Abstract
The eukaryotic translation initiation factor 4A (eIF-4A) possesses an in vitro helicase activity that allows the unwinding of double-stranded RNA. This activity is dependent on ATP hydrolysis and the presence of another translation initiation factor, eIF-4B. These two initiation factors are thought to unwind mRNA secondary structures in preparation for ribosome binding and initiation of translation. To further characterize the function of eIF-4A in cellular translation and its interaction with other elements of the translation machinery, we have isolated mutations in the TIF1 and TIF2 genes encoding eIF-4A in Saccharomyces cerevisiae. We show that three highly conserved domains of the D-E-A-D protein family, encoding eIF-4A and other RNA helicases, are essential for protein function. Only in rare cases could we make a conservative substitution without affecting cell growth. The mutants show a clear correlation between their growth and in vivo translation rates. One mutation that results in a temperature-sensitive phenotype reveals an immediate decrease in translation activity following a shift to the nonpermissive temperature. These in vivo results confirm previous in vitro data demonstrating an absolute dependence of translation on the TIF1 and TIF2 gene products.
Full text
PDF
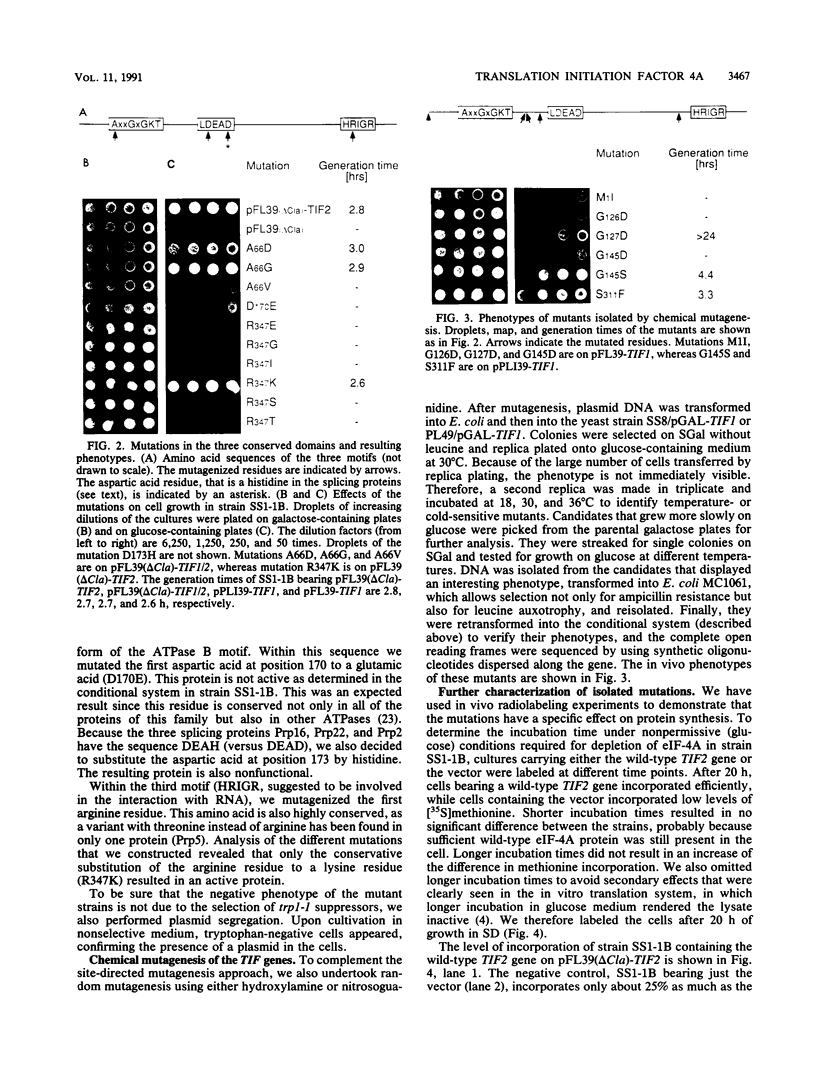







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altmann M., Müller P. P., Pelletier J., Sonenberg N., Trachsel H. A mammalian translation initiation factor can substitute for its yeast homologue in vivo. J Biol Chem. 1989 Jul 25;264(21):12145–12147. [PubMed] [Google Scholar]
- Altmann M., Sonenberg N., Trachsel H. Translation in Saccharomyces cerevisiae: initiation factor 4E-dependent cell-free system. Mol Cell Biol. 1989 Oct;9(10):4467–4472. doi: 10.1128/mcb.9.10.4467. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banroques J., Delahodde A., Jacq C. A mitochondrial RNA maturase gene transferred to the yeast nucleus can control mitochondrial mRNA splicing. Cell. 1986 Sep 12;46(6):837–844. doi: 10.1016/0092-8674(86)90065-6. [DOI] [PubMed] [Google Scholar]
- Blum S., Mueller M., Schmid S. R., Linder P., Trachsel H. Translation in Saccharomyces cerevisiae: initiation factor 4A-dependent cell-free system. Proc Natl Acad Sci U S A. 1989 Aug;86(16):6043–6046. doi: 10.1073/pnas.86.16.6043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burgess S., Couto J. R., Guthrie C. A putative ATP binding protein influences the fidelity of branchpoint recognition in yeast splicing. Cell. 1990 Mar 9;60(5):705–717. doi: 10.1016/0092-8674(90)90086-t. [DOI] [PubMed] [Google Scholar]
- Chang T. H., Arenas J., Abelson J. Identification of five putative yeast RNA helicase genes. Proc Natl Acad Sci U S A. 1990 Feb;87(4):1571–1575. doi: 10.1073/pnas.87.4.1571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen J. H., Lin R. J. The yeast PRP2 protein, a putative RNA-dependent ATPase, shares extensive sequence homology with two other pre-mRNA splicing factors. Nucleic Acids Res. 1990 Nov 11;18(21):6447–6447. doi: 10.1093/nar/18.21.6447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cigan A. M., Feng L., Donahue T. F. tRNAi(met) functions in directing the scanning ribosome to the start site of translation. Science. 1988 Oct 7;242(4875):93–97. doi: 10.1126/science.3051379. [DOI] [PubMed] [Google Scholar]
