Abstract
FSP27 [cell death-inducing DFFA-like effector c (CIDEC) in humans] is a protein associated with lipid droplets that downregulates the fatty acid oxidation (FAO) rate when it is overexpressed. However, little is known about its physiological role in liver. Here, we show that fasting regulates liver expression of Fsp27 in a time-dependent manner. Thus, during the initial stages of fasting, a maximal induction of 800-fold was achieved, whereas during the later phase of fasting, Fsp27 expression decreased. The early response to fasting can be explained by a canonical PKA-CREB-CRTC2 signaling pathway because: i) CIDEC expression was induced by forskolin, ii) Fsp27 promoter activity was increased by CREB, and iii) Fsp27 expression was upregulated in the liver of Sirt1 knockout animals. Interestingly, pharmacological (etomoxir) or genetic (Hmgcs2 interference) inhibition of the FAO rate increases the in vivo expression of Fsp27 during fasting. Similarly, CIDEC expression was upregulated in HepG2 cells by either etomoxir or HMGCS2 interference. Our data indicate that there is a kinetic mechanism of autoregulation between short- and long-term fasting, by which free FAs delivered to the liver during early fasting are accumulated/exported by FSP27/CIDEC, whereas over longer periods of fasting, they are degraded in the mitochondria through the carnitine palmitoyl transferase system.
Keywords: liver lipid droplets, CRTC2, SIRT1, fatty acid
During fasting, the liver adapts its metabolism to produce the glucose needed to maintain normoglycemia. Gluconeogenesis is effective when the necessary reducing power (NADH) is made available through the mitochondrial oxidation of FAs (1), which are mobilized from white adipose tissue (WAT) during fasting. Coupling of the two processes is based on the ability of the liver to synthesize ketone bodies, which are soluble products of the incomplete oxidation of FAs that may replace glucose as an energy source, but also increase the fatty acid oxidation (FAO) rate (2).
This intricate process is orchestrated by hormones like insulin, glucagon, or glucocorticoids. Thus, changes in the levels of these hormones affect the gene transcription program of the liver, which can be divided into two temporally distinct phases (3). At the onset of fasting (<12–18 h), hormonal changes produce an increase in both the level and activity of the transcriptional coactivator CRTC2 (CREB-regulated transcription coactivator 2; previously TORC2), which induces an increase in the expression of hepatic gluconeogenic genes (4). With sustained fasting (>12–18 h), SIRT1 becomes activated and deacetylates CRTC2. This event allows the ubiquitination of the protein and targets it for degradation (5). The activation of SIRT1 also results in the deacetylation of the CREB target gene PGC-1α, which is associated with an increase in the activity of this transcriptional coactivator, providing a mechanism to further amplify gluconeogenic gene expression in response to prolonged fasting (6). At the same time, FAs play a major role in maintaining high levels of transcription of the peroxisome proliferator-activated receptor (PPAR)-responsive genes (7, 8).
The increased delivery of FAs to the liver is not an exclusive characteristic of fasting. A high-fat diet produces an increase in liver FAs that leads to steatosis, which is accompanied by the expression of proteins characteristic of lipid droplets. One of these accumulated-liver proteins is the fat-specific protein 27 (FSP27, also known as cell death-inducing DFFA-like effector c (CIDEC in humans), which belongs to a family of proteins that play critical roles in controlling metabolic homeostasis (9, 10). Fsp27 mRNA has been detected in fatty livers, where an excess of lipids accumulates and large lipid droplets are formed (11, 12). More recently, FSP27 was shown to be a direct mediator of PPARγ-dependent hepatic steatosis (12), which suggests that expression of FSP27 may promote lipid-droplet formation in hepatocytes. Interestingly, forced expression of FSP27 in hepatocytes significantly decreased the activity of mitochondrial β-oxidation (12), whereas long-term intermittent fasting induces Fsp27 in WAT (13).
In physiological conditions, FAO is mainly regulated in liver throughout the carnitine palmitoyl transferase (CPT) system (1). The mitochondrial β-oxidation of FAs generates the NADH and ATP needed for gluconeogenesis, and therefore it is an important process in the establishment of considerable liver glucose output during fasting. In agreement, pharmacological treatment with etomoxir, an inhibitor of CPT1 [the key regulatory enzyme of the CPT system (14)], reduces gluconeogenesis and liver glucose output (15).
We have recently shown that downregulation of HMGCS2 [the step-limiting enzyme of ketogenesis (16)] by RNAi attenuates PPARα-dependent stimulation of the FAO rate in the HepG2 cell line (2). We found that expression of a specific shRNA in vivo reduced hepatic HMGCS2 activity by 50%, which correlates with a 20% decrease in liver FAO in fasted animals. In this condition, microarray analysis showed that Fsp27/CIDE (D. Haro and P. Marrero, unpublished observations) was one of the genes that were most upregulated by blocking ketogenesis. Therefore, we studied the expression pattern of Fsp27 during adaptation to fasting, and noted that it was highly induced (∼250-fold and ∼800-fold), mainly in the early period (6 h and 15 h of fasting, respectively). Over longer periods of fasting (24 h), the expression of PPARα target genes remained high, but the expression of Fsp27 decreased 4-fold with respect to its levels after 15 h of fasting. Importantly, we showed that pharmacological inhibition of FAO also upregulated Fsp27/CIDEC in mice liver or HepG2 cells. Additionally, we have reported that this gene is sensitive to both CREB and SIRT1 activity, which could explain its induction during the early stages of fasting.
EXPERIMENTAL PROCEDURES
Plasmids
For the reporter assays, Fsp27 promoter (−2025/+18 relative to the transcription start site) was amplified by PCR from mouse genomic DNA with the oligonucleotides forward (5′-TTAACGCGTCTGCAACTCATTCTGTAGCCC) and reverse (5′-TTACTCGAGGGCAAT ACCGCGTGGCCAG), and cloned in pGL3-basic vector (Promega) using the restriction sites MluI and XhoI, respectively (in bold in the primer sequence). The mutations in the CREB-identified sequences in the mouse Fsp27 promoter were made by site-directed mutagenesis, carried out using the QuickChange™Site-Directed Mutagenesis commercial kit (Stratagene) following the manufacturer's instructions. The mutants were generated by point mutations replacing the original sequences TGACTTCA (CRE1 site, −375/−366) and CGTCA (CRE2 half-site, −1,792/−1,787) by TGAGTATC (mut 1) and ATCGC (mut 2), respectively, in both sense and antisense orientations. Human HMGCS2 promoter (−1,149/+28) was used as a positive control to PPARα transactivation. Empty pGL3-basic was used as a negative control. Mouse PPARα expression vector (pSG5-PPARα) was a kind gift from Dr. S. Green, Macclesfield, UK. Mouse PPARγ2 expression vector (pSVSport-PPARγ) was a kind gift from Dr. B. M. Spiegelman, Harvard Medical School, Boston, MA. The pcDNA3-CREB expression plasmid was subcloned from pSV-CREB in the pcDNA3 empty vector with XbaI/HindIII restriction sites. pSV-CREB and pRSV-KCREB, expressing the wild-type and the dominant-negative form of CREB, respectively, were kindly provided by Dr. R. H. Goodman, Vollum Institute, Portland, OR.
