Abstract
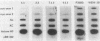
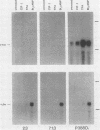
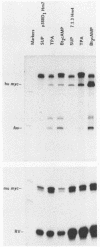
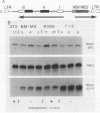
As in tumors with c-myc chromosomal translocations, c-myc retrovirus-induced monocyte tumors constitutively express an activated form of c-myc (the proviral gene), whereas the normal endogenous c-myc genes are transcriptionally silent. Treatment of these retrovirus-induced tumor cells with a number of bioactive chemicals and growth factors that are known to induce c-myc expression in cells of the monocyte lineage failed to induce the endogenous c-myc gene. In contrast, the same treatments induced the c-fos gene in both tumors and a control macrophage line. To investigate c-myc suppression further, a normal copy of the human c-myc gene was introduced into tumor and control cell lines by using a retrovirus with self-inactivating long terminal repeats. This transduced normal gene was expressed at equivalent levels in all cells, regardless of the state of endogenous c-myc gene expression, and was strongly induced by agents that induce the normal gene in the control cells. These results indicate that the signal transduction pathways that normally activate the c-myc gene are functional in myc-induced tumor cells and suggest that endogenous c-myc is actively suppressed. An examination of the c-myc locus itself showed that the lack of transcriptional activity correlated with the absence of several prominent DNase I-hypersensitive sites in the 5'-flanking region of the gene but without loss of general DNase sensitivity. Furthermore, analysis of 22 methylation-sensitive restriction enzyme sites in the 5'-flanking region, first exon, and first intron indicated that the silent c-myc genes remained in the same unmethylated state as did actively expressed genes. Thus, c-myc suppression does not appear to result from the most frequently described mechanisms of gene inactivation.
Full text
PDF


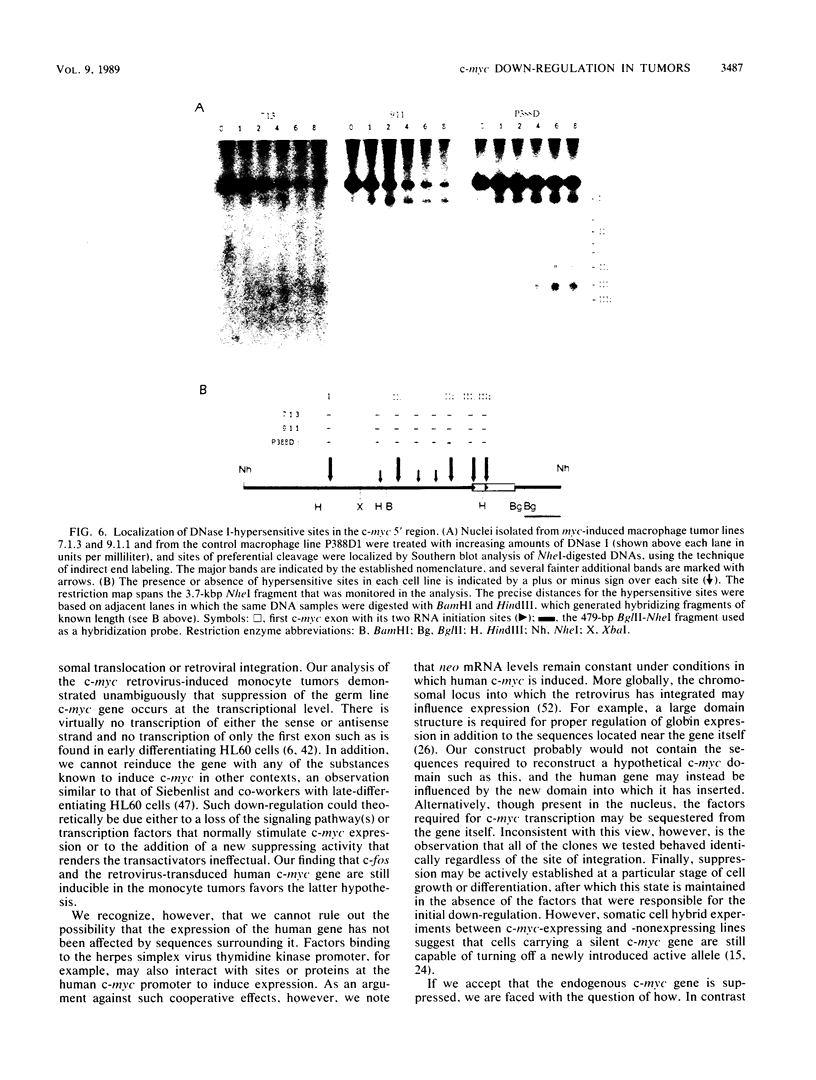



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adams J. M., Gerondakis S., Webb E., Corcoran L. M., Cory S. Cellular myc oncogene is altered by chromosome translocation to an immunoglobulin locus in murine plasmacytomas and is rearranged similarly in human Burkitt lymphomas. Proc Natl Acad Sci U S A. 1983 Apr;80(7):1982–1986. doi: 10.1073/pnas.80.7.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Armelin H. A., Armelin M. C., Kelly K., Stewart T., Leder P., Cochran B. H., Stiles C. D. Functional role for c-myc in mitogenic response to platelet-derived growth factor. Nature. 1984 Aug 23;310(5979):655–660. doi: 10.1038/310655a0. [DOI] [PubMed] [Google Scholar]
- Baumbach W. R., Keath E. J., Cole M. D. A mouse c-myc retrovirus transforms established fibroblast lines in vitro and induces monocyte-macrophage tumors in vivo. J Virol. 1986 Aug;59(2):276–283. doi: 10.1128/jvi.59.2.276-283.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baumbach W. R., Stanley E. R., Cole M. D. Induction of clonal monocyte-macrophage tumors in vivo by a mouse c-myc retrovirus: rearrangement of the CSF-1 gene as a secondary transforming event. Mol Cell Biol. 1987 Feb;7(2):664–671. doi: 10.1128/mcb.7.2.664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bentley D. L., Groudine M. A block to elongation is largely responsible for decreased transcription of c-myc in differentiated HL60 cells. Nature. 1986 Jun 12;321(6071):702–706. doi: 10.1038/321702a0. [DOI] [PubMed] [Google Scholar]
