Abstract
BACKGROUND & AIMS
Cholangiocarcinoma, the second most common liver cancer, can be classified as intra-hepatic cholangiocarcinoma (ICC) or extrahepatic cholangiocarcinoma. We performed an integrative genomic analysis of ICC samples from a large series of patients.
METHODS
We performed a gene expression profile, high-density single-nucleotide polymorphism array, and mutation analyses using formalin-fixed ICC samples from 149 patients. Associations with clinicopathologic traits and patient outcomes were examined for 119 cases. Class discovery was based on a non-negative matrix factorization algorithm and significant copy number variations were identified by GISTIC analysis. Gene set enrichment analysis was used to identify signaling pathways activated in specific molecular classes of tumors, and to analyze their genomic overlap with hepatocellular carcinoma (HCC).
RESULTS
We identified 2 main biological classes of ICC. The inflammation class (38% of ICCs) is characterized by activation of inflammatory signaling pathways, overexpression of cytokines, and STAT3 activation. The proliferation class (62%) is characterized by activation of oncogenic signaling pathways (including RAS, mitogen-activated protein kinase, and MET), DNA amplifications at 11q13.2, deletions at 14q22.1, mutations in KRAS and BRAF, and gene expression signatures previously associated with poor outcomes for patients with HCC. Copy number variation– based clustering was able to refine these molecular groups further. We identified high-level amplifications in 5 regions, including 1p13 (9%) and 11q13.2 (4%), and several focal deletions, such as 9p21.3 (18%) and 14q22.1 (12% in coding regions for the SAV1 tumor suppressor). In a complementary approach, we identified a gene expression signature that was associated with reduced survival times of patients with ICC; this signature was enriched in the proliferation class (P < .001).
CONCLUSIONS
We used an integrative genomic analysis to identify 2 classes of ICC. The proliferation class has specific copy number alterations, many features of the poor-prognosis signatures for HCC, and is associated with worse outcome. Different classes of ICC, based on molecular features, therefore might require different treatment approaches.
Keywords: Liver Cancer, Molecular Classification, Whole-Genome Profiling, Prognosis
Liver cancer is the third leading cause of cancer mortality worldwide, and cholangiocarcinoma represents the second most common type after hepatocellular carcinoma (HCC), accounting for 10%–15% of all primary liver malignancies.1 Cholangiocarcinoma is clinically classified into intrahepatic cholangiocarcinoma (ICC), arising from the small bile ducts within the liver, and extrahepatic cholangiocarcinoma (ECC), from the ductal epithelium of the extrahepatic bile duct. Both entities have distinct genetic and histologic features, risk factors, and clinical outcomes.2 Epidemiologic data suggest that ICC incidence has increased significantly in recent years, with great geographic variability, whereas ECC has remained stable or slightly declined.3 Surgical resection is the only potentially curative therapy for ICC patients, but most of them are diagnosed at an unresectable stage, for which only palliative chemotherapy is available.4 – 6
Chronic inflammation facilitates cholangiocyte transformation through a multistep process involving several factors, such as transforming growth factor-β and cytokines.7 In particular, interleukin (IL)-6 has been proposed as an important oncogenic driver in cholangiocarcinoma.8,9 Activating mutations of KRAS,10,11 promoter hypermethylation of CDKN2A,10 and overexpression of MET12 and HER213 also have been described. Nevertheless, there is limited understanding on the pathogenesis of this neoplasm, and no established molecular classification is available to date.
Unlike many other solid tumors, no molecular targeted agents have been approved for the treatment of this neoplasm. Phase III trials exploring molecular targeted therapies or chemotherapeutical regimens specific for ICC are lacking. Most studies include patients with biliary tract cancers, in which a small proportion are ICCs, as is the case with a recent trial testing cisplatin and gemcitabine.5 Sorafenib, a multiple tyrosine kinase inhibitor improving survival in advanced HCC patients,14 showed limited activity in such types of tumors in phase II studies.15 Hence, the identification of novel therapeutic targets remains an urgent need. To this end, genome-wide, unbiased characterization of molecular aberrations provides important information that would allow a better understanding of this disease. DNA copy number alterations represent a common mechanism driving oncogene and tumor-suppressor alterations. Consequently, a high-resolution genome-wide map should facilitate the identification of candidate drivers in ICC.
