Abstract
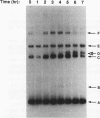
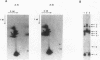
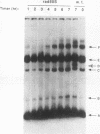
We have used nonessential circular minichromosomes to monitor sister chromatid exchange during yeast meiosis. Genetic analysis shows that a 64-kb circular minichromosome undergoes sister chromatid exchange during 40% of meioses. This frequency is not reduced by the presence of a homologous linear minichromosome. Furthermore, sister chromatid exchange can be stimulated by the presence of a 12-kb ARG4 DNA fragment, which contains initiation sites for meiotic gene conversion. Using physical analysis, we have directly identified a product of sister chromatid exchange: a head-to-tail dimer form of a circular minichromosome. This dimer form is absent in a rad50S mutant strain, which is deficient in processing of the ends of meiosis-specific double-stranded breaks into single-stranded DNA tails. Our studies suggest that meiotic sister chromatid exchange is stimulated by the same mechanism as meiotic homolog exchange.
Full text
PDF
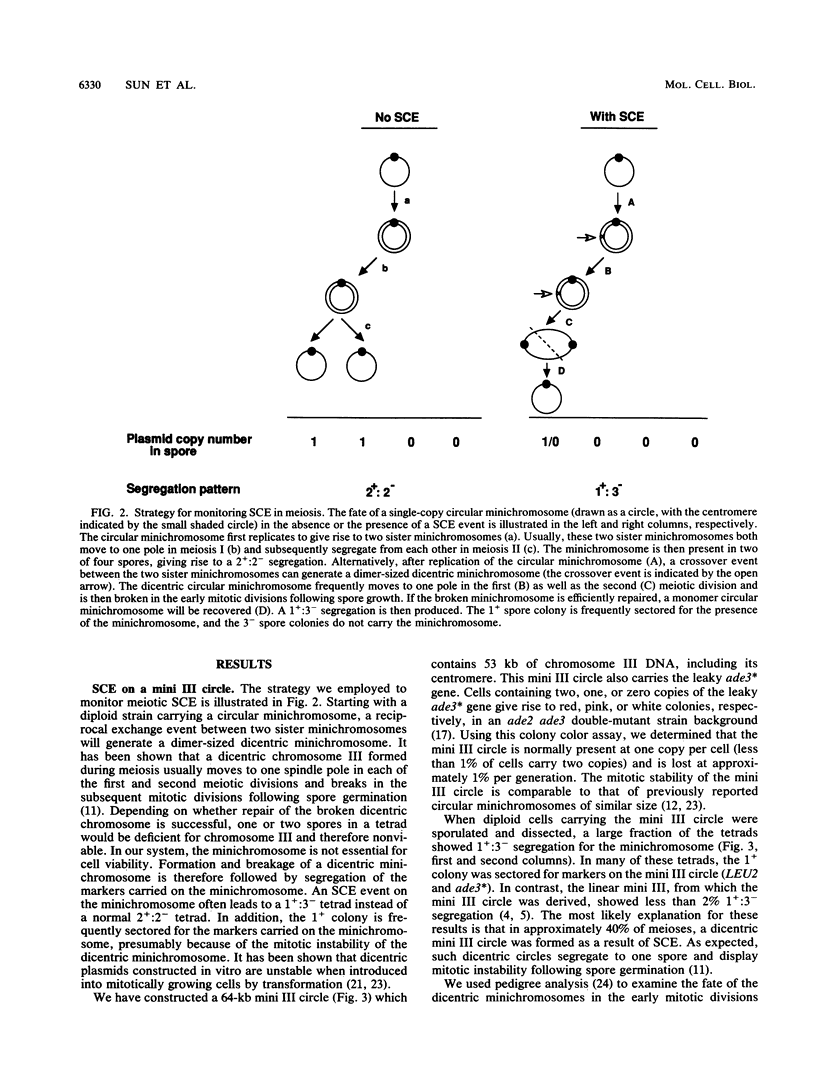
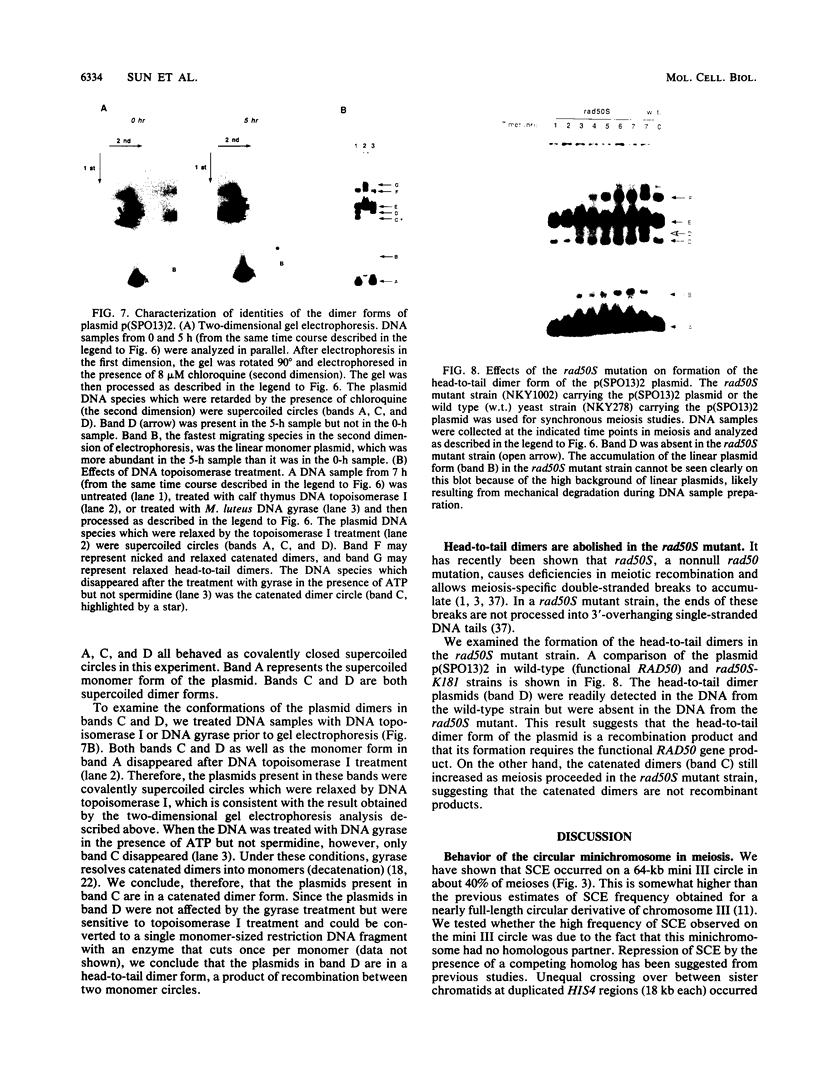

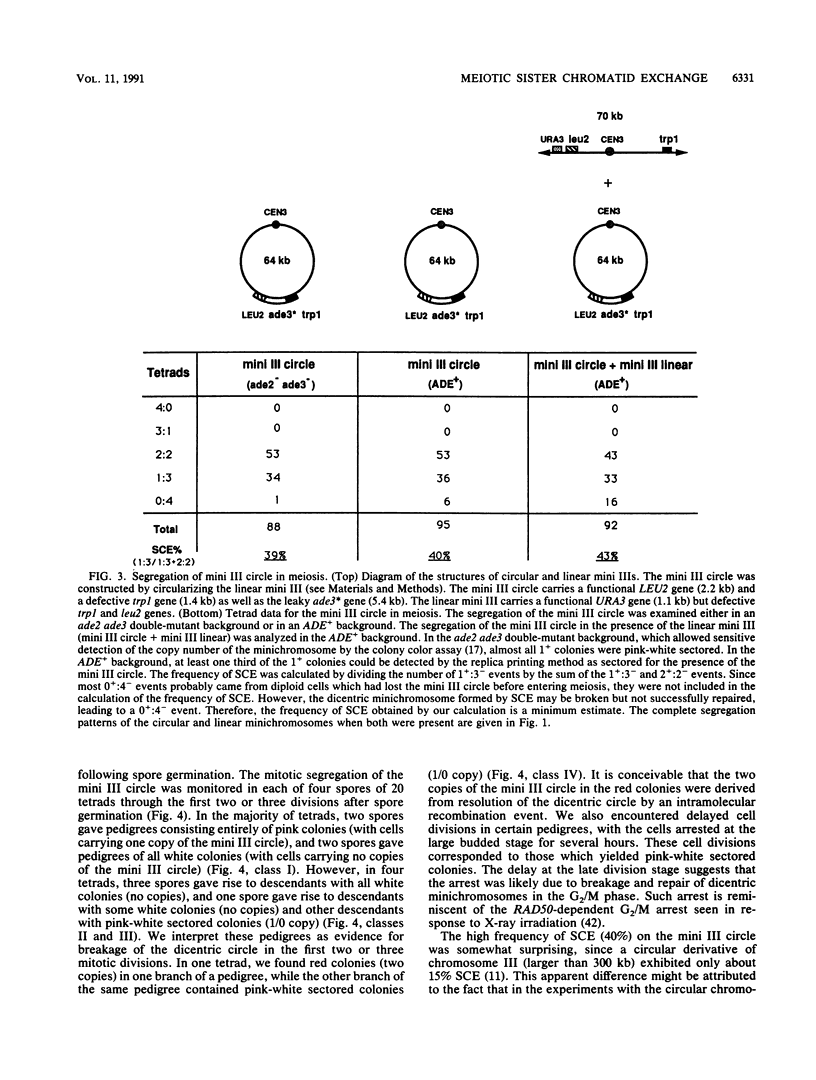





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alani E., Padmore R., Kleckner N. Analysis of wild-type and rad50 mutants of yeast suggests an intimate relationship between meiotic chromosome synapsis and recombination. Cell. 1990 May 4;61(3):419–436. doi: 10.1016/0092-8674(90)90524-i. [DOI] [PubMed] [Google Scholar]
- Cao L., Alani E., Kleckner N. A pathway for generation and processing of double-strand breaks during meiotic recombination in S. cerevisiae. Cell. 1990 Jun 15;61(6):1089–1101. doi: 10.1016/0092-8674(90)90072-m. [DOI] [PubMed] [Google Scholar]
- Dawson D. S., Murray A. W., Szostak J. W. An alternative pathway for meiotic chromosome segregation in yeast. Science. 1986 Nov 7;234(4777):713–717. doi: 10.1126/science.3535068. [DOI] [PubMed] [Google Scholar]