- Bentley D. L., Groudine M. Novel promoter upstream of the human c-myc gene and regulation of c-myc expression in B-cell lymphomas. Mol Cell Biol. 1986 Oct;6(10):3481–3489. doi: 10.1128/mcb.6.10.3481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bravo R., Neuberg M., Burckhardt J., Almendral J., Wallich R., Müller R. Involvement of common and cell type-specific pathways in c-fos gene control: stable induction of cAMP in macrophages. Cell. 1987 Jan 30;48(2):251–260. doi: 10.1016/0092-8674(87)90428-4. [DOI] [PubMed] [Google Scholar]
- Cedar H. DNA methylation and gene activity. Cell. 1988 Apr 8;53(1):3–4. doi: 10.1016/0092-8674(88)90479-5. [DOI] [PubMed] [Google Scholar]
- Chung J., Sinn E., Reed R. R., Leder P. Trans-acting elements modulate expression of the human c-myc gene in Burkitt lymphoma cells. Proc Natl Acad Sci U S A. 1986 Oct;83(20):7918–7922. doi: 10.1073/pnas.83.20.7918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Church G. M., Gilbert W. Genomic sequencing. Proc Natl Acad Sci U S A. 1984 Apr;81(7):1991–1995. doi: 10.1073/pnas.81.7.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cleveland J. L., Huleihel M., Bressler P., Siebenlist U., Akiyama L., Eisenman R. N., Rapp U. R. Negative regulation of c-myc transcription involves myc family proteins. Oncogene Res. 1988;3(4):357–375. [PubMed] [Google Scholar]
- Cole M. D. The myc oncogene: its role in transformation and differentiation. Annu Rev Genet. 1986;20:361–384. doi: 10.1146/annurev.ge.20.120186.002045. [DOI] [PubMed] [Google Scholar]
- Coppola J. A., Cole M. D. Constitutive c-myc oncogene expression blocks mouse erythroleukaemia cell differentiation but not commitment. Nature. 1986 Apr 24;320(6064):760–763. doi: 10.1038/320760a0. [DOI] [PubMed] [Google Scholar]
- Croce C. M., Erikson J., ar-Rushdi A., Aden D., Nishikura K. Translocated c-myc oncogene of Burkitt lymphoma is transcribed in plasma cells and repressed in lymphoblastoid cells. Proc Natl Acad Sci U S A. 1984 May;81(10):3170–3174. doi: 10.1073/pnas.81.10.3170. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dani C., Blanchard J. M., Piechaczyk M., El Sabouty S., Marty L., Jeanteur P. Extreme instability of myc mRNA in normal and transformed human cells. Proc Natl Acad Sci U S A. 1984 Nov;81(22):7046–7050. doi: 10.1073/pnas.81.22.7046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dani C., Mechti N., Piechaczyk M., Lebleu B., Jeanteur P., Blanchard J. M. Increased rate of degradation of c-myc mRNA in interferon-treated Daudi cells. Proc Natl Acad Sci U S A. 1985 Aug;82(15):4896–4899. doi: 10.1073/pnas.82.15.4896. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean M., Levine R. A., Ran W., Kindy M. S., Sonenshein G. E., Campisi J. Regulation of c-myc transcription and mRNA abundance by serum growth factors and cell contact. J Biol Chem. 1986 Jul 15;261(20):9161–9166. [PubMed] [Google Scholar]
