pnas.79.21.6465. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattori K., Angel P., Le Beau M. M., Karin M. Structure and chromosomal localization of the functional intronless human JUN protooncogene. Proc Natl Acad Sci U S A. 1988 Dec;85(23):9148–9152. doi: 10.1073/pnas.85.23.9148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson S. T., Petes T. D. Instability of simple sequence DNA in Saccharomyces cerevisiae. Mol Cell Biol. 1992 Jun;12(6):2749–2757. doi: 10.1128/mcb.12.6.2749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hinnebusch A. G. Evidence for translational regulation of the activator of general amino acid control in yeast. Proc Natl Acad Sci U S A. 1984 Oct;81(20):6442–6446. doi: 10.1073/pnas.81.20.6442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hope I. A., Struhl K. Functional dissection of a eukaryotic transcriptional activator protein, GCN4 of yeast. Cell. 1986 Sep 12;46(6):885–894. doi: 10.1016/0092-8674(86)90070-x. [DOI] [PubMed] [Google Scholar]
- Hope I. A., Struhl K. GCN4, a eukaryotic transcriptional activator protein, binds as a dimer to target DNA. EMBO J. 1987 Sep;6(9):2781–2784. doi: 10.1002/j.1460-2075.1987.tb02573.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jentsch S., McGrath J. P., Varshavsky A. The yeast DNA repair gene RAD6 encodes a ubiquitin-conjugating enzyme. Nature. 1987 Sep 10;329(6135):131–134. doi: 10.1038/329131a0. [DOI] [PubMed] [Google Scholar]
- Jones J. S., Weber S., Prakash L. The Saccharomyces cerevisiae RAD18 gene encodes a protein that contains potential zinc finger domains for nucleic acid binding and a putative nucleotide binding sequence. Nucleic Acids Res. 1988 Jul 25;16(14B):7119–7131. doi: 10.1093/nar/16.14.7119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kakizuka A., Miller W. H., Jr, Umesono K., Warrell R. P., Jr, Frankel S. R., Murty V. V., Dmitrovsky E., Evans R. M. Chromosomal translocation t(15;17) in human acute promyelocytic leukemia fuses RAR alpha with a novel putative transcription factor, PML. Cell. 1991 Aug 23;66(4):663–674. doi: 10.1016/0092-8674(91)90112-c. [DOI] [PubMed] [Google Scholar]
- Kashi Y., Tikochinsky Y., Genislav E., Iraqi F., Nave A., Beckmann J. S., Gruenbaum Y., Soller M. Large restriction fragments containing poly-TG are highly polymorphic in a variety of vertebrates. Nucleic Acids Res. 1990 Mar 11;18(5):1129–1132. doi: 10.1093/nar/18.5.1129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Landschulz W. H., Johnson P. F., McKnight S. L. The leucine zipper: a hypothetical structure common to a new class of DNA binding proteins. Science. 1988 Jun 24;240(4860):1759–1764. doi: 10.1126/science.3289117. [DOI] [PubMed] [Google Scholar]
- Laurent B. C., Yang X., Carlson M. An essential Saccharomyces cerevisiae gene homologous to SNF2 encodes a helicase-related protein in a new family. Mol Cell Biol. 1992 Apr;12(4):1893–1902. doi: 10.1128/mcb.12.4.1893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence C. W., Christensen R. B. Ultraviolet-induced reversion of cyc1 alleles in radiation sensitive strains of yeast. II. rev2 mutant strains. Genetics. 1978 Oct;90(2):213–226. doi: 10.1093/genetics/90.2.213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lawrence C. W. Mutagenesis in Saccharomyces cerevisiae. Adv Genet. 1982;21:173–254. doi: 10.1016/s0065-2660(08)60299-0. [DOI] [PubMed] [Google Scholar]
- Lemontt J. F. Mutants of yeast defective in mutation induced by ultraviolet light. Genetics. 1971 May;68(1):21–33. doi: 10.1093/genetics/68.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lemontt J. F. Pathways of ultraviolet mutability in Saccharomyces cerevisiae. I. Some properties of double mutants involving uvs9 and rev. Mutat Res. 1971 Dec;13(4):311–317. doi: 10.1016/0027-5107(71)90041-8. [DOI] [PubMed] [Google Scholar]
