Abstract
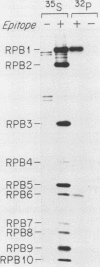
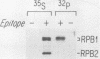
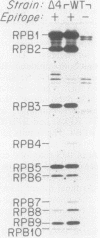
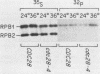
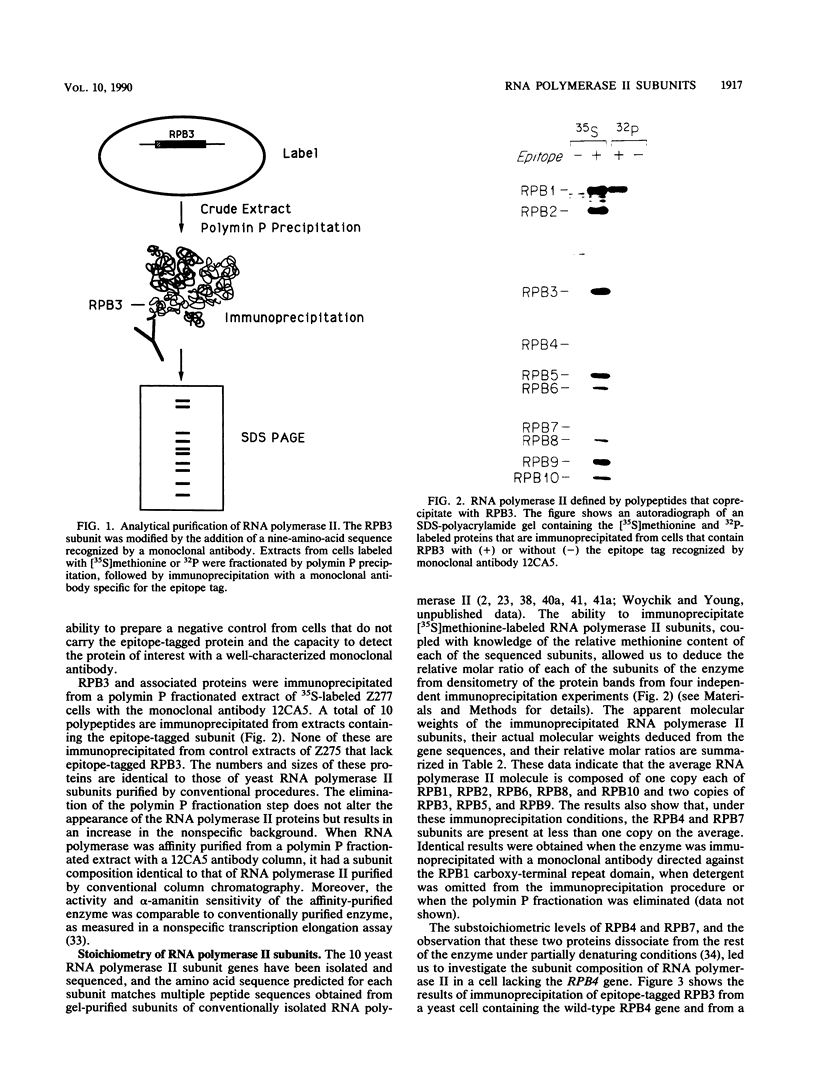
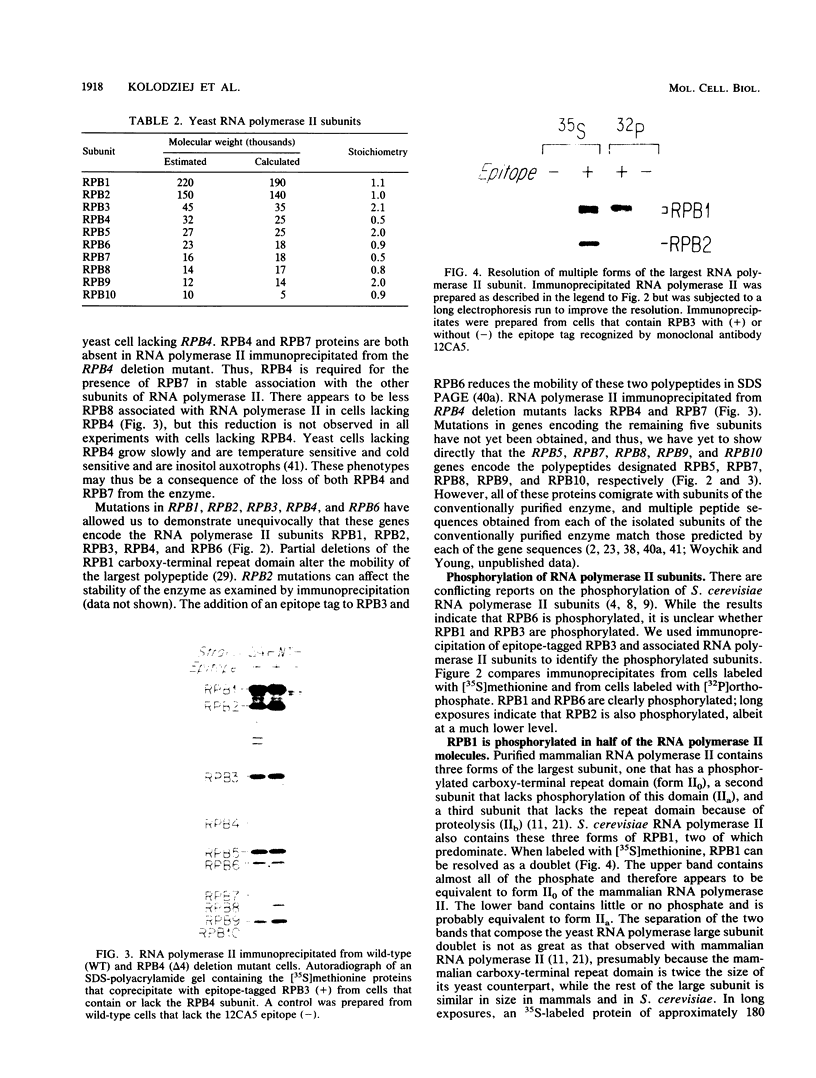
RNA polymerase II subunit composition, stoichiometry, and phosphorylation were investigated in Saccharomyces cerevisiae by attaching an epitope coding sequence to a well-characterized RNA polymerase II subunit gene (RPB3) and by immunoprecipitating the product of this gene with its associated polypeptides. The immunopurified enzyme catalyzed alpha-amanitin-sensitive RNA synthesis in vitro. The 10 polypeptides that immunoprecipitated were identical in size and number to those previously described for RNA polymerase II purified by conventional column chromatography. The relative stoichiometry of the subunits was deduced from knowledge of the sequence of the subunits and from the extent of labeling with [35S]methionine. Immunoprecipitation from 32P-labeled cell extracts revealed that three of the subunits, RPB1, RPB2, and RPB6, are phosphorylated in vivo. Phosphorylated and unphosphorylated forms of RPB1 could be distinguished; approximately half of the RNA polymerase II molecules contained a phosphorylated RPB1 subunit. These results more precisely define the subunit composition and phosphorylation of a eucaryotic RNA polymerase II enzyme.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ahearn J. M., Jr, Bartolomei M. S., West M. L., Cisek L. J., Corden J. L. Cloning and sequence analysis of the mouse genomic locus encoding the largest subunit of RNA polymerase II. J Biol Chem. 1987 Aug 5;262(22):10695–10705. [PubMed] [Google Scholar]
- Allison L. A., Moyle M., Shales M., Ingles C. J. Extensive homology among the largest subunits of eukaryotic and prokaryotic RNA polymerases. Cell. 1985 Sep;42(2):599–610. doi: 10.1016/0092-8674(85)90117-5. [DOI] [PubMed] [Google Scholar]
- Bell G. I., Valenzuela P., Rutter W. J. Phosphorylation of yeast DNA-dependent RNA polymerases in vivo and in vitro. Isolation of enzymes and identification of phosphorylated subunits. J Biol Chem. 1977 May 10;252(9):3082–3091. [PubMed] [Google Scholar]
- Biggs J., Searles L. L., Greenleaf A. L. Structure of the eukaryotic transcription apparatus: features of the gene for the largest subunit of Drosophila RNA polymerase II. Cell. 1985 Sep;42(2):611–621. doi: 10.1016/0092-8674(85)90118-7. [DOI] [PubMed] [Google Scholar]