- Dony C., Kessel M., Gruss P. Post-transcriptional control of myc and p53 expression during differentiation of the embryonal carcinoma cell line F9. Nature. 1985 Oct 17;317(6038):636–639. doi: 10.1038/317636a0. [DOI] [PubMed] [Google Scholar]
- Dunnick W., Baumgartner J., Fradkin L., Schultz C., Szurek P. Methylation of plasmacytoma c-myc genes. Gene. 1985;39(2-3):287–292. doi: 10.1016/0378-1119(85)90325-7. [DOI] [PubMed] [Google Scholar]
- Dyson P. J., Littlewood T. D., Forster A., Rabbitts T. H. Chromatin structure of transcriptionally active and inactive human c-myc alleles. EMBO J. 1985 Nov;4(11):2885–2891. doi: 10.1002/j.1460-2075.1985.tb04018.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dyson P. J., Rabbitts T. H. Chromatin structure around the c-myc gene in Burkitt lymphomas with upstream and downstream translocation points. Proc Natl Acad Sci U S A. 1985 Apr;82(7):1984–1988. doi: 10.1073/pnas.82.7.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fahrlander P. D., Piechaczyk M., Marcu K. B. Chromatin structure of the murine c-myc locus: implications for the regulation of normal and chromosomally translocated genes. EMBO J. 1985 Dec 1;4(12):3195–3202. doi: 10.1002/j.1460-2075.1985.tb04065.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feo S., ar-Rushdi A., Huebner K., Finan J., Nowell P. C., Clarkson B., Croce C. M. Suppression of the normal mouse c-myc oncogene in human lymphoma cells. Nature. 1985 Feb 7;313(6002):493–495. doi: 10.1038/313493a0. [DOI] [PubMed] [Google Scholar]
- Greenberg M. E., Ziff E. B. Stimulation of 3T3 cells induces transcription of the c-fos proto-oncogene. Nature. 1984 Oct 4;311(5985):433–438. doi: 10.1038/311433a0. [DOI] [PubMed] [Google Scholar]
- Grosveld F., van Assendelft G. B., Greaves D. R., Kollias G. Position-independent, high-level expression of the human beta-globin gene in transgenic mice. Cell. 1987 Dec 24;51(6):975–985. doi: 10.1016/0092-8674(87)90584-8. [DOI] [PubMed] [Google Scholar]
- Hay N., Bishop J. M., Levens D. Regulatory elements that modulate expression of human c-myc. Genes Dev. 1987 Sep;1(7):659–671. doi: 10.1101/gad.1.7.659. [DOI] [PubMed] [Google Scholar]
- Jones T. R., Cole M. D. Rapid cytoplasmic turnover of c-myc mRNA: requirement of the 3' untranslated sequences. Mol Cell Biol. 1987 Dec;7(12):4513–4521. doi: 10.1128/mcb.7.12.4513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kakkis E., Calame K. A plasmacytoma-specific factor binds the c-myc promoter region. Proc Natl Acad Sci U S A. 1987 Oct;84(20):7031–7035. doi: 10.1073/pnas.84.20.7031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kakkis E., Prehn J., Calame K. An active chromatin structure acquired by translocated c-myc genes. Mol Cell Biol. 1986 Apr;6(4):1357–1361. doi: 10.1128/mcb.6.4.1357. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keath E. J., Caimi P. G., Cole M. D. Fibroblast lines expressing activated c-myc oncogenes are tumorigenic in nude mice and syngeneic animals. Cell. 1984 Dec;39(2 Pt 1):339–348. doi: 10.1016/0092-8674(84)90012-6. [DOI] [PubMed] [Google Scholar]