Herein, we apply genomic methodologies to archived fixed tumor tissues obtained from a large series of ICC patients with annotated clinical data. We report the presence of 2 highly distinct molecular classes identified through transcriptional profiling, suggest class-specific oncogenic mechanisms, and uncover their prognostic impact on clinical outcome.
Materials and Methods
Patients and Tumor Samples
A total of 153 formalin-fixed, paraffin-embedded (FFPE) samples were obtained from ICC patients resected between 1995 and 2007 at 3 centers from the HCC Genomic Consortium: IRCCS Istituto Nazionale Tumori (Milan), Mount Sinai School of Medicine (New York), and Hospital Clinic (Barcelona). Pathologic diagnosis of ICC was confirmed by 2 independent liver pathologists (M.S. and S.N.T.). A total of 149 good-quality samples were analyzed for molecular characterization to define molecular-based classes. Clinical follow-up data were available for 119 patients and were used to identify prognostic classes (Supplementary Figure 1). The study protocol was approved at each center’s Institutional Review Board.
Gene Expression, Single-Nucleotide Polymorphism Array, Fluorescence In Situ Hybridization, and Immunohistochemistry
See Supplementary Materials and Methods section.
Bioinformatics and Clinical Data Analysis
See Supplementary Materials and Methods section.
Results
Genomic Profiling Identifies 2 ICC Classes With Distinct Molecular Alterations
We performed whole-genome expression profiling in a final cohort of 149 ICC patients. The non-negative matrix factorization– based unsupervised clustering16 revealed 2 distinct and robust classes, which we named proliferation (92 of 149; 62%) and inflammation (57 of 149; 38%), based on subsequent functional characterization (Figure 1A, Supplementary Figure 2). Leave-one-out cross-validation analysis further confirmed the robustness of the non-negative matrix factorization– based classification (Supplementary Table 1) and identified 1402 overexpressed genes in the proliferation and 163 overexpressed genes in the inflammation classes (Supplementary Table 2). Patients in the proliferation class showed more aggressive tumors characterized by poorer histologic differentiation and intraneural invasion (Table 1). No significant differences were found for sex, age, tumor size, TNM stage, satellites, and macrovascular invasion.
Figure 1.
Molecular classes of ICC with distinct clinical outcome. Non-negative matrix factorization–based algorithm identified 2 robust classes: proliferation (orange) and inflammation (green) classes. (A) The heatmap shows unsupervised clustering of 149 ICCs based on the whole-genome expression. High and low expression levels are represented in red and blue, respectively. The top 100 differentially expressed genes are presented in the heat-map. (B) Kaplan–Meier estimates of overall survival (n = 119); (C) Kaplan–Meier plot of overall recurrence (n = 113). Patients in the proliferation class (red) showed shorter survival and earlier recurrence.
Table 1.