Animal experiments
For the fasting kinetics experiment, 10 week-old C57BL/6J male mice (supplied by Charles River) were used. Mice were either fed ad libitum (ZT12) with a standard chow diet or fasted for 6 h (ZT18), 15 h (ZT3), or 24 h (ZT12) and euthanized at the indicated Zeitgeber time (ZT).
For the etomoxir treatment experiment, 8 week-old C57BL/6J male mice were injected intraperitoneally with a single dose of etomoxir (50 mg/kg body weight) for 16 h. Control mice were injected with the vehicle (water). Mice were fasted for the last 6 h of treatment and then euthanized at ZT18.
For HMGCS2 knockdown experiments, adenoviruses encoding shRNA control or shRNA Hmgcs2 were administered to 9 week-old C57BL/6J mice by tail-vein injection (4 × 109 pfu/animal). Nine days after injection, mice were fasted for 15 h and euthanized at ZT3 (i.e., 3 h after the onset of the 12 h light span). Blood was collected by cardiac puncture and kept on ice until centrifugation (1,500 g, 15 min at 4°C). The serum obtained was stored at 4°C until analysis.
Sirt1 liver-specific knockout (Sirt1-LKO) mice were a kind gift from Dr. L. Guarente (17). Four month-old Sirt1-LKO mice and their age-matched littermate Lox controls (Cre−/−, Sirt1flox/flox) were fasted for 15 h overnight, and euthanized at ZT3.
Glycemia was assessed in mice using an Ascencia Elite XL glucometer and strips (Bayer) to measure glucose in the blood sampled from the heart after isoflurane inhalation (anesthesia) and opening of the cardiac cavity.
Livers were extracted and used for mitochondrial protein extraction or immediately snap-frozen in liquid nitrogen and stored at −80°C until analysis. All mice were housed in cages on a 12:12 h light:dark cycle at controlled temperature (25 ± 1°C). All experimental protocols with mice were performed with the approval of the animal ethics committee of the Universitat de Barcelona, Barcelona, Spain.
Adenovirus generation
For the generation of adenovirus encoding shRNA Hmgcs2, the sequence against mouse Hmgcs2 GGCTTCTGTTCAGTCCAGGAGGACATCAA encoded in the pGFP-V-RS-shRNA vector (Origene, GI579478) was first tested in vitro (data not shown). To construct the recombinant adenovirus, a cassette containing the U6 promoter and shRNA sequence was subcloned from pGFP-V-RS-shRNA to the pDUAL-BASIC-EGFP shuttle vector with EcoRI and HindIII restriction sites. Further adenovirus construction, purification by CsCl gradient, and titration were performed by Vector Biolabs. An adenovirus encoding scrambled shRNA was used as a control.
Ketone body determination
The concentration of total ketone bodies in mice serum was determined using the Autokit Total Ketone Bodies (Wako; Germany), following the manufacturer's instructions, as described previously (2). They are expressed as mM of 3-hydroxybutyrate.
Cell culture and treatments
The human hepatocellular carcinoma cells, HepG2, were cultured at 37°C in a humidified atmosphere containing 5% CO2 in Eagle's MEM supplemented with 100 U/ml penicillin G, 100 µg/ml streptomycin, and heat-inactivated 10% FBS. Etomoxir, Wy14643, GW9662, forskolin, and H89 were obtained from Sigma-Aldrich. Rosiglitazone was from Alexis.
siRNA transfection
HepG2 cells were seeded 24 h before transfection at a density of 4 × 105 cells/well in 6-well plates. Specific siGENOME SMARTpool against HMGCS2 (M-010179-01) and SIRT1 (T2004-01), and siGENOME NonTargeting siRNA #1 (D-001210-01-05), used as a control, were purchased from Thermo Scientific Dharmacon. A concentration of 10 nM was transfected with Dharmafect 4 (Thermo Scientific Dharmacon), according to the manufacturer's instructions. Cells were harvested 72 h posttransfection, and successful knockdown was assessed by real-time PCR.
Mitochondria isolation
Fresh liver (∼1 g) was homogenized in 5 ml of homogenization buffer (250 mM saccharose, 0.1 mM EDTA, and 5 mM Tris-HCl, pH 7.4) using a teflon pestle. The supernatant of the centrifugation (350 g, 10 min at 4°C) of homogenized tissue was again centrifuged at 15,000 g for 15 min at 4°C. The resulting pellet was washed in homogenization buffer, resuspended in 2 ml of resuspension buffer (0.4 mM DTT, 1.5% Triton X-100, 100 mM Tris-HCl, pH 8) and desalted by dialysis using a Dialysis sack, average width 25 mm (1.0 in), against 1:1,000 dialysis solution (20 mM KH2PO4, 12 mM EDTA). Mitochondrial protein was quantified following the Bradford reaction and stored at −80°C.
Enzymatic activity assay
The HMGCS activity determination was carried out as described previously (16). Briefly, HMGCS activity was measured as the incorporation of [1-14C]acetyl-CoA into HMG-CoA at 30°C for 10 min. The reaction was initiated by adding 30 µg of dialyzed mitochondrial protein extract to the reaction mixture (100 mM Tris-HCl, 1 mM EDTA, 20 µM acetoacetyl-CoA, 200 µM and 4,000 cpm/nmol [14C]acetyl-CoA) in a final volume of 200 µl. After 10 min, the enzymatic reaction was stopped by adding 300 µl of 6 N hydrochloric acid, and the mixture was incubated for 2 h at 100°C. To determine the amount of [14C]HMG-CoA formed, nonvolatile radioactivity was recovered from the vials with water, diluted in Ecolite scintillation liquid, and counted in an automatic analyzer. HMGCS-specific activity is expressed as nmol of produced HMG-CoA per min per mg of protein assayed.
The CPT1A activity was measured in liver extracts by subtraction of CPT2 activity from total CPT activity as previously described (18). CPT total activity determination was carried out by the forward exchange method using l-[3H]carnitine. In a total volume of 0.5 ml, the standard enzyme assay mixture contained 0.2 mM of l-[3H]carnitine (∼5,000 dpm/nmol), 80 μM palmitoyl-CoA, 20 mM HEPES (pH 7.0), 1% FA-free albumin, and 60 mM KCl, with (CPT2 activity) or without (total CPT activity) 50 μM malonyl-CoA (Sigma-Aldrich), the physiological inhibitor of CPT1A activity (14). Reactions were initiated by addition of 0.1 mg liver postnuclear extracts (see below). The reaction was linear up to 4 min, and all incubations were done at 30°C for 3 min. Reactions were stopped by addition of 6% perchloric acid and were then centrifuged at 2,300 rpm for 5 min. The resulting pellet was suspended in water, and the product [3H]palmitoylcarnitine was extracted with butanol at low pH. After centrifugation at 3,000 rpm for 3 min, an aliquot of the butanol phase was transferred to a vial for radioactive counting.