- Game J. C., Sitney K. C., Cook V. E., Mortimer R. K. Use of a ring chromosome and pulsed-field gels to study interhomolog recombination, double-strand DNA breaks and sister-chromatid exchange in yeast. Genetics. 1989 Dec;123(4):695–713. doi: 10.1093/genetics/123.4.695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Game J. C., Zamb T. J., Braun R. J., Resnick M., Roth R. M. The Role of Radiation (rad) Genes in Meiotic Recombination in Yeast. Genetics. 1980 Jan;94(1):51–68. doi: 10.1093/genetics/94.1.51. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giroux C. N., Dresser M. E., Tiano H. F. Genetic control of chromosome synapsis in yeast meiosis. Genome. 1989;31(1):88–94. doi: 10.1139/g89-017. [DOI] [PubMed] [Google Scholar]
- Gottlieb S., Wagstaff J., Esposito R. E. Evidence for two pathways of meiotic intrachromosomal recombination in yeast. Proc Natl Acad Sci U S A. 1989 Sep;86(18):7072–7076. doi: 10.1073/pnas.86.18.7072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guacci V., Kaback D. B. Distributive disjunction of authentic chromosomes in Saccharomyces cerevisiae. Genetics. 1991 Mar;127(3):475–488. doi: 10.1093/genetics/127.3.475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haber J. E., Thorburn P. C., Rogers D. Meiotic and mitotic behavior of dicentric chromosomes in Saccharomyces cerevisiae. Genetics. 1984 Feb;106(2):185–205. doi: 10.1093/genetics/106.2.185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hieter P., Mann C., Snyder M., Davis R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell. 1985 Feb;40(2):381–392. doi: 10.1016/0092-8674(85)90152-7. [DOI] [PubMed] [Google Scholar]
- Jackson J. A., Fink G. R. Meiotic recombination between duplicated genetic elements in Saccharomyces cerevisiae. Genetics. 1985 Feb;109(2):303–332. doi: 10.1093/genetics/109.2.303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaback D. B. Meiotic segregation of circular plasmid-minichromosomes from intact chromosomes in Saccharomyces cerevisiae. Curr Genet. 1989 Jun;15(6):385–392. doi: 10.1007/BF00376792. [DOI] [PubMed] [Google Scholar]
- Kane S. M., Roth R. Carbohydrate metabolism during ascospore development in yeast. J Bacteriol. 1974 Apr;118(1):8–14. doi: 10.1128/jb.118.1.8-14.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klapholz S., Waddell C. S., Esposito R. E. The role of the SPO11 gene in meiotic recombination in yeast. Genetics. 1985 Jun;110(2):187–216. doi: 10.1093/genetics/110.2.187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koshland D., Kent J. C., Hartwell L. H. Genetic analysis of the mitotic transmission of minichromosomes. Cell. 1985 Feb;40(2):393–403. doi: 10.1016/0092-8674(85)90153-9. [DOI] [PubMed] [Google Scholar]