Clinicopathologic Characteristics of the Patients According to ICC Classes
| Proliferation class | Inflammation class | Total a | P value | |
|---|---|---|---|---|
| n (%) | 72 (60) | 47 (40) | 119 | |
| Demographics | ||||
| Gender, n (%) | ||||
| Male | 43 (60) | 22 (47) | 65 (55) | .19 |
| Age, years | ||||
| Median (IQR) | 65 (56–72) | 59 (52–68) | 64 (54–70) | .13 |
| Race, n (%) | ||||
| Caucasian | 62 (93) | 46 (98) | 108 (95) | .13 |
| African American | 1 (1.5) | 1 (2) | 2 (1.8) | |
| Asian | 3 (4.5) | 0 (0) | 3 (2.7) | |
| Viral hepatitis, n (%) | ||||
| Hepatitis C | 10 (14) | 9 (19) | 19 (16) | .44 |
| Hepatitis B | 5 (7) | 6 (13) | 11 (9) | .33 |
| Cirrhosisζ, n (%) | 12 (17) | 8 (17) | 20 (17) | 1.00 |
| Jaundice, n (%) | 2 (3) | 4 (9) | 6 (5) | .41 |
| Bilirubin, n (%) | ||||
| >1 mg/dL | 10 (14) | 14 (30) | 24 (20) | .06 |
| ALT, n (%) | ||||
| >40 IU/L | 21 (29) | 10 (21) | 31 (26) | .29 |
| Albumin, n (%) | ||||
| <35 mg/dL | 7 (10) | 3 (6) | 10 (8) | .74 |
| Tumor feautures (pathologic) | ||||
| Tumor diameter, cm | ||||
| Median (IQR) | 7 (4–10) | 6 (4–8.4) | 6 (4–9) | .12 |
| Tumor number, n (%) | ||||
| Single | 63 (88) | 36 (77) | 99 (83) | |
| Multiple | 9 (12) | 11 (23) | 20 (17) | .14 |
| Cell differentiation, n (%) | ||||
| Well | 4 (7) | 14 (38) | 18 (19) | |
| Moderate-poor | 54 (96) | 23 (62) | 77 (81) | <.01 |
| Stage,b n (%) | ||||
| I + II | 57 (79) | 37 (78) | 94 (79) | |
| III + IV | 15 (21) | 9 (19) | 24 (21) | 1.00 |
| Macrovascular invasion, n (%) | 12 (17) | 4 (9) | 15 (13) | .27 |
| Invasion of peritoneum, n (%) | 4 (6) | 0 (0) | 4 (3) | .15 |
| Infiltration of resection margins, n (%) | ||||
| Positive | 39 (54) | 15 (32) | 49 (45) | .05 |
| Invasion of bile duct, n (%) | 4 (6) | 2 (4) | 5 (5) | 1.00 |
| Intraneural invasion, n (%) | 18 (25) | 2 (4) | 17 (17) | <.01 |
| Satellites, n (%) | 20 (28) | 12 (26) | 29 (27) | .83 |
| Events, n (%) | ||||
| Recurrence | 45 (62) | 29 (62) | 74 (62) | .55 |
| Death | 50 (69) | 30 (64) | 80 (67) | .55 |
IQR, interquartile range.
Variables included here had less than 10% of missing values except for cell differentiation (n = 23, 19% missing).
Data according to the AJCC TNM stage 7th edition.
Pathways Activated in the Proliferation Class
Gene set enrichment analysis showed that several oncogenic pathways were enriched in the proliferation class, including receptor tyrosine kinase (RTK) pathways such as epidermal growth factor, RAS, AKT, MET, and the angiogenesis-related vascular endothelial growth factor and platelet-derived growth factor (q < 0.05, Supplementary Table 3). HRAS pathway activation (q = 0.02) was identified by Ingenuity Pathway Analysis with RAS homolog and RALA as the main molecules in the major network (Figure 2A). A network including HDAC1 and mitogen-activated protein kinase also was prominent in this class (Supplementary Figure 3). Top biofunctions are shown in Supplementary Figure 4. Afterward, we evaluated the genomic similarity between tumors in these classes with published HCC profiles.17–20 HCC signatures of poor prognosis were found enriched in the proliferation class, including cluster A,17 the cholangiocarcinoma-like class,18 G3 (cell-cycle– dysregulated class),19 transforming growth factor-β/WNT-activated class (S1 class),20 and the AFP-positive class (S2 class)20 (q < 0.10; Supplementary Table 4).
Figure 2.