FA oxidation determination
FA oxidation was determined as previously described (19, 20). Briefly, hepatic postnuclear extracts were obtained by homogenization of ∼20 mg of frozen tissue in 1 ml of homogenization buffer (50 mM KH2PO4, 150 mM NaCl, 30 mM EDTA, 250 mM saccharose, 1 mM DTT) using a Dounce homogenizer. Following centrifugation at 250 g for 10 min at 4°C, the postnuclear supernatant was recovered and quantified by the Bradford method. FA oxidation was determined as pmol [14C]palmitoyl-CoA oxidized per min per mg protein. The reaction was performed at 37°C for 11 min by adding 15 μg of postnuclear extracts to the reaction mixture (200 mM NAD+, 10 mM FAD+, 1 mM DTT, 75 mg/l BSA, 1 mM l-carnitine, 100 mM CoA, 10 mM and 5,000 dpm/nmol palmitoyl CoA[palmitoyl-1-14C]). The reaction was stopped by adding 7% HClO4, and the mixture was incubated for 1 h at 4°C. Radiolabeled acid-soluble products were determined by counting the supernatant of the centrifugation (12,000 g, 2 min, 4°C) diluted in Ecolite scintillation liquid in an automatic analyzer.
Western blot analysis
Liver mitochondrial and whole-cell extracts were loaded in a 10–12% SDS-PAGE gel and then transferred to Immobilon-P membranes (Millipore; Bedford, MA) and probed with different antibodies. The antibodies used were mHMGCS2 (1:1,000, sc-33828) and VDAC (1:500, sc-32063) from Santa Cruz Biotechnologies, and FSP27 anti-serum [kindly provided by Dr. M. Kasuga, Japan (21)] and Tubulin (Calbiochem, CP06) for mitochondrial and whole-cell extracts, respectively. Detection was carried out using an ECL chemiluminescence detection kit for HRP (Biological Industries).
Real-time PCR analysis
Total RNA was extracted from HepG2 cells or liver by Tri-Reagent (Ambion) and was further treated with DNase I (Ambion). For real-time PCR analysis, cDNA was synthesized from total RNA by murine leukemia virus reverse transcriptase (Invitrogen) with random hexamers (Roche Diagnostics). cDNA was subjected to real-time PCR analysis using TaqMan universal PCR master mix (Invitrogen) and the specific gene expression Taqman probes from Applied Biosystems. For HepG2 cells, the following human gene probes were used: CIDEC Hs00535723_m1, HMGCS2 Hs00985427_m1, and PEPCK Hs00159918_m1. For mice experiments, mouse probes were used: Cact Mm00451571_m1, Cpt1a Mm00550438_m1, Cpt2 Mm00487202_m1, Fsp27 Mm00617672_m1, and Hmgcs2 Mm00550050_m1. Relative mRNA abundance was obtained by normalizing to 18S levels (Applied Biosystems).
Luciferase reporter assay
HepG2 cells were seeded at a density of 1.5 × 105 cells/well in 24-well plates and transfected using Lipofectamine LTX (Invitrogen), following the manufacturer's instructions. Four-hundred nanograms of reporter gene construct and 0.15–0.25 µg of the eukaryotic expression vector (pcDNA3, PPARα, PPARγ, or CREB) were cotransfected. The plasmid pRL-CMV (10 ng/well) was included as an internal transfection control. Cells were harvested after 48 h posttransfection using the passive lysis method (Promega), and luciferase assays were performed using the dual luciferase reporter assay system (Promega), following the manufacturer's recommendations. Firefly and Renilla luciferase activities were determined in a Berthold Sirius Luminometer.
Statistical analysis
All results are expressed as mean ±SEM. Significant differences were assessed using the two-tailed Student's t-test. P < 0.05 was considered statistically significant.
RESULTS
Fsp27 expression is induced in liver during the early steps of fasting
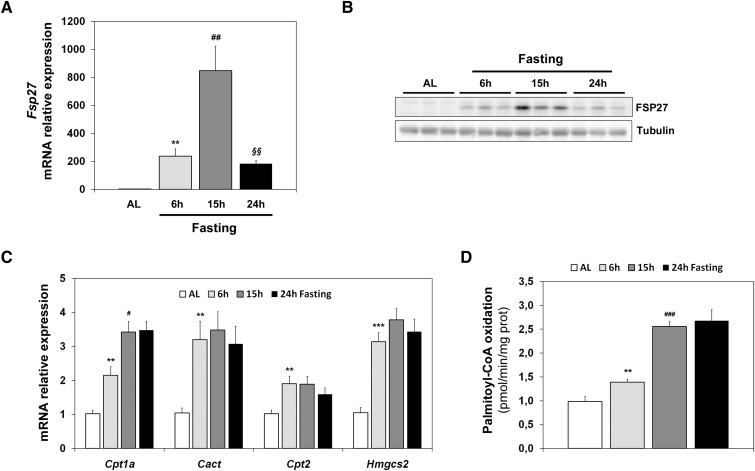
Figure 1A shows that Fsp27 is induced more than 800-fold in early fasting (15 h), but its expression is reduced during longer periods of starvation (24 h). A similar kinetic of FSP27 expression levels is observed for the protein in liver (Fig. 1B). This induction is not observed during the corresponding periods of time in mice fed ad libitum (data not shown). These results suggest that Fsp27 is specifically responsive to the signals that control the early fasting response, and that its expression could be attenuated when expression of genes that control FAO (CPT system [Cpt1a, Cact, Cpt2]) ketogenesis (Hmgcs2) is maintained (Fig. 1C), and FAO is firmly active (Fig. 1D).
Fig. 1.
Hepatic Fsp27 expression is induced by early fasting. Mice were fed ad libitum (AL) or subjected to 6 h, 15 h, or 24 h fasting. Fsp27 (A), Hmgcs2 (C), and other FAO genes (Cpt1a, Cact, and Cpt2) mRNA levels in liver. B: Western blot analysis of liver extracts showing FSP27 protein. Tubulin was used as a control. D: FAO flux measured ex vivo by incubation of liver extract with [14C]palomitoyl-CoA, monitored by acid-soluble metabolites. Results are means ± SEM for each group (n = 5 mice). **P < 0.01, ***P < 0.001 relative to AL; #P < 0.05, ##P < 0.01, ###P < 0.001 relative to 6 h fasting; §§P < 0.01 relative to 15 h fasting.