- Levinson G., Gutman G. A. High frequencies of short frameshifts in poly-CA/TG tandem repeats borne by bacteriophage M13 in Escherichia coli K-12. Nucleic Acids Res. 1987 Jul 10;15(13):5323–5338. doi: 10.1093/nar/15.13.5323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder P., Lasko P. F., Ashburner M., Leroy P., Nielsen P. J., Nishi K., Schnier J., Slonimski P. P. Birth of the D-E-A-D box. Nature. 1989 Jan 12;337(6203):121–122. doi: 10.1038/337121a0. [DOI] [PubMed] [Google Scholar]
- Luria S. E., Delbrück M. Mutations of Bacteria from Virus Sensitivity to Virus Resistance. Genetics. 1943 Nov;28(6):491–511. doi: 10.1093/genetics/28.6.491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison A., Christensen R. B., Alley J., Beck A. K., Bernstine E. G., Lemontt J. F., Lawrence C. W. REV3, a Saccharomyces cerevisiae gene whose function is required for induced mutagenesis, is predicted to encode a nonessential DNA polymerase. J Bacteriol. 1989 Oct;171(10):5659–5667. doi: 10.1128/jb.171.10.5659-5667.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen P. J., McMaster G. K., Trachsel H. Cloning of eukaryotic protein synthesis initiation factor genes: isolation and characterization of cDNA clones encoding factor eIF-4A. Nucleic Acids Res. 1985 Oct 11;13(19):6867–6880. doi: 10.1093/nar/13.19.6867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patarca R., Freeman G. J., Schwartz J., Singh R. P., Kong Q. T., Murphy E., Anderson Y., Sheng F. Y., Singh P., Johnson K. A. rpt-1, an intracellular protein from helper/inducer T cells that regulates gene expression of interleukin 2 receptor and human immunodeficiency virus type 1. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2733–2737. doi: 10.1073/pnas.85.8.2733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perry L. J., Rixon F. J., Everett R. D., Frame M. C., McGeoch D. J. Characterization of the IE110 gene of herpes simplex virus type 1. J Gen Virol. 1986 Nov;67(Pt 11):2365–2380. doi: 10.1099/0022-1317-67-11-2365. [DOI] [PubMed] [Google Scholar]
- Prakash L. Characterization of postreplication repair in Saccharomyces cerevisiae and effects of rad6, rad18, rev3 and rad52 mutations. Mol Gen Genet. 1981;184(3):471–478. doi: 10.1007/BF00352525. [DOI] [PubMed] [Google Scholar]
- Prakash L., Prakash S. Isolation and characterization of MMS-sensitive mutants of Saccharomyces cerevisiae. Genetics. 1977 May;86(1):33–55. doi: 10.1093/genetics/86.1.33. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray B. K., Lawson T. G., Kramer J. C., Cladaras M. H., Grifo J. A., Abramson R. D., Merrick W. C., Thach R. E. ATP-dependent unwinding of messenger RNA structure by eukaryotic initiation factors. J Biol Chem. 1985 Jun 25;260(12):7651–7658. [PubMed] [Google Scholar]
- Reynolds P., Higgins D. R., Prakash L., Prakash S. The nucleotide sequence of the RAD3 gene of Saccharomyces cerevisiae: a potential adenine nucleotide binding amino acid sequence and a nonessential acidic carboxyl terminal region. Nucleic Acids Res. 1985 Apr 11;13(7):2357–2372. doi: 10.1093/nar/13.7.2357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose M., Botstein D. Construction and use of gene fusions to lacZ (beta-galactosidase) that are expressed in yeast. Methods Enzymol. 1983;101:167–180. doi: 10.1016/0076-6879(83)01012-5. [DOI] [PubMed] [Google Scholar]
- Salvato M. S., Shimomaye E. M. The completed sequence of lymphocytic choriomeningitis virus reveals a unique RNA structure and a gene for a zinc finger protein. Virology. 1989 Nov;173(1):1–10. doi: 10.1016/0042-6822(89)90216-x. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schatz D. G., Oettinger M. A., Baltimore D. The V(D)J recombination activating gene, RAG-1. Cell. 1989 Dec 22;59(6):1035–1048. doi: 10.1016/0092-8674(89)90760-5. [DOI] [PubMed] [Google Scholar]