- Cigan A. M., Pabich E. K., Feng L., Donahue T. F. Yeast translation initiation suppressor sui2 encodes the alpha subunit of eukaryotic initiation factor 2 and shares sequence identity with the human alpha subunit. Proc Natl Acad Sci U S A. 1989 Apr;86(8):2784–2788. doi: 10.1073/pnas.86.8.2784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Company M., Arenas J., Abelson J. Requirement of the RNA helicase-like protein PRP22 for release of messenger RNA from spliceosomes. Nature. 1991 Feb 7;349(6309):487–493. doi: 10.1038/349487a0. [DOI] [PubMed] [Google Scholar]
- Dalbadie-McFarland G., Abelson J. PRP5: a helicase-like protein required for mRNA splicing in yeast. Proc Natl Acad Sci U S A. 1990 Jun;87(11):4236–4240. doi: 10.1073/pnas.87.11.4236. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dombroski A. J., Brennan C. A., Spear P., Platt T. Site-directed alterations in the ATP-binding domain of rho protein affect its activities as a termination factor. J Biol Chem. 1988 Dec 15;263(35):18802–18809. [PubMed] [Google Scholar]
- Donahue T. F., Cigan A. M., Pabich E. K., Valavicius B. C. Mutations at a Zn(II) finger motif in the yeast eIF-2 beta gene alter ribosomal start-site selection during the scanning process. Cell. 1988 Aug 26;54(5):621–632. doi: 10.1016/s0092-8674(88)80006-0. [DOI] [PubMed] [Google Scholar]
- Dorer D. R., Christensen A. C., Johnson D. H. A novel RNA helicase gene tightly linked to the Triplo-lethal locus of Drosophila. Nucleic Acids Res. 1990 Sep 25;18(18):5489–5494. doi: 10.1093/nar/18.18.5489. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Etchison D., Smith K. Variations in cap-binding complexes from uninfected and poliovirus-infected HeLa cells. J Biol Chem. 1990 May 5;265(13):7492–7500. [PubMed] [Google Scholar]
- Ford M. J., Anton I. A., Lane D. P. Nuclear protein with sequence homology to translation initiation factor eIF-4A. Nature. 1988 Apr 21;332(6166):736–738. doi: 10.1038/332736a0. [DOI] [PubMed] [Google Scholar]
- Fry D. C., Kuby S. A., Mildvan A. S. ATP-binding site of adenylate kinase: mechanistic implications of its homology with ras-encoded p21, F1-ATPase, and other nucleotide-binding proteins. Proc Natl Acad Sci U S A. 1986 Feb;83(4):907–911. doi: 10.1073/pnas.83.4.907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goyer C., Altmann M., Trachsel H., Sonenberg N. Identification and characterization of cap-binding proteins from yeast. J Biol Chem. 1989 May 5;264(13):7603–7610. [PubMed] [Google Scholar]
- Grifo J. A., Abramson R. D., Satler C. A., Merrick W. C. RNA-stimulated ATPase activity of eukaryotic initiation factors. J Biol Chem. 1984 Jul 10;259(13):8648–8654. [PubMed] [Google Scholar]
- Guarente L., Yocum R. R., Gifford P. A GAL10-CYC1 hybrid yeast promoter identifies the GAL4 regulatory region as an upstream site. Proc Natl Acad Sci U S A. 1982 Dec;79(23):7410–7414. doi: 10.1073/pnas.79.23.7410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gururajan R., Perry-O'Keefe H., Melton D. A., Weeks D. L. The Xenopus localized messenger RNA An3 may encode an ATP-dependent RNA helicase. Nature. 1991 Feb 21;349(6311):717–719. doi: 10.1038/349717a0. [DOI] [PubMed] [Google Scholar]
- Hirling H., Scheffner M., Restle T., Stahl H. RNA helicase activity associated with the human p68 protein. Nature. 1989 Jun 15;339(6225):562–564. doi: 10.1038/339562a0. [DOI] [PubMed] [Google Scholar]
- Hodgman T. C. A new superfamily of replicative proteins. Nature. 1988 May 5;333(6168):22–23. doi: 10.1038/333022b0. [DOI] [PubMed] [Google Scholar]