Downregulation of FAO increases Fsp27/CIDEC expression
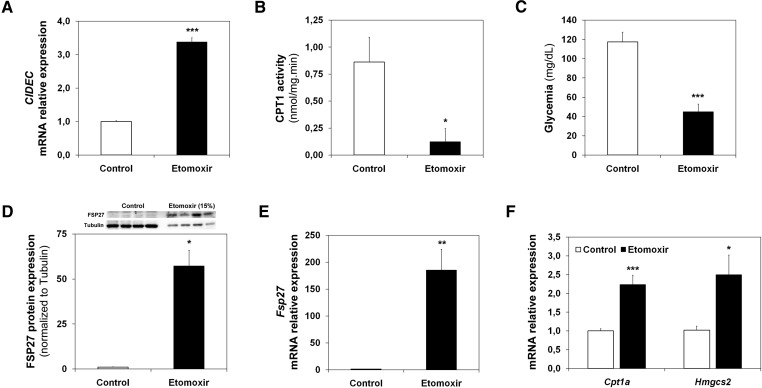
Next, we studied whether the FAO rate affected Fsp27/CIDEC expression. Figure 2A shows that 10 μM etomoxir, a CPT1 inhibitor that blocks FAO (14), induces CIDEC mRNA levels in HepG2 cells. Figure 2B shows that CPT1A activity was inhibited in the liver of short-term (6 h)-starved animals that were intraperitoneally injected with etomoxir. Consistently, the etomoxir-treated animals were hypoglycemic (Fig. 2C). In addition, the etomoxir treatment induced FSP27 expression more than 50-fold (Fig. 2D). This induction correlates with an increase (more than 150-fold) of the Fsp27 mRNA (Fig. 2E). As expected, Fig. 2F shows that the pharmacological treatment induced, to a lesser extent, the hepatic expression of Cpt1a and Hmgcs2, which control FAO and ketogenesis, respectively (22).
Fig. 2.
Fsp27/CIDEC is induced by etomoxir. A: CIDEC mRNA levels in HepG2 cells treated with 10 µM etomoxir for 6 h (mean of three independent experiments). C57BL/6J male animals were treated for 16 h with etomoxir and fasted for the last 6 h of treatment. B: Liver CPT1A activity. C: Blood glucose levels. D: FSP27 and tubulin levels (top) and protein quantification (bottom) in 50 μg (Control) or 7.5 μg (Etomoxir) of liver postnuclear extracts. E: Liver Fsp27 mRNA levels. F: Liver Cpt1a and Hmgcs2 mRNA levels. Results are means of ± SEM for each group (n = 4–5 mice). *P < 0.05, **P < 0.01, ***P < 0.001 relative to Control.
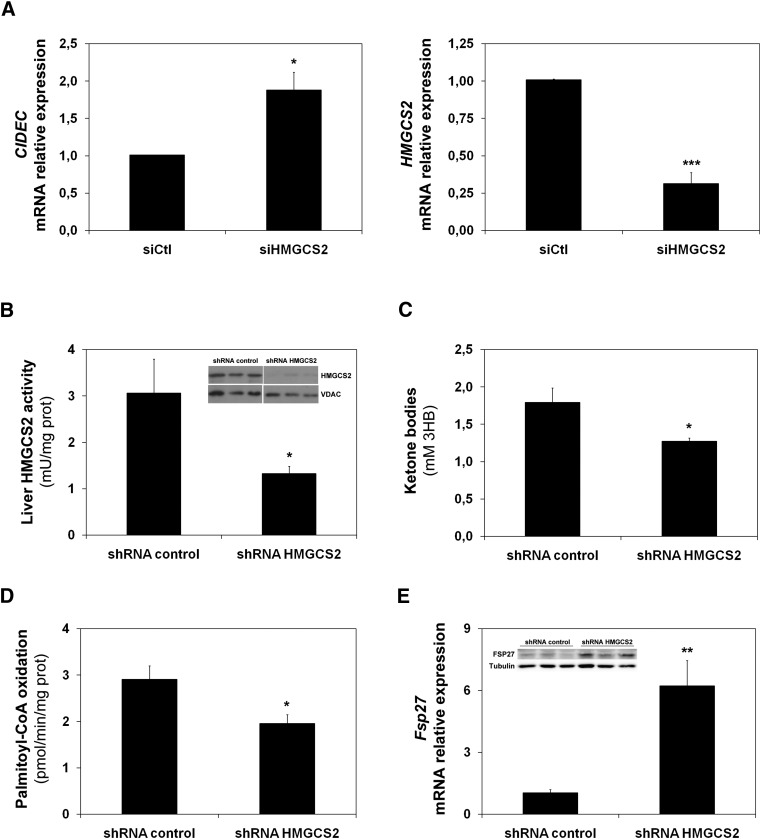
Recently, we have shown that HMGCS2 also regulates the FAO rate in HepG2 cells (2). Therefore, we used the interference of HMGCS2 to downregulate FAO in HepG2 cells and mouse liver. Figure 3A (left) shows that CIDEC mRNA expression was induced by siRNA-mediated downregulation of HMGCS2 (right). The induction of Fsp27 was also observed in a mouse model. Figure 3B shows that a 50% reduction in HMGCS2 activity was achieved by tail-vein injection of an adenovirus that expresses a specific shRNA for HMGCS2 (see insert in Fig. 3B). As expected, the low enzymatic activity correlated with a 25% reduction in plasma ketone bodies (Fig. 3C). Consistently, the interference of HMGCS2 in vivo produced both a 20% reduction of FAO (Fig. 3D) and an increase of Fsp27 mRNA and protein levels (Fig. 3E).
Fig. 3.
Fsp27/CIDEC is induced by HMGCS2 knockdown. A: CIDEC (left panel) and HMGCS2 (right panel) mRNA levels in HepG2 cells treated with a siRNA control or siRNA of HMGCS2 (2). C57BL/6J mice were injected through the tail vein with adenoviruses expressing either a control (shRNA control) or a specific shRNA (Hmgcs2). B: Enzymatic activity in mitochondria from liver of 15 h-fasted animals. The insert shows a Western blot analysis from mitochondrial protein extract for HMGCS2 and the mitochondrial marker VDAC. C: Total ketone bodies, expressed as 3-HB, detected in serum. D: Palmitate oxidation from mice liver extracts measured by acid-soluble metabolites. E: Fsp27 expression in liver. The insert shows a representative Western blot from liver extracts for FSP27 and tubulin. Results are means of ± SEM for each group (n = 5 mice). *P < 0.05, **P < 0.01, ***P < 0.001 relative to siRNA or shRNA control.
Taken together, these results indicate that Fsp27/CIDEC expression is regulated by the FAO rate.
