Abstract
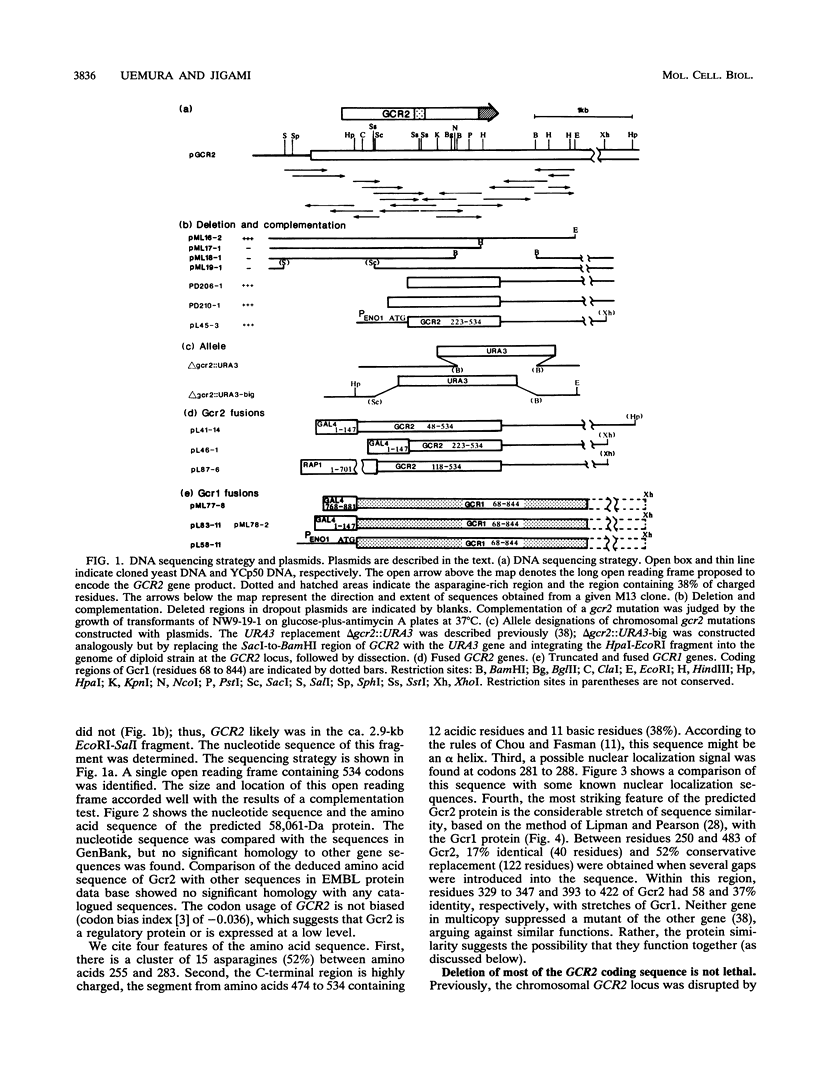
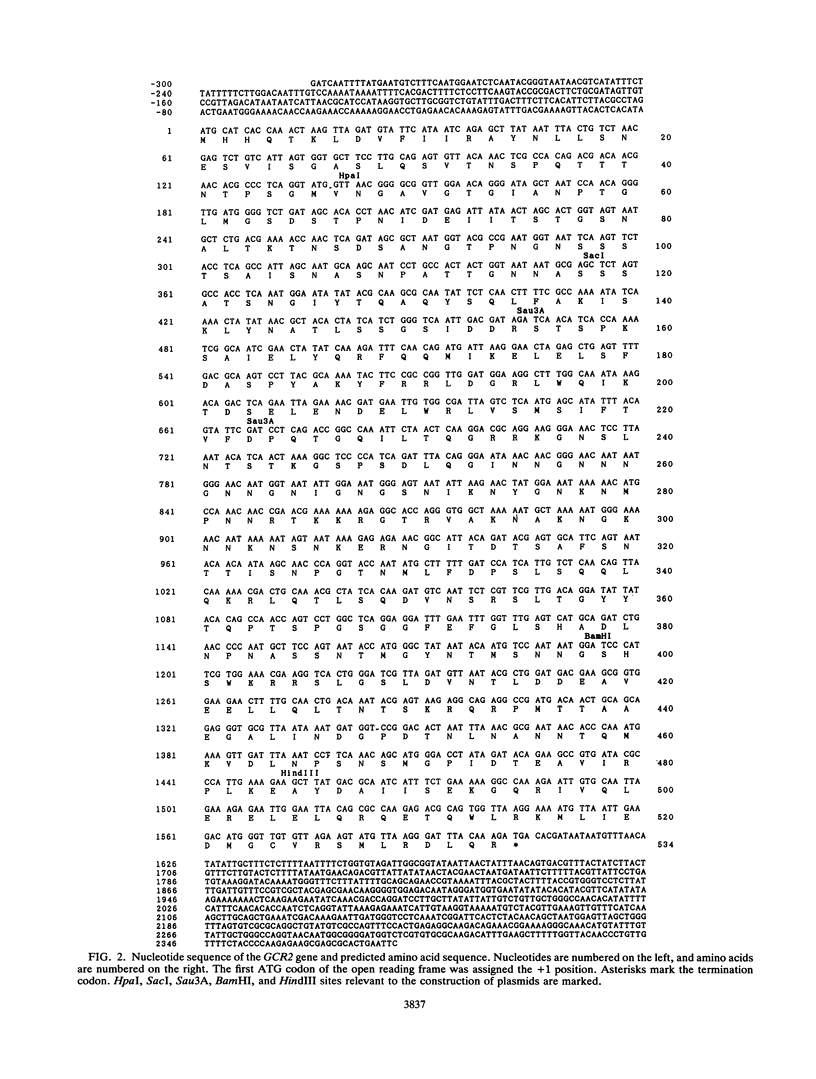
The Saccharomyces cerevisiae GCR2 gene affects expression of most of the glycolytic genes. We report the nucleotide sequence of GCR2, which can potentially encode a 58,061-Da protein. There is a small cluster of asparagines near the center and a C-terminal region that would be highly charged but overall neutral. Fairly homologous regions were found between Gcr2 and Gcr1 proteins. To test potential interactions, the genetic method of S. Fields and O. Song (Nature [London] 340:245-246, 1989), which uses protein fusions of candidate gene products with, respectively, the N-terminal DNA-binding domain of Gal4 and the C-terminal activation domain II, assessing restoration of Gal4 function, was used. In a delta gal4 delta gal80 strain, double transformation by plasmids containing, respectively, a Gal4 (transcription-activating region)/Gcr1 fusion and a Gal4 (DNA-binding domain)/Gcr2 fusion activated lacZ expression from an integrated GAL1/lacZ fusion, indicating reconstitution of functional Gal4 through the interaction of Gcr1 and Gcr2 proteins. The Gal4 (transcription-activating region)/Gcr1 fusion protein alone complemented the defects of both gcr1 and gcr2 strains. Furthermore, a Rap1/Gcr2 fusion protein partially complemented the defects of gcr1 strains. These results suggest that Gcr2 has transcriptional activation activity and that the GCR1 and GCR2 gene products function together.
Full text
PDF






Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baker H. V. GCR1 of Saccharomyces cerevisiae encodes a DNA binding protein whose binding is abolished by mutations in the CTTCC sequence motif. Proc Natl Acad Sci U S A. 1991 Nov 1;88(21):9443–9447. doi: 10.1073/pnas.88.21.9443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker H. V. Glycolytic gene expression in Saccharomyces cerevisiae: nucleotide sequence of GCR1, null mutants, and evidence for expression. Mol Cell Biol. 1986 Nov;6(11):3774–3784. doi: 10.1128/mcb.6.11.3774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennetzen J. L., Hall B. D. Codon selection in yeast. J Biol Chem. 1982 Mar 25;257(6):3026–3031. [PubMed] [Google Scholar]
- Brendel V., Karlin S. Association of charge clusters with functional domains of cellular transcription factors. Proc Natl Acad Sci U S A. 1989 Aug;86(15):5698–5702. doi: 10.1073/pnas.86.15.5698. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brindle P. K., Holland J. P., Willett C. E., Innis M. A., Holland M. J. Multiple factors bind the upstream activation sites of the yeast enolase genes ENO1 and ENO2: ABFI protein, like repressor activator protein RAP1, binds cis-acting sequences which modulate repression or activation of transcription. Mol Cell Biol. 1990 Sep;10(9):4872–4885. doi: 10.1128/mcb.10.9.4872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchman A. R., Kimmerly W. J., Rine J., Kornberg R. D. Two DNA-binding factors recognize specific sequences at silencers, upstream activating sequences, autonomously replicating sequences, and telomeres in Saccharomyces cerevisiae. Mol Cell Biol. 1988 Jan;8(1):210–225. doi: 10.1128/mcb.8.1.210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Buchman A. R., Lue N. F., Kornberg R. D. Connections between transcriptional activators, silencers, and telomeres as revealed by functional analysis of a yeast DNA-binding protein. Mol Cell Biol. 1988 Dec;8(12):5086–5099. doi: 10.1128/mcb.8.12.5086. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers A., Stanway C., Kingsman A. J., Kingsman S. M. The UAS of the yeast PGK gene is composed of multiple functional elements. Nucleic Acids Res. 1988 Sep 12;16(17):8245–8260. doi: 10.1093/nar/16.17.8245. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers A., Stanway C., Tsang J. S., Henry Y., Kingsman A. J., Kingsman S. M. ARS binding factor 1 binds adjacent to RAP1 at the UASs of the yeast glycolytic genes PGK and PYK1. Nucleic Acids Res. 1990 Sep 25;18(18):5393–5399. doi: 10.1093/nar/18.18.5393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers A., Tsang J. S., Stanway C., Kingsman A. J., Kingsman S. M. Transcriptional control of the Saccharomyces cerevisiae PGK gene by RAP1. Mol Cell Biol. 1989 Dec;9(12):5516–5524. doi: 10.1128/mcb.9.12.5516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou P. Y., Fasman G. D. Prediction of the secondary structure of proteins from their amino acid sequence. Adv Enzymol Relat Areas Mol Biol. 1978;47:45–148. doi: 10.1002/9780470122921.ch2. [DOI] [PubMed] [Google Scholar]
- Clifton D., Fraenkel D. G. The gcr (glycolysis regulation) mutation of Saccharomyces cerevisiae. J Biol Chem. 1981 Dec 25;256(24):13074–13078. [PubMed] [Google Scholar]
- Clifton D., Weinstock S. B., Fraenkel D. G. Glycolysis mutants in Saccharomyces cerevisiae. Genetics. 1978 Jan;88(1):1–11. doi: 10.1093/genetics/88.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fields S., Song O. A novel genetic system to detect protein-protein interactions. Nature. 1989 Jul 20;340(6230):245–246. doi: 10.1038/340245a0. [DOI] [PubMed] [Google Scholar]