- Kreuzer K. N., Cozzarelli N. R. Formation and resolution of DNA catenanes by DNA gyrase. Cell. 1980 May;20(1):245–254. doi: 10.1016/0092-8674(80)90252-4. [DOI] [PubMed] [Google Scholar]
- Malone R. E., Esposito R. E. Recombinationless meiosis in Saccharomyces cerevisiae. Mol Cell Biol. 1981 Oct;1(10):891–901. doi: 10.1128/mcb.1.10.891. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mann C., Davis R. W. Instability of dicentric plasmids in yeast. Proc Natl Acad Sci U S A. 1983 Jan;80(1):228–232. doi: 10.1073/pnas.80.1.228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizuuchi K., Fisher L. M., O'Dea M. H., Gellert M. DNA gyrase action involves the introduction of transient double-strand breaks into DNA. Proc Natl Acad Sci U S A. 1980 Apr;77(4):1847–1851. doi: 10.1073/pnas.77.4.1847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray A. W., Schultes N. P., Szostak J. W. Chromosome length controls mitotic chromosome segregation in yeast. Cell. 1986 May 23;45(4):529–536. doi: 10.1016/0092-8674(86)90284-9. [DOI] [PubMed] [Google Scholar]
- Murray A. W., Szostak J. W. Chromosome segregation in mitosis and meiosis. Annu Rev Cell Biol. 1985;1:289–315. doi: 10.1146/annurev.cb.01.110185.001445. [DOI] [PubMed] [Google Scholar]
- Murray A. W., Szostak J. W. Pedigree analysis of plasmid segregation in yeast. Cell. 1983 Oct;34(3):961–970. doi: 10.1016/0092-8674(83)90553-6. [DOI] [PubMed] [Google Scholar]
- Nicolas A., Treco D., Schultes N. P., Szostak J. W. An initiation site for meiotic gene conversion in the yeast Saccharomyces cerevisiae. Nature. 1989 Mar 2;338(6210):35–39. doi: 10.1038/338035a0. [DOI] [PubMed] [Google Scholar]
- Orr-Weaver T. L., Szostak J. W., Rothstein R. J. Genetic applications of yeast transformation with linear and gapped plasmids. Methods Enzymol. 1983;101:228–245. doi: 10.1016/0076-6879(83)01017-4. [DOI] [PubMed] [Google Scholar]