Fsp27 is not a PPARα target gene in HepG2 cells
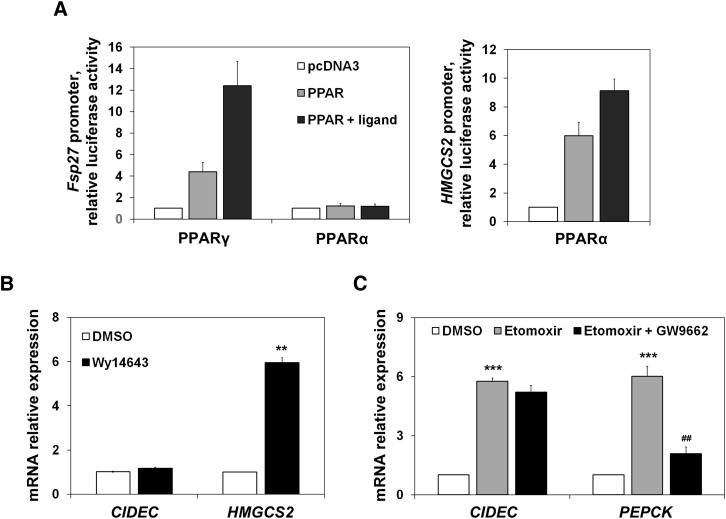
To investigate whether PPARα was responsible for Fsp27 induction during fasting, we used an Fsp27 promoter luciferase assay, and also studied the effect of a PPARα synthetic ligand (Wy14643) on CIDEC mRNA levels in HepG2 cells. Figure 4A shows that, as expected (12), Fsp27 is a PPARγ target gene, although it is not activated by PPARα in a reporter assay in which HMGCS2 promoter [a bona fide PPARα target gene (2)] was used as a control. Additionally, Fig. 4B shows that a PPARα synthetic ligand (Wy14643) was unable to induce CIDEC expression. To further study whether PPARγ was involved in Fsp27/CIDEC expression changes affected by the FAO rate, we used a specific PPARγ antagonist (GW9662). Figure 4C shows that although the antagonist could blunt the etomoxir effect on PEPCK expression, it was unable to attenuate the CIDEC induction mediated by etomoxir.
Fig. 4.
Fsp27/CIDEC is not a PPARα target gene. A: HepG2 cells transfected for 48 h with mouse Fsp27 promoter constructs cloned in pGL3-basic and cotransfected with pcDNA3, PPARγ, or PPARα expression vectors, represented by fold activation to pcDNA3. Twenty-four hours before harvesting, cells were treated with 10 µM of PPARγ and PPARα ligands, rosiglitazone, and Wy14643, respectively (closed bars). Human HMGCS2 promoter cotransfected with PPARα and treated or not with its ligand was used as a positive control. The mean of two independent experiments performed in duplicate are shown. B: CIDEC and HMGCS2 mRNA levels in HepG2 cells treated with the PPARα synthetic ligand Wy14643 (10 µM) for 24 h. C: CIDEC and PEPCK mRNA levels in HepG2 cells treated with etomoxir and a PPARγ antagonist (GW9662) (10 µM each) for 6 h. Results are means of ± SEM (n = 3–4 independent experiments). **P < 0.01, ***P < 0.001 relative to control (DMSO-treated cells); ##P < 0.01 relative to etomoxir-treated cells.
Fsp27 is activated by CREB, and its early induction by fasting is repressed by SIRT1
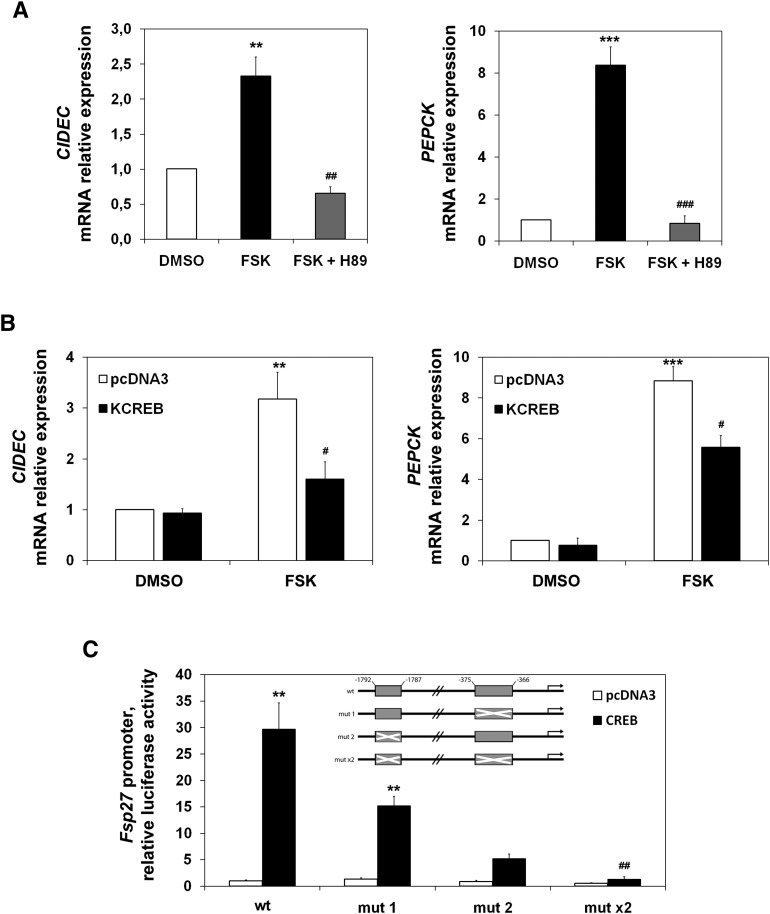
During short-term fasting, the increase in circulating pancreatic glucagon stimulates the gluconeogenic program through the activation of the cAMP pathway, leading to the upregulation of gluconeogenic genes. Figure 5A (left) shows that forskolin, an adenylate cyclase activator, induced CIDEC mRNA in HepG2 cells. This induction was abolished by a PKA inhibitor (H89). Figure 5A (right) shows the expected effect of this cAMP stimulator on PEPCK expression. Figure 5B (left) shows that a plasmid expressing KCREB (a dominant repressor of CREB activity) could also blunt forskolin induction of the CIDEC endogenous gene. To understand the molecular mechanism underlying Fsp27/CIDEC early fasting-mediated expression in liver, we analyzed the sequence of the 5′ flanking region of the mouse Fsp27 gene, and found two putative CREB response elements (CRE) starting at positions −1,787 (CRE2) and −366 (CRE1) upstream of the transcription start site. The upstream site (CRE2) is a conserved half-site motif (CGTCA), whereas the downstream site (CRE1) is an eight-base-pair element (TGACTTCA), partially conserved from the canonical palindrome (TGACGTCA). To test whether the Fsp27 gene transcription was induced by CREB, we made several constructs with the luciferase gene as a reporter (Fig. 5C, insert). We transfected HepG2 cells with these constructs and an expression vector for CREB. Figure 5C shows that CREB overexpression induced the wild-type reporter, and this induction was diminished when either of the single CREs was mutated (mut 1 or mut 2), or totally abolished when both elements were simultaneously mutated (mut ×2). These results identify the CRE1 and CRE2 sequences as CREB-responsive elements in the Fsp27 mouse gene.