- Keath E. J., Kelekar A., Cole M. D. Transcriptional activation of the translocated c-myc oncogene in mouse plasmacytomas: similar RNA levels in tumor and proliferating normal cells. Cell. 1984 Jun;37(2):521–528. doi: 10.1016/0092-8674(84)90382-9. [DOI] [PubMed] [Google Scholar]
- Kelly K., Cochran B. H., Stiles C. D., Leder P. Cell-specific regulation of the c-myc gene by lymphocyte mitogens and platelet-derived growth factor. Cell. 1983 Dec;35(3 Pt 2):603–610. doi: 10.1016/0092-8674(83)90092-2. [DOI] [PubMed] [Google Scholar]
- Knight E., Jr, Anton E. D., Fahey D., Friedland B. K., Jonak G. J. Interferon regulates c-myc gene expression in Daudi cells at the post-transcriptional level. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1151–1154. doi: 10.1073/pnas.82.4.1151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lachman H. M., Skoultchi A. I. Expression of c-myc changes during differentiation of mouse erythroleukaemia cells. Nature. 1984 Aug 16;310(5978):592–594. doi: 10.1038/310592a0. [DOI] [PubMed] [Google Scholar]
- Leder P., Battey J., Lenoir G., Moulding C., Murphy W., Potter H., Stewart T., Taub R. Translocations among antibody genes in human cancer. Science. 1983 Nov 18;222(4625):765–771. doi: 10.1126/science.6356357. [DOI] [PubMed] [Google Scholar]
- Lipp M., Schilling R., Wiest S., Laux G., Bornkamm G. W. Target sequences for cis-acting regulation within the dual promoter of the human c-myc gene. Mol Cell Biol. 1987 Apr;7(4):1393–1400. doi: 10.1128/mcb.7.4.1393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mann R., Mulligan R. C., Baltimore D. Construction of a retrovirus packaging mutant and its use to produce helper-free defective retrovirus. Cell. 1983 May;33(1):153–159. doi: 10.1016/0092-8674(83)90344-6. [DOI] [PubMed] [Google Scholar]
- Mitchell R. L., Zokas L., Schreiber R. D., Verma I. M. Rapid induction of the expression of proto-oncogene fos during human monocytic differentiation. Cell. 1985 Jan;40(1):209–217. doi: 10.1016/0092-8674(85)90324-1. [DOI] [PubMed] [Google Scholar]
- Moore J. P., Todd J. A., Hesketh T. R., Metcalfe J. C. c-fos and c-myc gene activation, ionic signals, and DNA synthesis in thymocytes. J Biol Chem. 1986 Jun 25;261(18):8158–8162. [PubMed] [Google Scholar]
- Müller R., Curran T., Müller D., Guilbert L. Induction of c-fos during myelomonocytic differentiation and macrophage proliferation. Nature. 1985 Apr 11;314(6011):546–548. doi: 10.1038/314546a0. [DOI] [PubMed] [Google Scholar]
- Nepveu A., Marcu K. B. Intragenic pausing and anti-sense transcription within the murine c-myc locus. EMBO J. 1986 Nov;5(11):2859–2865. doi: 10.1002/j.1460-2075.1986.tb04580.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piechaczyk M., Yang J. Q., Blanchard J. M., Jeanteur P., Marcu K. B. Posttranscriptional mechanisms are responsible for accumulation of truncated c-myc RNAs in murine plasma cell tumors. Cell. 1985 Sep;42(2):589–597. doi: 10.1016/0092-8674(85)90116-3. [DOI] [PubMed] [Google Scholar]