- Petes T. D. Unequal meiotic recombination within tandem arrays of yeast ribosomal DNA genes. Cell. 1980 Mar;19(3):765–774. doi: 10.1016/s0092-8674(80)80052-3. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Schultes N. P., Szostak J. W. A poly(dA.dT) tract is a component of the recombination initiation site at the ARG4 locus in Saccharomyces cerevisiae. Mol Cell Biol. 1991 Jan;11(1):322–328. doi: 10.1128/mcb.11.1.322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schultes N. P., Szostak J. W. Decreasing gradients of gene conversion on both sides of the initiation site for meiotic recombination at the ARG4 locus in yeast. Genetics. 1990 Dec;126(4):813–822. doi: 10.1093/genetics/126.4.813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun H., Treco D., Schultes N. P., Szostak J. W. Double-strand breaks at an initiation site for meiotic gene conversion. Nature. 1989 Mar 2;338(6210):87–90. doi: 10.1038/338087a0. [DOI] [PubMed] [Google Scholar]
- Sun H., Treco D., Szostak J. W. Extensive 3'-overhanging, single-stranded DNA associated with the meiosis-specific double-strand breaks at the ARG4 recombination initiation site. Cell. 1991 Mar 22;64(6):1155–1161. doi: 10.1016/0092-8674(91)90270-9. [DOI] [PubMed] [Google Scholar]
- Sundin O., Varshavsky A. Terminal stages of SV40 DNA replication proceed via multiply intertwined catenated dimers. Cell. 1980 Aug;21(1):103–114. doi: 10.1016/0092-8674(80)90118-x. [DOI] [PubMed] [Google Scholar]
- Wagstaff J. E., Klapholz S., Waddell C. S., Jensen L., Esposito R. E. Meiotic exchange within and between chromosomes requires a common Rec function in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Dec;5(12):3532–3544. doi: 10.1128/mcb.5.12.3532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H. T., Frackman S., Kowalisyn J., Esposito R. E., Elder R. Developmental regulation of SPO13, a gene required for separation of homologous chromosomes at meiosis I. Mol Cell Biol. 1987 Apr;7(4):1425–1435. doi: 10.1128/mcb.7.4.1425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J. C., Peck L. J., Becherer K. DNA supercoiling and its effects on DNA structure and function. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 1):85–91. doi: 10.1101/sqb.1983.047.01.011. [DOI] [PubMed] [Google Scholar]
- Weinert T. A., Hartwell L. H. The RAD9 gene controls the cell cycle response to DNA damage in Saccharomyces cerevisiae. Science. 1988 Jul 15;241(4863):317–322. doi: 10.1126/science.3291120. [DOI] [PubMed] [Google Scholar]