Fig. 5.
Fsp27/CIDEC is a CREB target gene. A: CIDEC (left panel) and PEPCK (right panel) mRNA levels in HepG2 cells pretreated for 1 h with 50 µM H89 (PKA inhibitor) and treated with 10 µM forskolin (FSK) in OPTI-MEM (Invitrogen), or vehicle (DMSO) for 6 h. B: CIDEC (left panel) and PEPCK (right panel) mRNA levels in HepG2 cells transfected for 48 h with pcDNA3 or KCREB and treated with 10 µM forskolin (FSK) or vehicle (DMSO) for the last 6 h before harvesting. Results are means of ± SEM (n = 3 independent experiments). **P < 0.01, ***P < 0.001 relative to control, #P < 0.05, relative to FSK-treated cells transfected with empty vector; ##P < 0.01 and ###P < 0.001 relative to FSK-treated cells. C: HepG2 cells transfected for 48 h with pGL3b (control) or mouse Fsp27 promoter constructs cloned in pGL3-basic and cotransfected with either pcDNA3 or pcDNA3-CREB expression vectors, represented by fold activation to pcDNA3 (WT promoter construct). A scheme of the mouse Fsp27 promoter-luciferase reporter construct is shown (shadow box indicates CRE elements, crossed shadow box indicates the construct in which CRE elements were mutated). pGL3-basic activation was subtracted from each condition. Results are means of four independent experiments performed in duplicate (*P < 0.05, **P < 0.01, ***P < 0.001 relative to pcDNA3; #P < 0.05, ##P < 0.01 relative to CREB activation of WT promoter).
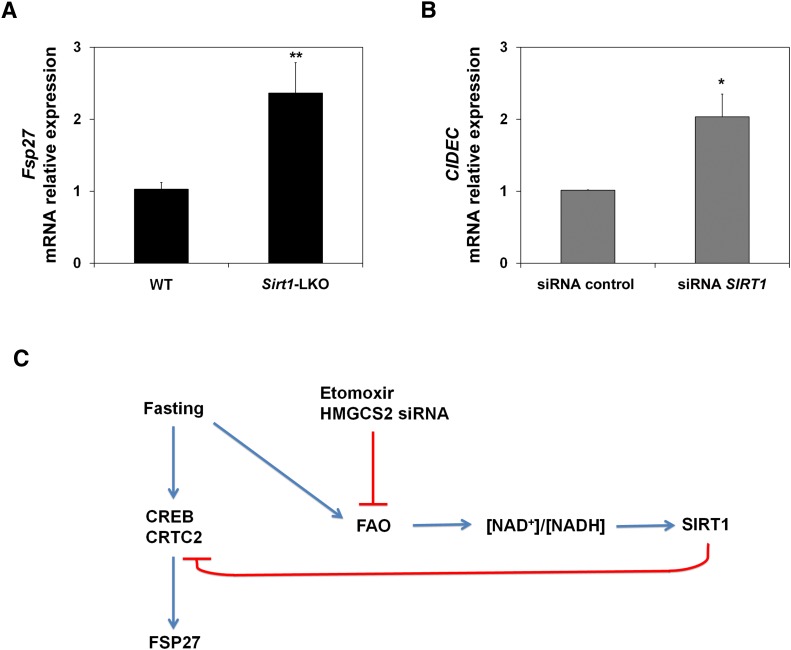
Once gene expression is activated, hepatic SIRT1 activity reaches maximal levels after 18 h of fasting, which promotes a switch between early and late fasting signaling programs (5). To further understand the expression of Fsp27 during fasting, we analyzed its expression in Sirt1 liver-specific knockout mice (Sirt1-LKO) (17). Figure 6A shows that after 15 h fasting, Fsp27 was induced to a higher level in the liver of Sirt1-LKO mice than in the wild-type animals. Figure 6B shows that SIRT1 interference in HepG2 cells (2) also induced the expression of the orthologous CIDEC gene. These data suggest that SIRT1 activity could repress the expression of Fsp27 during late fasting.
Fig. 6.
Fsp27/CIDEC is regulated by SIRT1 activity. A: Fsp27 mRNA levels in liver of wild-type (WT) and Sirt1-LKO mice fasted for 15 h. Data are the mean ± SEM (n = 5–6 mice). **P < 0.01 relative to WT. B: CIDEC mRNA levels in HepG2 cells treated with an siRNA control or siRNA of SIRT1. Results are means of ± SEM (n = 4 independent experiments). *P < 0.05 relative to control. C: Schematic model of the Fsp27/CIDEC regulatory pathway under early fasting.
Taken together, these data indicate that Fsp27 is under the control of the early-fasting induction program. Indeed, during late fasting, SIRT1 expression could be involved in FSP27 downregulation during this period and could also be mediating the effect of FAO rate on its expression (Fig. 6C).
DISCUSSION
In this article, we show that Fsp27 is highly induced during early fasting (6–15 h) (Fig. 1A, B). It is noteworthy that Fsp27 expression was diminished in late fasting (24 h), whereas the expression of the CPT system (Cpt1a, Cpt2, and Cact) and Hmgcs2, which, respectively, regulate FAO and ketogenesis, remained unchanged (Fig. 1C). Therefore, we speculate that Fsp27 is expressed shortly after food removal, whereas the program for FAO adaptation is still in progress (Fig. 1D).
FSP27/CIDEC is a lipid droplet-associated protein that is expressed in WAT or in steatotic liver. Therefore, our finding of hepatic fasting induction of Fsp27 was unexpected. However, a futile cycle of triglyceride/FA has been proposed in fasting (23, 24). In humans, more than 60% of the FAs that reach the liver during fasting are reesterified (25, 26). Indeed, fasting in mice also leads to triglyceride accumulation in the liver. Furthermore, another member of the CIDE family, CIDEB, may be involved in the maturation of VLDL by interacting with apoB-100/-48 (27). Our data suggest that FSP27/CIDEC protein could play a role in the accumulation/export of the newly synthesized triglyceride in the early steps of the fasting adaptation process.
FSP27 expression in liver represses FAO and decreases triglyceride turnover (12, 28). Here we show that in HepG2 cells and mouse liver, pharmacological (etomoxir, Fig. 2) or genetic (RNAi for HMGCS2, Fig. 3) inhibition of FAO induces Fsp27/CIDEC expression. These data suggest that there is a competition for free FAs in the liver between the systems that accumulate and store them (FSP27), and those that oxidize them [the CPT system and ketogenesis (HMGCS2)]. Therefore, we propose a kinetic mechanism of auto-regulation between short- and long-term fasting, by which free FAs delivered to the liver during early fasting are accumulated/exported (FSP27/CIDEC), whereas over longer periods of fasting, FAs are oxidized through the CPT system in the mitochondria.