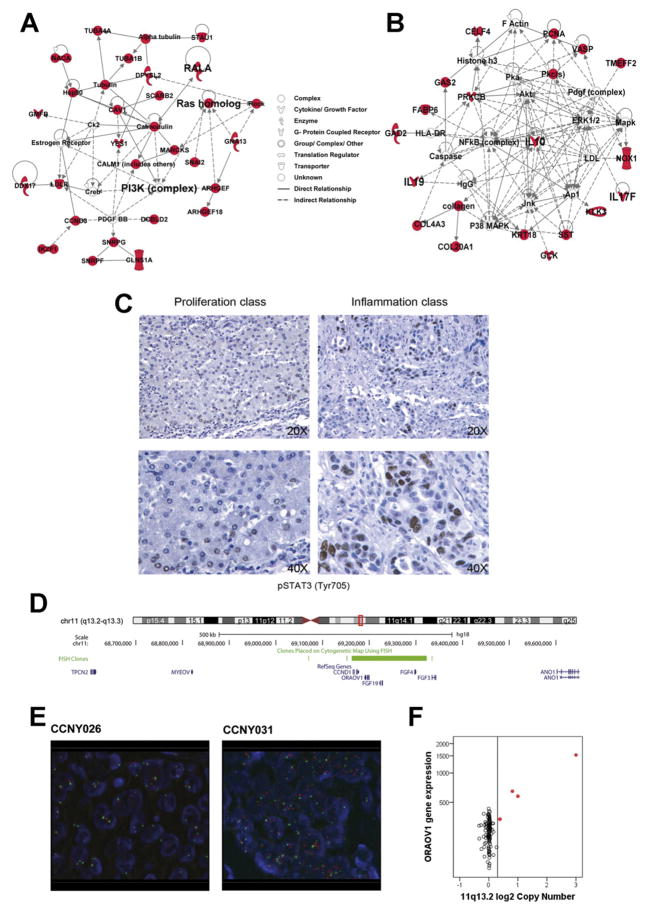
Dysregulated gene networks and validation of HLA at 11q13.2 in ICC molecular classes. Ingenuity Pathway Analysis results for (A) proliferation and (B) inflammation classes are presented. Network analysis of deregulated genes in the proliferation class shows the presence of 3 central nodes (ie, PI3K, RALA, and RAS homolog). In the inflammation class one top-scored network related to inflammation and including several cytokines (eg, IL-10, IL-19) was identified. A node represents a gene or gene product, and an edge indicates the relationship between nodes. Solid lines symbolize direct interaction and dotted lines stand for indirect interaction. (C) Immunohistochemical STAT3-Tyr705 phosphorylation staining in inflammation (right panel) and proliferation (left panel) class representative samples. Images were captured with 20× (upper panel) and 40× (lower panel) objectives. Nuclear positive staining was enriched in tumors of the inflammation class (77% positive patients in the inflammation vs 58% in the proliferation subclass). (D) Annotated genes mapped in the minimal overlapping region at 11q13.2 and the genomic location of the fluorescence in situ hybridization probe (green bar) used for confirmation of 11q13.2 HLA in 5 patients from the proliferation class. (E) HLA at 11q13.2 was validated by FISH. Red: BAC probe centered on the region of interest; green: BAC probe CEP-11 for the centromere of chromosome 11. Representative images are displayed: 1 patient without HLA (left panel) or with HLA (right panel). (F) ORAOV1 gene expression is increased in the 5 patients harboring 11q13.2 HLA. Each dot represents the corresponding copy number and expression level for a single tumor. Horizontal axis, log2 copy number for SNP probes in the minimal region at 11q13.2. Vertical axis, normalized gene expression level for each tumor.
Pathways Activated in the Inflammation Class
Tumors clustering within the inflammation class were characterized by induction of immune response–related pathways, such as dendritic cells and cytokine pathways (q < 0.05; Supplementary Table 5). Notably, overexpression of cytokines belonging to the Th2 subtype, such as IL-4 and IL-10, along with down-regulation of Th1 cytokines, was observed (Supplementary Figure 5A). Consistently, Ingenuity Pathway Analysis revealed a strong association with IL-17F, IL-17A, and chemokine signaling (Supplementary Figure 5B). One top scored network related to inflammation and IL-10 was identified (Figure 2B). Members of the STAT family are key transducers of cytokine signaling, especially the oncogene STAT3.21 Tumors in this class were enriched for nuclear pSTAT3 (77% vs 58% in the proliferation class; P = .02; Figure 2C). Immunostaining against the active subunit of nuclear factor-κB revealed that it was activated in 9%–11% of patients, with no differences between classes. Tumor-in-filtrating immune cells did not differ in number between our classes (data not shown).
