Abstract
We have conducted an extensive linker substitution analysis of the polyadenylation signal from a pea rbcS gene. From these studies, we can identify at least two, and perhaps three, distinct classes of cis element involved in mRNA 3' end formation in this gene. One of these, termed the far-upstream element, is located between 60 and 120 nt upstream from its associated polyadenylation sites and appears to be largely composed of a series of UG motifs. A second, termed the near-upstream element, is more proximate to poly(A) sites and may be functionally analogous to the mammalian polyadenylation signal AAUAAA, even though the actual sequences involved may not be AAUAAA. The third possible class is the putative cleavage and polyadenylation site itself. We find that the rbcS-E9 far-upstream element can replace the analogous element in another plant polyadenylation signal, that from cauliflower mosaic virus, and that one near-upstream element can function with either of two poly(A) sites. Thus, these different cis elements are largely interchangeable. Our studies indicate that a cellular plant gene possesses upstream elements distinct from AAUAAA that are involved in mRNA 3' end formation and that plant genes probably have modular, multicomponent polyadenylation signals.
Full text
PDF



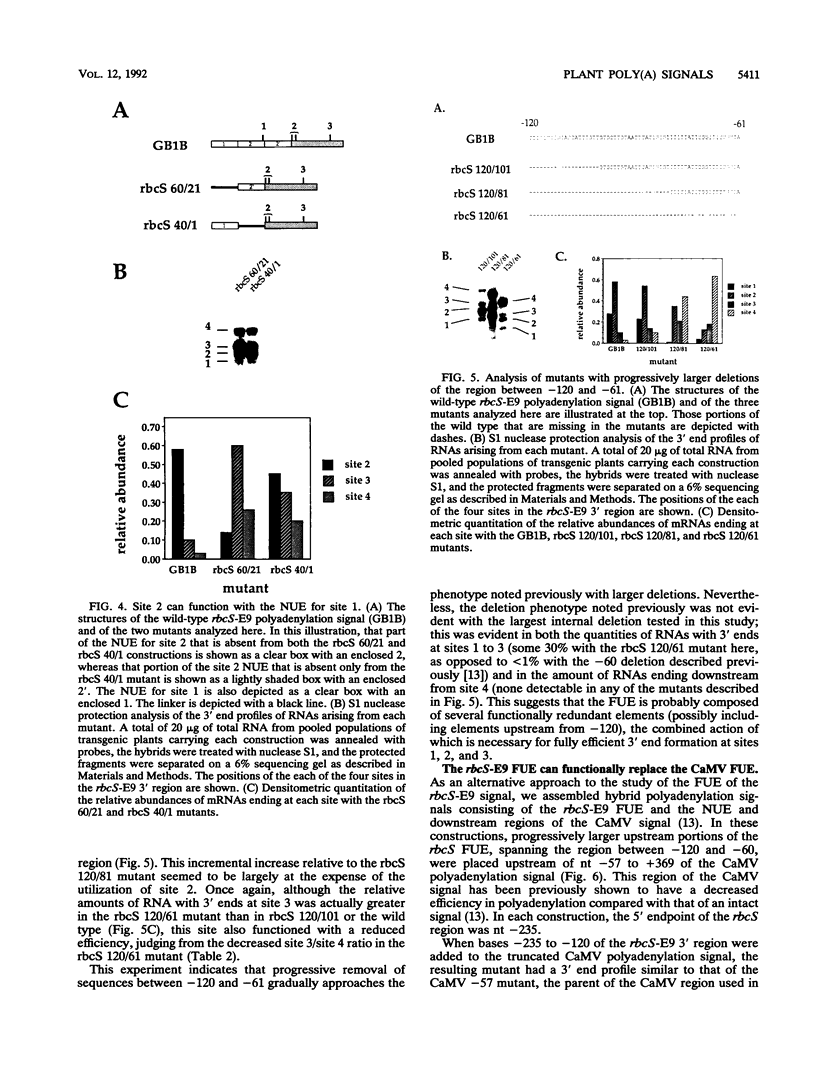



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Abe A., Hiraoka Y., Fukasawa T. Signal sequence for generation of mRNA 3' end in the Saccharomyces cerevisiae GAL7 gene. EMBO J. 1990 Nov;9(11):3691–3697. doi: 10.1002/j.1460-2075.1990.tb07581.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carswell S., Alwine J. C. Efficiency of utilization of the simian virus 40 late polyadenylation site: effects of upstream sequences. Mol Cell Biol. 1989 Oct;9(10):4248–4258. doi: 10.1128/mcb.9.10.4248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeZazzo J. D., Imperiale M. J. Sequences upstream of AAUAAA influence poly(A) site selection in a complex transcription unit. Mol Cell Biol. 1989 Nov;9(11):4951–4961. doi: 10.1128/mcb.9.11.4951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilmartin G. M., Nevins J. R. An ordered pathway of assembly of components required for polyadenylation site recognition and processing. Genes Dev. 1989 Dec;3(12B):2180–2190. doi: 10.1101/gad.3.12b.2180. [DOI] [PubMed] [Google Scholar]
- Guerineau F., Brooks L., Mullineaux P. Effect of deletions in the cauliflower mosaic virus polyadenylation sequence on the choice of the polyadenylation sites in tobacco protoplasts. Mol Gen Genet. 1991 Apr;226(1-2):141–144. doi: 10.1007/BF00273597. [DOI] [PubMed] [Google Scholar]
- Hunt A. G. Identification and characterization of cryptic polyadenylation sites in the 3' region of a pea ribulose-1,5-bisphosphate carboxylase small subunit gene. DNA. 1988 Jun;7(5):329–336. doi: 10.1089/dna.1.1988.7.329. [DOI] [PubMed] [Google Scholar]
- Hunt A. G., MacDonald M. H. Deletion analysis of the polyadenylation signal of a pea ribulose-1,5-bisphosphate carboxylase small-subunit gene. Plant Mol Biol. 1989 Aug;13(2):125–138. doi: 10.1007/BF00016132. [DOI] [PubMed] [Google Scholar]