- Schiestl R. H., Petes T. D. Integration of DNA fragments by illegitimate recombination in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1991 Sep 1;88(17):7585–7589. doi: 10.1073/pnas.88.17.7585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streisinger G., Okada Y., Emrich J., Newton J., Tsugita A., Terzaghi E., Inouye M. Frameshift mutations and the genetic code. This paper is dedicated to Professor Theodosius Dobzhansky on the occasion of his 66th birthday. Cold Spring Harb Symp Quant Biol. 1966;31:77–84. doi: 10.1101/sqb.1966.031.01.014. [DOI] [PubMed] [Google Scholar]
- Sung P., Higgins D., Prakash L., Prakash S. Mutation of lysine-48 to arginine in the yeast RAD3 protein abolishes its ATPase and DNA helicase activities but not the ability to bind ATP. EMBO J. 1988 Oct;7(10):3263–3269. doi: 10.1002/j.1460-2075.1988.tb03193.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung P., Prakash L., Matson S. W., Prakash S. RAD3 protein of Saccharomyces cerevisiae is a DNA helicase. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8951–8955. doi: 10.1073/pnas.84.24.8951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung P., Prakash L., Weber S., Prakash S. The RAD3 gene of Saccharomyces cerevisiae encodes a DNA-dependent ATPase. Proc Natl Acad Sci U S A. 1987 Sep;84(17):6045–6049. doi: 10.1073/pnas.84.17.6045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sung P., Prakash S., Prakash L. The RAD6 protein of Saccharomyces cerevisiae polyubiquitinates histones, and its acidic domain mediates this activity. Genes Dev. 1988 Nov;2(11):1476–1485. doi: 10.1101/gad.2.11.1476. [DOI] [PubMed] [Google Scholar]
- Tagawa M., Sakamoto T., Shigemoto K., Matsubara H., Tamura Y., Ito T., Nakamura I., Okitsu A., Imai K., Taniguchi M. Expression of novel DNA-binding protein with zinc finger structure in various tumor cells. J Biol Chem. 1990 Nov 15;265(32):20021–20026. [PubMed] [Google Scholar]
- Takahashi M., Inaguma Y., Hiai H., Hirose F. Developmentally regulated expression of a human "finger"-containing gene encoded by the 5' half of the ret transforming gene. Mol Cell Biol. 1988 Apr;8(4):1853–1856. doi: 10.1128/mcb.8.4.1853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tamkun J. W., Deuring R., Scott M. P., Kissinger M., Pattatucci A. M., Kaufman T. C., Kennison J. A. brahma: a regulator of Drosophila homeotic genes structurally related to the yeast transcriptional activator SNF2/SWI2. Cell. 1992 Feb 7;68(3):561–572. doi: 10.1016/0092-8674(92)90191-e. [DOI] [PubMed] [Google Scholar]
- Tautz D. Genomic finger printing goes simple. Bioessays. 1990 Jan;12(1):44–46. doi: 10.1002/bies.950120111. [DOI] [PubMed] [Google Scholar]
- Tautz D., Renz M. Simple sequences are ubiquitous repetitive components of eukaryotic genomes. Nucleic Acids Res. 1984 May 25;12(10):4127–4138. doi: 10.1093/nar/12.10.4127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thiem S. M., Miller L. K. A baculovirus gene with a novel transcription pattern encodes a polypeptide with a zinc finger and a leucine zipper. J Virol. 1989 Nov;63(11):4489–4497. doi: 10.1128/jvi.63.11.4489-4497.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker J. E., Saraste M., Runswick M. J., Gay N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1982;1(8):945–951. doi: 10.1002/j.1460-2075.1982.tb01276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walmsley R. M., Szostak J. W., Petes T. D. Is there left-handed DNA at the ends of yeast chromosomes? Nature. 1983 Mar 3;302(5903):84–86. doi: 10.1038/302084a0. [DOI] [PubMed] [Google Scholar]
- Weber J. L., May P. E. Abundant class of human DNA polymorphisms which can be typed using the polymerase chain reaction. Am J Hum Genet. 1989 Mar;44(3):388–396. [PMC free article] [PubMed] [Google Scholar]
- van Straaten F., Müller R., Curran T., Van Beveren C., Verma I. M. Complete nucleotide sequence of a human c-onc gene: deduced amino acid sequence of the human c-fos protein. Proc Natl Acad Sci U S A. 1983 Jun;80(11):3183–3187. doi: 10.1073/pnas.80.11.3183. [DOI] [PMC free article] [PubMed] [Google Scholar]