Landscape of Chromosomal Aberrations in ICC
The GISTIC algorithm identified frequent and high-magnitude DNA copy number gain for chromosome 1q (32%) and 7p (25%), and loss for 6q (52%), 9q (45%), 9p (42%), 3p (41%), 13q (38%), 14q (36%), 8p (30%), 21q (20%), 17p (21%), 4q (18%), 1p (16%), 18q (15%), and 11p (13%), which were distributed equally among classes (Supplementary Figures 6 and 7A, Supplementary Table 6). We found 5 regions with high-level amplification (HLA), which affected 23% of tumors mostly from the proliferation class (Table 2, Supplementary Figure 7B). One HLA at 11q13.2, including 6 genes (ie, CCND1, FGF family members, ORAOV1) was observed in 5 patients of the proliferation class (5 of 128; 4%) and was confirmed by fluorescence in situ hybridization (Figure 2D and E, Supplementary Table 7). The ORAOV1 gene was overexpressed significantly in all samples (P = .001; Figure 2F). Another amplified locus was 1p13.1 (11 of 128; 9%), spanning several transcriptional repressors (eg, TRIM45, TTF2, and VTCN1; Table 2). In addition, HLAs at 1p31.3 and 6q12 were identified in 6% (8 of 128) and 5% (6 of 128) of patients, respectively. The former covers only ROR1, a poorly characterized RTK-like orphan receptor,22 whereas the latter includes PHF3, a PHD finger protein.
Table 2.
Most Relevant Copy Number Alterations in ICC Samples (n = 128)
| Cytoband | Gene no | Wide peak boundaries | High-level amplifications
|
|||
|---|---|---|---|---|---|---|
| Amplification q-value | Proliferation class (n = 88)
|
Inflammation class (n = 40)
|
All samples (n = 128)
|
|||
| Patients harboring amplification, n (%) | Patients harboring amplification, n (%) | Total frequency, n (%) | ||||
| 1p13.1 | TRIM45 | chr1:64134549–64223024 | 30.001 | 9 (10) | 2 (5) | 11 (9) |
| TTF2 | ||||||
| VTCN1 | ||||||
| hsa-mir-942 | ||||||
| 1p31.3 | ROR1 | chr1:117422805–117574724 | 0.15 | 6 (7) | 2 (5) | 8 (6) |
| 6q12 | PHF3 | chr6:64401474–64683286 | 0.15 | 6 (7) | 0 (0) | 6 (5) |
| 11q13.2 | CCND1 | chr11:68555478–69647117 | 30.001 | 5 (6) | 0 (0) | 5 (4) |
| GF3 | ||||||
| FGF4 | ||||||
| FGF19 | ||||||
| MYEOV | ||||||
| ORAOV1 | ||||||
| 12q12 | TMEM16A | chr12:43835040–43925043 | 0.15 | 7 (8) | 1 (3) | 8 (6) |
| TPCN2 | ||||||
| Focal deletions
|
||||||
| 9p21.3 | CDKN2A | chr9:21855843–22441030 | 5.36E-07 | 17 (19) | 6 (15) | 23 (18) |
| CDKNB | ||||||
| 9p21.3 | ELAVL2 | chr9:22441032–25673219 | 0.067876 | 11 (13) | 6 (15) | 17 (13) |
| TUSC1 | ||||||
| 10q23.2 | PTEN | chr10:89299367–90034324 | 0.10106 | 1 (1) | 2 (5) | 3 (2) |
| PAPSS2 | ||||||
| ATAD1 | ||||||
| 12p13.2 | BCL2L14 | chr12:11393730–12374584 | 0.17564 | 2 (2) | 2 (5) | 4 (3) |
| 14q22.1 | SAV1 | chr14:50157014–50265114 | 3.45E-08 | 11 (13) | 4 (10) | 15 (12) |
NOTE. See Supplementary Tables 8 and 9 for a comprehensive description of the 30 focal deletions identified.