- Hyman L. E., Seiler S. H., Whoriskey J., Moore C. L. Point mutations upstream of the yeast ADH2 poly(A) site significantly reduce the efficiency of 3'-end formation. Mol Cell Biol. 1991 Apr;11(4):2004–2012. doi: 10.1128/mcb.11.4.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irniger S., Egli C. M., Braus G. H. Different classes of polyadenylation sites in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jun;11(6):3060–3069. doi: 10.1128/mcb.11.6.3060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Joshi C. P. Putative polyadenylation signals in nuclear genes of higher plants: a compilation and analysis. Nucleic Acids Res. 1987 Dec 10;15(23):9627–9640. doi: 10.1093/nar/15.23.9627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacDonald M. H., Mogen B. D., Hunt A. G. Characterization of the polyadenylation signal from the T-DNA-encoded octopine synthase gene. Nucleic Acids Res. 1991 Oct 25;19(20):5575–5581. doi: 10.1093/nar/19.20.5575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McLauchlan J., Gaffney D., Whitton J. L., Clements J. B. The consensus sequence YGTGTTYY located downstream from the AATAAA signal is required for efficient formation of mRNA 3' termini. Nucleic Acids Res. 1985 Feb 25;13(4):1347–1368. doi: 10.1093/nar/13.4.1347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mogen B. D., MacDonald M. H., Graybosch R., Hunt A. G. Upstream sequences other than AAUAAA are required for efficient messenger RNA 3'-end formation in plants. Plant Cell. 1990 Dec;2(12):1261–1272. doi: 10.1105/tpc.2.12.1261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osborne B. I., Guarente L. Mutational analysis of a yeast transcriptional terminator. Proc Natl Acad Sci U S A. 1989 Jun;86(11):4097–4101. doi: 10.1073/pnas.86.11.4097. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Proudfoot N. Poly(A) signals. Cell. 1991 Feb 22;64(4):671–674. doi: 10.1016/0092-8674(91)90495-k. [DOI] [PubMed] [Google Scholar]
- Russnak R. H. Regulation of polyadenylation in hepatitis B viruses: stimulation by the upstream activating signal PS1 is orientation-dependent, distance-independent, and additive. Nucleic Acids Res. 1991 Dec 11;19(23):6449–6456. doi: 10.1093/nar/19.23.6449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russnak R., Ganem D. Sequences 5' to the polyadenylation signal mediate differential poly(A) site use in hepatitis B viruses. Genes Dev. 1990 May;4(5):764–776. doi: 10.1101/gad.4.5.764. [DOI] [PubMed] [Google Scholar]
- Russo P., Li W. Z., Hampsey D. M., Zaret K. S., Sherman F. Distinct cis-acting signals enhance 3' endpoint formation of CYC1 mRNA in the yeast Saccharomyces cerevisiae. EMBO J. 1991 Mar;10(3):563–571. doi: 10.1002/j.1460-2075.1991.tb07983.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanfaçon H., Brodmann P., Hohn T. A dissection of the cauliflower mosaic virus polyadenylation signal. Genes Dev. 1991 Jan;5(1):141–149. doi: 10.1101/gad.5.1.141. [DOI] [PubMed] [Google Scholar]
- Sanfaçon H., Hohn T. Proximity to the promoter inhibits recognition of cauliflower mosaic virus polyadenylation signal. Nature. 1990 Jul 5;346(6279):81–84. doi: 10.1038/346081a0. [DOI] [PubMed] [Google Scholar]
- Sheets M. D., Ogg S. C., Wickens M. P. Point mutations in AAUAAA and the poly (A) addition site: effects on the accuracy and efficiency of cleavage and polyadenylation in vitro. Nucleic Acids Res. 1990 Oct 11;18(19):5799–5805. doi: 10.1093/nar/18.19.5799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Valsamakis A., Zeichner S., Carswell S., Alwine J. C. The human immunodeficiency virus type 1 polyadenylylation signal: a 3' long terminal repeat element upstream of the AAUAAA necessary for efficient polyadenylylation. Proc Natl Acad Sci U S A. 1991 Mar 15;88(6):2108–2112. doi: 10.1073/pnas.88.6.2108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wickens M. How the messenger got its tail: addition of poly(A) in the nucleus. Trends Biochem Sci. 1990 Jul;15(7):277–281. doi: 10.1016/0968-0004(90)90054-f. [DOI] [PubMed] [Google Scholar]