We investigated whether PPARα, a master regulator of fasting in liver, could be one of the signals that trigger the induction of Fsp27 in fasting. However, the fasting expression pattern of Fsp27 appears not to be related to PPARα signaling, inasmuch as the liver expression pattern of Fsp27 is different from that of bona fide (Cpt1a, Cact, Cpt2, Hmgcs2) PPARα target genes (Fig. 1A, C). In addition, CIDEC endogenous expression is not stimulated by the PPARα ligand in HepG2 cells (Fig. 4B), and the Fsp27 promoter is sensitive to PPARγ, but not to PPARα expression in both HEK293 (12) and HepG2 (Fig. 4A) cell lines. Interestingly, a PPARγ antagonist does not impair the stimulation by etomoxir of CIDEC expression (Fig. 4C). Therefore, our data suggest that inhibition of FAO in liver could alter gene expression through a mechanism different from that of increased levels of free FAs. However, in addition to the PPAR family already studied (12), other transcription factors could be modulated by FAs, and we cannot completely rule out the possibility that lipid accumulation is also responsible for the Fsp27/CIDEC induction.
The two main intracellular signals that regulate the fasting-associated program of gene expression are cAMP and NAD+. SIRT1 has emerged as a protein that can interconnect both intracellular signals, inasmuch as it is a NAD+-consuming deacetylase that was recently shown to be able to deacetylate and attenuate CREB activity (29). Interestingly, CIDEC gene expression was upregulated by forskolin in a hepatoma cell line (Fig. 5A), and this effect was mediated by PKA and CREB (Fig. 5A, B). Consistently, mouse Fsp27 promoter was activated by CREB (Fig. 5B) and upregulated in the liver of Sirt1-LKO mice (Fig. 6A). Recently, it has been suggested that a switch from early gene activation via CRTC2 to late action of FOXO1 is critical for transcriptional regulation in fasting (5). CRTC2 is a coactivator that is responsible for CREB-mediated transcriptional activation of the hepatic gluconeogenic program (30). During prolonged fasting, CRTC2 is deacetylated by the NAD+-dependent enzyme SIRT1, which allows its ubiquitination and degradation in the proteosome to suppress CREB-CRTC2 signaling (5). Therefore, we propose that expression of Fsp27 during early fasting fits the model in which CREB-mediated expression is attenuated in long fasting periods (Fig. 6C).
SIRT1 regulation could also explain why Fsp27/CIDEC expression is induced when FAO is attenuated (Figs. 2 and 3). Recently, SIRT1 has been proposed as a link between protein acetylation and metabolism (31). Thus, during fasting, oxidative metabolism (FAO and ketogenesis) may be expected to pull forward, through the NADH and ATP supply, an anabolic process like gluconeogenesis. In addition to glucose, this will, in turn, generate NAD+ and stimulate SIRT1 (an NAD+-consuming enzyme) activity (2). Therefore, Fsp27/CIDEC, a gene that is repressed by SIRT1, will be upregulated by a blunted FAO (Fig. 6C).
In conclusion, here we show that a lipid droplet-associated protein, Fsp27/CIDEC, was expressed in liver in a physiological situation like the early stages of fasting. We also describe that Fsp27 is a CREB target gene that is upregulated when FAO/ketogenesis is impaired, thus providing new insight into the regulation of fasting metabolism.
Acknowledgments
The authors are truly grateful to Dr. Leonard Guarente and Dr. Hung-Chun Chang (Department of Biology, Massachusetts Institute of Technology) for liver-specific Sirt1 knockout mice and to Dr. Masato Kasuga and Dr. Yoshikazu Tamori (Kobe University Graduate School of Medicine, Japan) for providing FSP27 anti-serum. The authors also thank Carlos Del Rosario Rabadán and Albert Pérez Martí for generating the Fsp27 promoter construct and Western blot assistance, respectively.
Footnotes
Abbreviations:
- CIDEC
- cell death-inducing DFFA-like effector c
- CPT
- carnitine palmitoyl transferase
- FAO
- fatty acid oxidation
- PPAR
- peroxisome proliferator-activated receptor
- WAT
- white adipose tissue
- ZT
- Zeitgeber time
This project was supported by Grants BFU2007-67322/BMC (P.F.M) and SAF2010-15217 (D.H.) from Spain's Ministerio de Educación y Ciencia and by funding from the Catalan Government (Ajut de Suport als Grups de Recerca de Catalunya 2005SGR00857 and 2009SGR163). A.V-B. was supported by a scholarship from the Catalan Government (Ajut al Personal Investigador FPI 2007-2011), and A.L.D.S-C. was supported by the Portuguese Government's Fundação para a Ciência e a Tecnologia.
REFERENCES
- 1.McGarry J. D., Foster D. W. 1980. Regulation of hepatic fatty acid oxidation and ketone body production. Annu. Rev. Biochem. 49: 395–420 [DOI] [PubMed] [Google Scholar]
- 2.Vilà-Brau A., De Sousa-Coelho A. L., Mayordomo C., Haro D., Marrero P. F. 2011. Human HMGCS2 regulates mitochondrial fatty acid oxidation and FGF21 expression in HepG2 cell line. J. Biol. Chem. 286: 20423–20430 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dominy J. E., Lee Y., Gerhart-Hines Z., Puigserver P. 2010. Nutrient-dependent regulation of PGC-1alpha's acetylation state and metabolic function through the enzymatic activities of Sirt1/GCN5. Biochim. Biophys. Acta. 1804: 1676–1683 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Le Lay J., Tuteja G., White P., Dhir R., Ahima R., Kaestner K. H. 2009. CRTC2 (TORC2) contributes to the transcriptional response to fasting in the liver but is not required for the maintenance of glucose homeostasis. Cell Metab. 10: 55–62 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Liu Y., Dentin R., Chen D., Hedrick S., Ravnskjaer K., Schenk S., Milne J., Meyers D. J., Cole P., Yates J., et al. 2008. A fasting inducible switch modulates gluconeogenesis via activator/coactivator exchange. Nature. 456: 269–273 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Altarejos J. Y., Montminy M. 2011. CREB and the CRTC co-activators: sensors for hormonal and metabolic signals. Nat. Rev. Mol. Cell Biol. 12: 141–151 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Leone T. C., Weinheimer C. J., Kelly D. P. 1999. A critical role for the peroxisome proliferator-activated receptor alpha (PPARalpha) in the cellular fasting response: the PPARalpha-null mouse as a model of fatty acid oxidation disorders. Proc. Natl. Acad. Sci. USA. 96: 7473–7478 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Sanderson L. M., Boekschoten M. V., Desvergne B., Müller M., Kersten S. 2010. Transcriptional profiling reveals divergent roles of PPARalpha and PPARbeta/delta in regulation of gene expression in mouse liver. Physiol. Genomics. 41: 42–52 [DOI] [PubMed] [Google Scholar]