- Iggo R. D., Lane D. P. Nuclear protein p68 is an RNA-dependent ATPase. EMBO J. 1989 Jun;8(6):1827–1831. doi: 10.1002/j.1460-2075.1989.tb03577.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iggo R., Picksley S., Southgate J., McPheat J., Lane D. P. Identification of a putative RNA helicase in E.coli. Nucleic Acids Res. 1990 Sep 25;18(18):5413–5417. doi: 10.1093/nar/18.18.5413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jamieson D. J., Rahe B., Pringle J., Beggs J. D. A suppressor of a yeast splicing mutation (prp8-1) encodes a putative ATP-dependent RNA helicase. Nature. 1991 Feb 21;349(6311):715–717. doi: 10.1038/349715a0. [DOI] [PubMed] [Google Scholar]
- Lasko P. F., Ashburner M. The product of the Drosophila gene vasa is very similar to eukaryotic initiation factor-4A. Nature. 1988 Oct 13;335(6191):611–617. doi: 10.1038/335611a0. [DOI] [PubMed] [Google Scholar]
- Lawson T. G., Lee K. A., Maimone M. M., Abramson R. D., Dever T. E., Merrick W. C., Thach R. E. Dissociation of double-stranded polynucleotide helical structures by eukaryotic initiation factors, as revealed by a novel assay. Biochemistry. 1989 May 30;28(11):4729–4734. doi: 10.1021/bi00437a033. [DOI] [PubMed] [Google Scholar]
- Leroy P., Alzari P., Sassoon D., Wolgemuth D., Fellous M. The protein encoded by a murine male germ cell-specific transcript is a putative ATP-dependent RNA helicase. Cell. 1989 May 19;57(4):549–559. doi: 10.1016/0092-8674(89)90125-6. [DOI] [PubMed] [Google Scholar]
- Linder P., Lasko P. F., Ashburner M., Leroy P., Nielsen P. J., Nishi K., Schnier J., Slonimski P. P. Birth of the D-E-A-D box. Nature. 1989 Jan 12;337(6203):121–122. doi: 10.1038/337121a0. [DOI] [PubMed] [Google Scholar]
- Linder P., Prat A. Baker's yeast, the new work horse in protein synthesis studies: analyzing eukaryotic translation initiation. Bioessays. 1990 Nov;12(11):519–526. doi: 10.1002/bies.950121103. [DOI] [PubMed] [Google Scholar]
- Linder P., Slonimski P. P. An essential yeast protein, encoded by duplicated genes TIF1 and TIF2 and homologous to the mammalian translation initiation factor eIF-4A, can suppress a mitochondrial missense mutation. Proc Natl Acad Sci U S A. 1989 Apr;86(7):2286–2290. doi: 10.1073/pnas.86.7.2286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Linder P., Slonimski P. P. Sequence of the genes TIF1 and TIF2 from Saccharomyces cerevisiae coding for a translation initiation factor. Nucleic Acids Res. 1988 Nov 11;16(21):10359–10359. doi: 10.1093/nar/16.21.10359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen P. J., McMaster G. K., Trachsel H. Cloning of eukaryotic protein synthesis initiation factor genes: isolation and characterization of cDNA clones encoding factor eIF-4A. Nucleic Acids Res. 1985 Oct 11;13(19):6867–6880. doi: 10.1093/nar/13.19.6867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nielsen P. J., Trachsel H. The mouse protein synthesis initiation factor 4A gene family includes two related functional genes which are differentially expressed. EMBO J. 1988 Jul;7(7):2097–2105. doi: 10.1002/j.1460-2075.1988.tb03049.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nishi K., Morel-Deville F., Hershey J. W., Leighton T., Schnier J. An eIF-4A-like protein is a suppressor of an Escherichia coli mutant defective in 50S ribosomal subunit assembly. Nature. 1988 Dec 1;336(6198):496–498. doi: 10.1038/336496a0. [DOI] [PubMed] [Google Scholar]
- Planta R. J., Raué H. A. Control of ribosome biogenesis in yeast. Trends Genet. 1988 Mar;4(3):64–68. doi: 10.1016/0168-9525(88)90042-x. [DOI] [PubMed] [Google Scholar]