- Bird D. M., Riddle D. L. Molecular cloning and sequencing of ama-1, the gene encoding the largest subunit of Caenorhabditis elegans RNA polymerase II. Mol Cell Biol. 1989 Oct;9(10):4119–4130. doi: 10.1128/mcb.9.10.4119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bréant B., Buhler J. M., Sentenac A., Fromageot P. On the phosphorylation of yeast RNA polymerases A and B. Eur J Biochem. 1983 Feb 1;130(2):247–251. doi: 10.1111/j.1432-1033.1983.tb07143.x. [DOI] [PubMed] [Google Scholar]
- Buhler J. M., Iborra F., Sentenac A., Fromageot P. Structural studies on yeast RNA polymerases. Existence of common subunits in RNA polymerases A(I) and B(II). J Biol Chem. 1976 Mar 25;251(6):1712–1717. [PubMed] [Google Scholar]
- Buhler J. M., Iborra F., Sentenac A., Fromageot P. The presence of phosphorylated subunits in yeast RNA polymerases A and B. FEBS Lett. 1976 Nov 15;72(1):37–41. doi: 10.1016/0014-5793(76)80893-9. [DOI] [PubMed] [Google Scholar]
- Cadena D. L., Dahmus M. E. Messenger RNA synthesis in mammalian cells is catalyzed by the phosphorylated form of RNA polymerase II. J Biol Chem. 1987 Sep 15;262(26):12468–12474. [PubMed] [Google Scholar]
- Carroll S. B., Stollar B. D. Conservation of a DNA-binding site in the largest subunit of eukaryotic RNA polymerase II. J Mol Biol. 1983 Nov 5;170(3):777–790. doi: 10.1016/s0022-2836(83)80131-4. [DOI] [PubMed] [Google Scholar]
- Cho J. M., Kimball A. P. Probes of eukaryotic DNA-dependent RNA polymerase II-I. Binding of 9-beta-D-arabinofuranosyl-6-mercaptopurine to the elongation subsite. Biochem Pharmacol. 1982 Aug 15;31(16):2575–2581. doi: 10.1016/0006-2952(82)90703-1. [DOI] [PubMed] [Google Scholar]
- Cisek L. J., Corden J. L. Phosphorylation of RNA polymerase by the murine homologue of the cell-cycle control protein cdc2. Nature. 1989 Jun 29;339(6227):679–684. doi: 10.1038/339679a0. [DOI] [PubMed] [Google Scholar]
- Evan G. I., Lewis G. K., Ramsay G., Bishop J. M. Isolation of monoclonal antibodies specific for human c-myc proto-oncogene product. Mol Cell Biol. 1985 Dec;5(12):3610–3616. doi: 10.1128/mcb.5.12.3610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falkenburg D., Dworniczak B., Faust D. M., Bautz E. K. RNA polymerase II of Drosophila. Relation of its 140,000 Mr subunit to the beta subunit of Escherichia coli RNA polymerase. J Mol Biol. 1987 Jun 20;195(4):929–937. doi: 10.1016/0022-2836(87)90496-7. [DOI] [PubMed] [Google Scholar]