- Gill G., Ptashne M. Mutants of GAL4 protein altered in an activation function. Cell. 1987 Oct 9;51(1):121–126. doi: 10.1016/0092-8674(87)90016-x. [DOI] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Henry Y. A., Chambers A., Tsang J. S., Kingsman A. J., Kingsman S. M. Characterisation of the DNA binding domain of the yeast RAP1 protein. Nucleic Acids Res. 1990 May 11;18(9):2617–2623. doi: 10.1093/nar/18.9.2617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hess B., Boiteux A., Krüger J. Cooperation of glycolytic enzymes. Adv Enzyme Regul. 1969;7:149–167. doi: 10.1016/0065-2571(69)90016-8. [DOI] [PubMed] [Google Scholar]
- Hill J. E., Myers A. M., Koerner T. J., Tzagoloff A. Yeast/E. coli shuttle vectors with multiple unique restriction sites. Yeast. 1986 Sep;2(3):163–167. doi: 10.1002/yea.320020304. [DOI] [PubMed] [Google Scholar]
- Hoffman C. S., Winston F. A ten-minute DNA preparation from yeast efficiently releases autonomous plasmids for transformation of Escherichia coli. Gene. 1987;57(2-3):267–272. doi: 10.1016/0378-1119(87)90131-4. [DOI] [PubMed] [Google Scholar]
- Holland M. J., Yokoi T., Holland J. P., Myambo K., Innis M. A. The GCR1 gene encodes a positive transcriptional regulator of the enolase and glyceraldehyde-3-phosphate dehydrogenase gene families in Saccharomyces cerevisiae. Mol Cell Biol. 1987 Feb;7(2):813–820. doi: 10.1128/mcb.7.2.813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huet J., Sentenac A. TUF, the yeast DNA-binding factor specific for UASrpg upstream activating sequences: identification of the protein and its DNA-binding domain. Proc Natl Acad Sci U S A. 1987 Jun;84(11):3648–3652. doi: 10.1073/pnas.84.11.3648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalderon D., Richardson W. D., Markham A. F., Smith A. E. Sequence requirements for nuclear location of simian virus 40 large-T antigen. Nature. 1984 Sep 6;311(5981):33–38. doi: 10.1038/311033a0. [DOI] [PubMed] [Google Scholar]
- Kalderon D., Roberts B. L., Richardson W. D., Smith A. E. A short amino acid sequence able to specify nuclear location. Cell. 1984 Dec;39(3 Pt 2):499–509. doi: 10.1016/0092-8674(84)90457-4. [DOI] [PubMed] [Google Scholar]
- Kawasaki G., Fraenkel D. G. Cloning of yeast glycolysis genes by complementation. Biochem Biophys Res Commun. 1982 Oct 15;108(3):1107–1122. doi: 10.1016/0006-291x(82)92114-3. [DOI] [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Lustig A. J., Kurtz S., Shore D. Involvement of the silencer and UAS binding protein RAP1 in regulation of telomere length. Science. 1990 Oct 26;250(4980):549–553. doi: 10.1126/science.2237406. [DOI] [PubMed] [Google Scholar]
- Ma J., Ptashne M. A new class of yeast transcriptional activators. Cell. 1987 Oct 9;51(1):113–119. doi: 10.1016/0092-8674(87)90015-8. [DOI] [PubMed] [Google Scholar]