Thirty regions showed focal deletions affecting 53% of ICCs, without apparent accumulation in either class (Supplementary Figure 7B and Supplementary Tables 8 and 9). Focal deletion at 9p21.3 was identified in 18% (23 of 128), a locus harboring CDKN2A and CDKN2B tumor suppressors (Table 2). Thirteen percent of patients (17 of 128) presented a second focal deletion at 9p21.3 covering 2 genes, ELAVL2 and TUSC1. A focal deletion at 12p13.2 was found in 3% of ICC patients (4 of 128; Table 2).23 Markedly, 1 of the 30 candidate regions covers only 1 gene, SAV1, within a minimal region from 50.16 to 50.26 Mb at 14q22.1 (15 of 128 patients; 11 from the proliferation class) and was associated significantly with reduced SAV1 gene expression (P = .006; Supplementary Figure 8A) This Hippo pathway gene plays a pivotal role in organ size control and tumor suppression.24 Expression of BIRC2 was overexpressed significantly in these samples, but no significant changes were observed for YAP1, LATS1/2, or BIRC5 (Supplementary Figure 8B–D).
Mutation Analysis
We sought to determine the association of previously reported mutations with the molecular classes. KRAS exon 1–activating mutations were observed in 12 patients (12 of 147; 8%), 8 from the proliferation class (8 of 90 vs 4 of 57 in the inflammation class; P = NS; Supplementary Table 10, Supplementary Figure 9A and B). Mutations at exon 15 of BRAF were found in 5 patients (5 of 141; 4%), 4 from the proliferation class (4 of 87 vs 1 of 54; P = NS, Supplementary Figure 9C and D). Three-point mutations were at V600E. Two mutations involving epidermal growth factor receptor (EGFR) (2 of 142; 1.4%) and 1 at exon 7 of TP53 all occurred in proliferation class patients.
Integration of Gene Expression, Copy Number, and Mutation Profiles Refine Proliferation and Inflammation Classes
The integration of DNA copy number variations (CNVs) and gene expression-based classes refined the class prediction. We found that the proliferation and inflammation classes could be divided further into 3 subgroups named P1–3 and I1–3, respectively (Figure 3, Supplementary Figure 10). Within the proliferation class (n = 87) we identified P1 (n = 35; 40%), P2 (n = 24; 28%), and P3 (n = 28; 32%). P1 was characterized by enrichment of KRAS mutations (6 of 35 vs 4 of 92 in the rest; P = .03), IGF1R and MET signaling pathways, and signatures of poor prognosis, including CC bad prognosis,25 G3,19 and S120 (false-discovery rate, < 0.25; Supplementary Table 11). A recently published CC signature with stem cell–like features26 was found enriched in P2 along with recurrence signatures. Finally, P3 showed chromosome 7p gains (17 of 28), EGFR overexpression (Supplementary Figure 11A and B; P < .001; Kruskal–Wallis test), EGFR mutations, and enrichment of an EGFR activation signature.27 Also, chromosome 1q gains (14 of 28 vs 27 of 99; P =.04) and several signatures of poor prognosis were found enriched in P3.
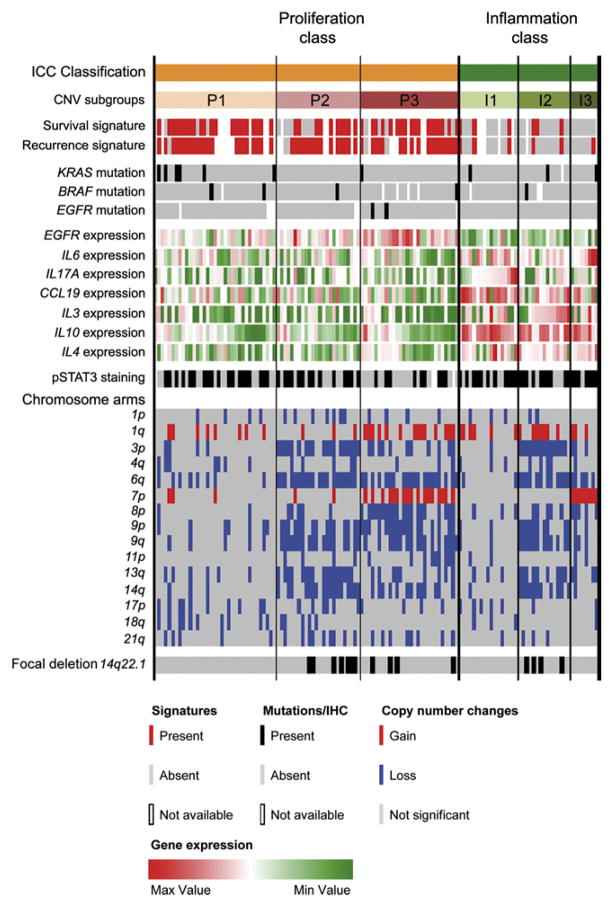
Figure 3.