- Field J., Nikawa J., Broek D., MacDonald B., Rodgers L., Wilson I. A., Lerner R. A., Wigler M. Purification of a RAS-responsive adenylyl cyclase complex from Saccharomyces cerevisiae by use of an epitope addition method. Mol Cell Biol. 1988 May;8(5):2159–2165. doi: 10.1128/mcb.8.5.2159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grummt I. Nucleotide sequence requirements for specific initiation of transcription by RNA polymerase I. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6908–6911. doi: 10.1073/pnas.79.22.6908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Julius D., Schekman R., Thorner J. Glycosylation and processing of prepro-alpha-factor through the yeast secretory pathway. Cell. 1984 Feb;36(2):309–318. doi: 10.1016/0092-8674(84)90224-1. [DOI] [PubMed] [Google Scholar]
- Kim W. Y., Dahmus M. E. Immunochemical analysis of mammalian RNA polymerase II subspecies. Stability and relative in vivo concentration. J Biol Chem. 1986 Oct 25;261(30):14219–14225. [PubMed] [Google Scholar]
- Klekamp M. S., Weil P. A. Specific transcription of homologous class III genes in yeast-soluble cell-free extracts. J Biol Chem. 1982 Jul 25;257(14):8432–8441. [PubMed] [Google Scholar]
- Kolodziej P., Young R. A. RNA polymerase II subunit RPB3 is an essential component of the mRNA transcription apparatus. Mol Cell Biol. 1989 Dec;9(12):5387–5394. doi: 10.1128/mcb.9.12.5387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Kuo C. L., Campbell J. L. Cloning of Saccharomyces cerevisiae DNA replication genes: isolation of the CDC8 gene and two genes that compensate for the cdc8-1 mutation. Mol Cell Biol. 1983 Oct;3(10):1730–1737. doi: 10.1128/mcb.3.10.1730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lue N. F., Kornberg R. D. Accurate initiation at RNA polymerase II promoters in extracts from Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8839–8843. doi: 10.1073/pnas.84.24.8839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nonet M., Scafe C., Sexton J., Young R. Eucaryotic RNA polymerase conditional mutant that rapidly ceases mRNA synthesis. Mol Cell Biol. 1987 May;7(5):1602–1611. doi: 10.1128/mcb.7.5.1602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nonet M., Sweetser D., Young R. A. Functional redundancy and structural polymorphism in the large subunit of RNA polymerase II. Cell. 1987 Sep 11;50(6):909–915. doi: 10.1016/0092-8674(87)90517-4. [DOI] [PubMed] [Google Scholar]
- Parker C. S., Topol J. A Drosophila RNA polymerase II transcription factor contains a promoter-region-specific DNA-binding activity. Cell. 1984 Feb;36(2):357–369. doi: 10.1016/0092-8674(84)90229-0. [DOI] [PubMed] [Google Scholar]