- Mortimer R. K., Hawthorne D. C. Genetic mapping in yeast. Methods Cell Biol. 1975;11:221–233. doi: 10.1016/s0091-679x(08)60325-8. [DOI] [PubMed] [Google Scholar]
- Nishizawa M., Araki R., Teranishi Y. Identification of an upstream activating sequence and an upstream repressible sequence of the pyruvate kinase gene of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1989 Feb;9(2):442–451. doi: 10.1128/mcb.9.2.442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Olesen J. T., Guarente L. The HAP2 subunit of yeast CCAAT transcriptional activator contains adjacent domains for subunit association and DNA recognition: model for the HAP2/3/4 complex. Genes Dev. 1990 Oct;4(10):1714–1729. doi: 10.1101/gad.4.10.1714. [DOI] [PubMed] [Google Scholar]
- Richardson W. D., Roberts B. L., Smith A. E. Nuclear location signals in polyoma virus large-T. Cell. 1986 Jan 17;44(1):77–85. doi: 10.1016/0092-8674(86)90486-1. [DOI] [PubMed] [Google Scholar]
- Shore D., Nasmyth K. Purification and cloning of a DNA binding protein from yeast that binds to both silencer and activator elements. Cell. 1987 Dec 4;51(5):721–732. doi: 10.1016/0092-8674(87)90095-x. [DOI] [PubMed] [Google Scholar]
- Uemura H., Fraenkel D. G. gcr2, a new mutation affecting glycolytic gene expression in Saccharomyces cerevisiae. Mol Cell Biol. 1990 Dec;10(12):6389–6396. doi: 10.1128/mcb.10.12.6389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uemura H., Jigami Y., Tanaka H., Toshimitsu N., Paterson M., Nakasato S. Nucleotide sequence of the 5' flanking region responsible for the enhancement of the expression of yeast enolase 1 gene. J Biochem. 1985 Sep;98(3):859–862. doi: 10.1093/oxfordjournals.jbchem.a135345. [DOI] [PubMed] [Google Scholar]
- Uemura H., Shiba T., Paterson M., Jigami Y., Tanaka H. Identification of a sequence containing the positive regulatory region of Saccharomyces cerevisiae gene ENO1. Gene. 1986;45(1):67–75. doi: 10.1016/0378-1119(86)90133-2. [DOI] [PubMed] [Google Scholar]
- Uemura H., Wickner R. B. Suppression of chromosomal mutations affecting M1 virus replication in Saccharomyces cerevisiae by a variant of a viral RNA segment (L-A) that encodes coat protein. Mol Cell Biol. 1988 Feb;8(2):938–944. doi: 10.1128/mcb.8.2.938. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wychowski C., Benichou D., Girard M. A domain of SV40 capsid polypeptide VP1 that specifies migration into the cell nucleus. EMBO J. 1986 Oct;5(10):2569–2576. doi: 10.1002/j.1460-2075.1986.tb04536.x. [DOI] [PMC free article] [PubMed] [Google Scholar]