CNV-based identification of further subgroups within the ICC classes. Non-negative matrix factorization clustering of copy number data identified the subgroups P1–3 and I1–3 within the proliferation and inflammation classes, respectively. Patients with copy number data are represented (n =127). Survival and recurrence signatures were enriched significantly in the proliferation class (patients in red, P < .001). Mutated patients are shown by black bars; KRAS mutations were enriched in the P1 subgroup (P = .03). Heatmap represents IL-6, EGFR, IL-17A, IL-3, IL-10, and I-L4 expression, highest in red and lowest in green; IL-6 and EGFR were found overexpressed in I3 and P3 (P <.001) whereas IL-17A and IL-3 were found overexpressed in I1 (P = .003) and I2 (P < .001) subgroups, respectively. CNV affecting broad arms are shown, gains are in red and losses are in blue. Only significant changes are represented. The presence of focal deletion of 14q22.1 and positive STAT3 staining is shown by black bars.
Within the inflammation class (n = 40), the subgroups I1 (n = 17; 42.5%), I2 (n = 15; 37.5%), and I3 (n = 8; 20%) were identified. The I3 subgroup presented with IL-6 gene overexpression (P = .001, Kruskal–Wallis test), and chromosome 7p gains (8 of 8 patients). I1 along with P1 present less chromosome instability as defined by the total number of arms gains and losses, with a mean of 2 events (either arms gain or loss) vs a mean of 6 events in the other subgroups (P < .001; Supplementary Figure 11C). Furthermore, even if the expression of most inflammation class–related cytokines, such as IL-10 and IL-4, was homogenous among I1–3 subgroups, I1 was characterized by IL-17A and CCL19 overexpression (P = .003 and P < .001 vs rest, respectively; Supplementary Figure 11D and E), whereas the I2 subgroup showed IL-3 over-expression (P < .001 vs rest; Supplementary Figure 11F).
Prognostic Relevance of the ICC Molecular Classification
Overall, characteristics of our cohort resemble those reported in previous studies.28 Patients were mostly Caucasian (108 of 119; 95%), with a similar gender distribution (65 of 119 males; 55%), a median age of 64 years, and without clinically significant underlying liver disease (cirrhosis in 17%; 20 of 119). On pathologic analysis, the median tumor size was 6 cm (25%–75%, 4 –9 cm), 17% (20 of 119) were multinodular, and 27% (29 of 119) had satellite lesions. Fifteen patients (13%) had macrovascular invasion, and 5 patients (5%) had bile duct invasion. Forty-nine patients (45%) had surgical margin infiltration. The median survival of the entire cohort was 34.3 months. One-, 3-, and 5-year overall survival rates were 85%, 49%, and 34%, respectively. The median time to recurrence in the cohort was 19 months; and 1-, 3-, and 5-year recurrence rates were 37%, 64%, and 71%, respectively. No differences were found between cirrhotic and noncirrhotic ICC patients regarding clinical behavior.