- Ray B. K., Lawson T. G., Kramer J. C., Cladaras M. H., Grifo J. A., Abramson R. D., Merrick W. C., Thach R. E. ATP-dependent unwinding of messenger RNA structure by eukaryotic initiation factors. J Biol Chem. 1985 Jun 25;260(12):7651–7658. [PubMed] [Google Scholar]
- Rhoads R. E. Cap recognition and the entry of mRNA into the protein synthesis initiation cycle. Trends Biochem Sci. 1988 Feb;13(2):52–56. doi: 10.1016/0968-0004(88)90028-x. [DOI] [PubMed] [Google Scholar]
- Rozen F., Edery I., Meerovitch K., Dever T. E., Merrick W. C., Sonenberg N. Bidirectional RNA helicase activity of eucaryotic translation initiation factors 4A and 4F. Mol Cell Biol. 1990 Mar;10(3):1134–1144. doi: 10.1128/mcb.10.3.1134. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rozen F., Pelletier J., Trachsel H., Sonenberg N. A lysine substitution in the ATP-binding site of eucaryotic initiation factor 4A abrogates nucleotide-binding activity. Mol Cell Biol. 1989 Sep;9(9):4061–4063. doi: 10.1128/mcb.9.9.4061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sachs A. B., Davis R. W. Translation initiation and ribosomal biogenesis: involvement of a putative rRNA helicase and RPL46. Science. 1990 Mar 2;247(4946):1077–1079. doi: 10.1126/science.2408148. [DOI] [PubMed] [Google Scholar]
- Schirmaier F., Philippsen P. Identification of two genes coding for the translation elongation factor EF-1 alpha of S. cerevisiae. EMBO J. 1984 Dec 20;3(13):3311–3315. doi: 10.1002/j.1460-2075.1984.tb02295.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnier J., Schwelberger H. G., Smit-McBride Z., Kang H. A., Hershey J. W. Translation initiation factor 5A and its hypusine modification are essential for cell viability in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jun;11(6):3105–3114. doi: 10.1128/mcb.11.6.3105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sonenberg N. Cap-binding proteins of eukaryotic messenger RNA: functions in initiation and control of translation. Prog Nucleic Acid Res Mol Biol. 1988;35:173–207. doi: 10.1016/s0079-6603(08)60614-5. [DOI] [PubMed] [Google Scholar]
- Stotz A., Linder P. The ADE2 gene from Saccharomyces cerevisiae: sequence and new vectors. Gene. 1990 Oct 30;95(1):91–98. doi: 10.1016/0378-1119(90)90418-q. [DOI] [PubMed] [Google Scholar]
- Sung P., Higgins D., Prakash L., Prakash S. Mutation of lysine-48 to arginine in the yeast RAD3 protein abolishes its ATPase and DNA helicase activities but not the ability to bind ATP. EMBO J. 1988 Oct;7(10):3263–3269. doi: 10.1002/j.1460-2075.1988.tb03193.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Séraphin B., Simon M., Boulet A., Faye G. Mitochondrial splicing requires a protein from a novel helicase family. Nature. 1989 Jan 5;337(6202):84–87. doi: 10.1038/337084a0. [DOI] [PubMed] [Google Scholar]
- Völker T. A., Iida S., Bickle T. A. A single gene coding for resistance to both fusidic acid and chloramphenicol. J Mol Biol. 1982 Jan 25;154(3):417–425. doi: 10.1016/s0022-2836(82)80004-1. [DOI] [PubMed] [Google Scholar]
- Wassarman D. A., Steitz J. A. RNA splicing. Alive with DEAD proteins. Nature. 1991 Feb 7;349(6309):463–464. doi: 10.1038/349463a0. [DOI] [PubMed] [Google Scholar]
- Yaffe M. P., Schatz G. Two nuclear mutations that block mitochondrial protein import in yeast. Proc Natl Acad Sci U S A. 1984 Aug;81(15):4819–4823. doi: 10.1073/pnas.81.15.4819. [DOI] [PMC free article] [PubMed] [Google Scholar]