- Rabbitts P. H., Forster A., Stinson M. A., Rabbitts T. H. Truncation of exon 1 from the c-myc gene results in prolonged c-myc mRNa stability. EMBO J. 1985 Dec 30;4(13B):3727–3733. doi: 10.1002/j.1460-2075.1985.tb04141.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Remmers E. F., Yang J. Q., Marcu K. B. A negative transcriptional control element located upstream of the murine c-myc gene. EMBO J. 1986 May;5(5):899–904. doi: 10.1002/j.1460-2075.1986.tb04301.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schuler G. D., Cole M. D. GM-CSF and oncogene mRNA stabilities are independently regulated in trans in a mouse monocytic tumor. Cell. 1988 Dec 23;55(6):1115–1122. doi: 10.1016/0092-8674(88)90256-5. [DOI] [PubMed] [Google Scholar]
- Siebenlist U., Bressler P., Kelly K. Two distinct mechanisms of transcriptional control operate on c-myc during differentiation of HL60 cells. Mol Cell Biol. 1988 Feb;8(2):867–874. doi: 10.1128/mcb.8.2.867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siebenlist U., Hennighausen L., Battey J., Leder P. Chromatin structure and protein binding in the putative regulatory region of the c-myc gene in Burkitt lymphoma. Cell. 1984 Jun;37(2):381–391. doi: 10.1016/0092-8674(84)90368-4. [DOI] [PubMed] [Google Scholar]
- Stanton L. W., Watt R., Marcu K. B. Translocation, breakage and truncated transcripts of c-myc oncogene in murine plasmacytomas. Nature. 1983 Jun 2;303(5916):401–406. doi: 10.1038/303401a0. [DOI] [PubMed] [Google Scholar]
- Taub R., Moulding C., Battey J., Murphy W., Vasicek T., Lenoir G. M., Leder P. Activation and somatic mutation of the translocated c-myc gene in burkitt lymphoma cells. Cell. 1984 Feb;36(2):339–348. doi: 10.1016/0092-8674(84)90227-7. [DOI] [PubMed] [Google Scholar]
- Tramontano D., Chin W. W., Moses A. C., Ingbar S. H. Thyrotropin and dibutyryl cyclic AMP increase levels of c-myc and c-fos mRNAs in cultured rat thyroid cells. J Biol Chem. 1986 Mar 25;261(9):3919–3922. [PubMed] [Google Scholar]
- Weintraub H. Assembly and propagation of repressed and depressed chromosomal states. Cell. 1985 Oct;42(3):705–711. doi: 10.1016/0092-8674(85)90267-3. [DOI] [PubMed] [Google Scholar]
- Westin E. H., Wong-Staal F., Gelmann E. P., Dalla-Favera R., Papas T. S., Lautenberger J. A., Eva A., Reddy E. P., Tronick S. R., Aaronson S. A. Expression of cellular homologues of retroviral onc genes in human hematopoietic cells. Proc Natl Acad Sci U S A. 1982 Apr;79(8):2490–2494. doi: 10.1073/pnas.79.8.2490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu C. The 5' ends of Drosophila heat shock genes in chromatin are hypersensitive to DNase I. Nature. 1980 Aug 28;286(5776):854–860. doi: 10.1038/286854a0. [DOI] [PubMed] [Google Scholar]
- Zinn K., DiMaio D., Maniatis T. Identification of two distinct regulatory regions adjacent to the human beta-interferon gene. Cell. 1983 Oct;34(3):865–879. doi: 10.1016/0092-8674(83)90544-5. [DOI] [PubMed] [Google Scholar]
- ar-Rushdi A., Nishikura K., Erikson J., Watt R., Rovera G., Croce C. M. Differential expression of the translocated and the untranslocated c-myc oncogene in Burkitt lymphoma. Science. 1983 Oct 28;222(4622):390–393. doi: 10.1126/science.6414084. [DOI] [PubMed] [Google Scholar]