- Riva M., Schäffner A. R., Sentenac A., Hartmann G. R., Mustaev A. A., Zaychikov E. F., Grachev M. A. Active site labeling of the RNA polymerases A, B, and C from yeast. J Biol Chem. 1987 Oct 25;262(30):14377–14380. [PubMed] [Google Scholar]
- Ruet A., Sentenac A., Fromageot P. A specific assay for yeast RNA polymerases in crude cell extracts. Eur J Biochem. 1978 Oct;90(2):325–330. doi: 10.1111/j.1432-1033.1978.tb12608.x. [DOI] [PubMed] [Google Scholar]
- Ruet A., Sentenac A., Fromageot P., Winsor B., Lacroute F. A mutation of the B220 subunit gene affects the structural and functional properties of yeast RNA polymerase B in vitro. J Biol Chem. 1980 Jul 10;255(13):6450–6455. [PubMed] [Google Scholar]
- Scafe C., Nonet M., Young R. A. RNA polymerase II mutants defective in transcription of a subset of genes. Mol Cell Biol. 1990 Mar;10(3):1010–1016. doi: 10.1128/mcb.10.3.1010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sentenac A. Eukaryotic RNA polymerases. CRC Crit Rev Biochem. 1985;18(1):31–90. doi: 10.3109/10409238509082539. [DOI] [PubMed] [Google Scholar]
- Sweetser D., Nonet M., Young R. A. Prokaryotic and eukaryotic RNA polymerases have homologous core subunits. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1192–1196. doi: 10.1073/pnas.84.5.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weil P. A., Luse D. S., Segall J., Roeder R. G. Selective and accurate initiation of transcription at the Ad2 major late promotor in a soluble system dependent on purified RNA polymerase II and DNA. Cell. 1979 Oct;18(2):469–484. doi: 10.1016/0092-8674(79)90065-5. [DOI] [PubMed] [Google Scholar]
- Wilson I. A., Niman H. L., Houghten R. A., Cherenson A. R., Connolly M. L., Lerner R. A. The structure of an antigenic determinant in a protein. Cell. 1984 Jul;37(3):767–778. doi: 10.1016/0092-8674(84)90412-4. [DOI] [PubMed] [Google Scholar]
- Woychik N. A., Liao S. M., Kolodziej P. A., Young R. A. Subunits shared by eukaryotic nuclear RNA polymerases. Genes Dev. 1990 Mar;4(3):313–323. doi: 10.1101/gad.4.3.313. [DOI] [PubMed] [Google Scholar]
- Woychik N. A., Young R. A. RNA polymerase II subunit RPB4 is essential for high- and low-temperature yeast cell growth. Mol Cell Biol. 1989 Jul;9(7):2854–2859. doi: 10.1128/mcb.9.7.2854. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A., Davis R. W. Yeast RNA polymerase II genes: isolation with antibody probes. Science. 1983 Nov 18;222(4625):778–782. doi: 10.1126/science.6356359. [DOI] [PubMed] [Google Scholar]
- Yura T., Ishihama A. Genetics of bacterial RNA polymerases. Annu Rev Genet. 1979;13:59–97. doi: 10.1146/annurev.ge.13.120179.000423. [DOI] [PubMed] [Google Scholar]