- 9.Gong J., Sun Z., Li P. 2009. CIDE proteins and metabolic disorders. Curr. Opin. Lipidol. 20: 121–126 [DOI] [PubMed] [Google Scholar]
- 10.Puri V., Konda S., Ranjit S., Aouadi M., Chawla A., Chouinard M., Chakladar A., Czech M. P. 2007. Fat-specific protein 27, a novel lipid droplet protein that enhances triglyceride storage. J. Biol. Chem. 282: 34213–34218 [DOI] [PubMed] [Google Scholar]
- 11.Yu S., Matsusue K., Kashireddy P., Cao W. Q., Yeldandi V., Yeldandi A. V., Rao M. S., Gonzalez F. J., Reddy J. K. 2003. Adipocyte-specific gene expression and adipogenic steatosis in the mouse liver due to peroxisome proliferator-activated receptor gamma1 (PPARgamma1) overexpression. J. Biol. Chem. 278: 498–505 [DOI] [PubMed] [Google Scholar]
- 12.Matsusue K., Kusakabe T., Noguchi T., Takiguchi S., Suzuki T., Yamano S., Gonzalez F. J. 2008. Hepatic steatosis in leptin-deficient mice is promoted by the PPARgamma target gene Fsp27. Cell Metab. 7: 302–311 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Karbowska J., Kochan Z. 2012. Intermittent fasting up-regulates Fsp27/Cidec gene expression in white adipose tissue. Nutrition. 28: 294–299 [DOI] [PubMed] [Google Scholar]
- 14.Declercq P. E., Falck J. R., Kuwajima M., Tyminski H., Foster D. W., McGarry J. D. 1987. Characterization of the mitochondrial carnitine palmitoyltransferase enzyme system. I. Use of inhibitors. J. Biol. Chem. 262: 9812–9821 [PubMed] [Google Scholar]
- 15.Ratheiser K., Schneeweiss B., Waldhäusl W., Fasching P., Korn A., Nowotny P., Rohac M., Wolf H. P. 1991. Inhibition by etomoxir of carnitine palmitoyltransferase I reduces hepatic glucose production and plasma lipids in non-insulin-dependent diabetes mellitus. Metabolism. 40: 1185–1190 [DOI] [PubMed] [Google Scholar]
- 16.Clinkenbeard K. D., Reed W. D., Mooney R. A., Lane M. D. 1975. Intracellular localization of the 3-hydroxy-3-methylglutaryl coenzme A cycle enzymes in liver. Separate cytoplasmic and mitochondrial 3-hydroxy-3-methylglutaryl coenzyme A generating systems for cholesterogenesis and ketogenesis. J. Biol. Chem. 250: 3108–3116 [PubMed] [Google Scholar]
- 17.Chen D., Bruno J., Easlon E., Lin S. J., Cheng H. L., Alt F. W., Guarente L. 2008. Tissue-specific regulation of SIRT1 by calorie restriction. Genes Dev. 22: 1753–1757 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nicot C., Napal L., Relat J., González S., Llebaria A., Woldegiorgis G., Marrero P. F., Haro D. 2004. C75 activates malonyl-CoA sensitive and insensitive components of the CPT system. Biochem. Biophys. Res. Commun. 325: 660–664 [DOI] [PubMed] [Google Scholar]
- 19.Vilà L., Rebollo A., Ađalsteisson G. S., Alegret M., Merlos M., Roglans N., Laguna J. C. 2011. Reduction of liver fructokinase expression and improved hepatic inflammation and metabolism in liquid fructose-fed rats after atorvastatin treatment. Toxicol. Appl. Pharmacol. 251: 32–40 [DOI] [PubMed] [Google Scholar]
- 20.Lazarow P. B. 1981. Assay of peroxisomal beta-oxidation of fatty acids. Methods Enzymol. 72: 315–319 [DOI] [PubMed] [Google Scholar]
- 21.Nishino N., Tamori Y., Tateya S., Kawaguchi T., Shibakusa T., Mizunoya W., Inoue K., Kitazawa R., Kitazawa S., Matsuki Y., et al. 2008. FSP27 contributes to efficient energy storage in murine white adipocytes by promoting the formation of unilocular lipid droplets. J. Clin. Invest. 118: 2808–2821 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Hegardt F. G., Serra D., Asins G. 1995. Influence of etomoxir on the expression of several genes in liver, testis and heart. Gen. Pharmacol. 26: 897–904 [DOI] [PubMed] [Google Scholar]
- 23.Hanson R. W., Reshef L. 2003. Glyceroneogenesis revisited. Biochimie. 85: 1199–1205 [DOI] [PubMed] [Google Scholar]
- 24.Reshef L., Olswang Y., Cassuto H., Blum B., Croniger C. M., Kalhan S. C., Tilghman S. M., Hanson R. W. 2003. Glyceroneogenesis and the triglyceride/fatty acid cycle. J. Biol. Chem. 278: 30413–30416 [DOI] [PubMed] [Google Scholar]
- 25.Kalhan S. C., Mahajan S., Burkett E., Reshef L., Hanson R. W. 2001. Glyceroneogenesis and the source of glycerol for hepatic triacylglycerol synthesis in humans. J. Biol. Chem. 276: 12928–12931 [DOI] [PubMed] [Google Scholar]
- 26.Jensen M. D., Ekberg K., Landau B. R. 2001. Lipid metabolism during fasting. Am. J. Physiol. Endocrinol. Metab. 281: E789–E793 [DOI] [PubMed] [Google Scholar]
- 27.Ye J., Li J. Z., Liu Y., Li X., Yang T., Ma X., Li Q., Yao Z., Li P. 2009. Cideb, an ER- and lipid droplet-associated protein, mediates VLDL lipidation and maturation by interacting with apolipoprotein B. Cell Metab. 9: 177–190 [DOI] [PubMed] [Google Scholar]
- 28.Matsusue K. 2010. A physiological role for fat specific protein 27/cell death-inducing DFF45-like effector C in adipose and liver. Biol. Pharm. Bull. 33: 346–350 [DOI] [PubMed] [Google Scholar]
- 29.Qiang L., Lin H. V., Kim-Muller J. Y., Welch C. L., Gu W., Accili D. 2011. Proatherogenic abnormalities of lipid metabolism in SirT1 transgenic mice are mediated through Creb deacetylation. Cell Metab. 14: 758–767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Koo S. H., Flechner L., Qi L., Zhang X., Screaton R. A., Jeffries S., Hedrick S., Xu W., Boussouar F., Brindle P., et al. 2005. The CREB coactivator TORC2 is a key regulator of fasting glucose metabolism. Nature. 437: 1109–1111 [DOI] [PubMed] [Google Scholar]
- 31.Guarente L. 2011. The logic linking protein acetylation and metabolism. Cell Metab. 14: 151–153 [DOI] [PubMed] [Google Scholar]