Survival Analysis
We sought to evaluate the prognostic impact of our molecular classes and focal copy number alterations along with known prognostic clinicopathologic variables. Patients in the proliferation class had significantly shorter survival (median, 24.3 vs 47.2 months in the inflammation class; P = .048; Figure 1B). Univariate analysis also identified sex, tumor size, tumor number, TNM stage, macrovascular invasion, invasion of peritoneum, intraneural invasion, and satellite lesions to be associated significantly with overall survival (Table 3). In multivariate analysis, male sex (hazard ratio [HR], 0.42; 95% confidence interval [CI], 0.25– 0.72; P < .01), tumor size (HR, 1.92; 95% CI, 1.13–3.27; P = .02), macrovascular invasion (HR, 3.17; 95% CI, 1.68 –5.95; P < .01), and satellite lesions (HR, 2.89; 95% CI, 1.69 – 4.91; P < .001) were independent predictors of overall survival (Table 3).
Table 3.
Univariate and Multivariate Analysis for Survival and Recurrence
| Variables | Overall survival
|
|
Overall recurrencea
|
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Univariate analysis
|
95% CI | Multivariate analysis
|
Univariate analysis
|
95% CI | Multivariate analysis
|
|||||||
| P value | HR | P value | HR | 95% CI | P value | HR | P value | HR | 95% CI | |||
| Male sex | <.01 | 0.47 | 0.30–0.75 | <.01 | 0.42 | 0.25–0.72 | .09 | 1.49 | 0.94–2.36 | |||
| Caucasian | .08 | 1.30 | 0.97–1.76 | .76 | 1.15 | 0.46–2.88 | ||||||
| Etiology, hepatitis B virus | .07 | 0.43 | 0.17–1.06 | .12 | 0.51 | 0.22–1.18 | ||||||
| Etiology, hepatitis C virus | .59 | 1.17 | 0.66–2.09 | .76 | 1.10 | 0.59–2.05 | ||||||
| Albumin level <3.5 g/L | .25 | 2.35 | 1.11–4.95 | .52 | 1.35 | 0.54–3.39 | ||||||
| Jaundice | .55 | 0.70 | 0.22–2.33 | .44 | 0.63 | 0.20–2.02 | ||||||
| Cirrhosis | .26 | 1.45 | 0.76–2.79 | .55 | 0.79 | 0.36–1.74 | ||||||
| Bilirubin level >1 mg/dL | .57 | 0.17 | 0.68–1.98 | .92 | 0.97 | 0.55–1.72 | ||||||
| ALT level > 40 IU/L | .75 | 1.09 | 0.64–1.84 | .82 | 1.06 | 0.63–1.80 | ||||||
| Tumor size > 6 cm | .02 | 1.69 | 1.08–2.63 | .02 | 1.92 | 1.13–3.27 | .02 | 1.79 | 1.12–2.85 | |||
| Tumor number | .03 | 1.82 | 1.07–3.08 | .48 | 1.25 | 0.67–2.33 | ||||||
| TNM stage (I + II vs III + IV) | <.001 | 2.50 | 1.49–4.18 | .04 | 1.82 | 1.04–3.19 | ||||||
| Macrovascular invasion | <.01 | 2.49 | 1.38–4.49 | <.01 | 3.17 | 1.68–5.95 | .04 | 2.00 | 1.04–3.82 | |||
| Invasion of visceral peritoneum | .02 | 4.25 | 1.32–13.63 | .02 | 4.03 | 1.25–12.94 | ||||||
| Intraneural invasion | <.01 | 2.41 | 1.40–4.15 | <.01 | 2.36 | 1.31–4.24 | .02 | 2.24 | 1.12–4.48 | |||
| Resection margins (positive) | .02 | 1.45 | 1.05–1.96 | .39 | 1.20 | 0.79–1.82 | ||||||
| Invasion of bile duct | .08 | 2.13 | 0.93–4.91 | .21 | 1.91 | 0.69–5.25 | ||||||
| Satellite lesions | <.001 | 3.75 | 2.34–5.98 | <.001 | 2.89 | 1.69–4.91 | <.001 | 3.87 | 2.31–6.48 | <.001 | 4.85 | 2.60–9.06 |
| ICC molecular classification | .048 | 1.59 | 1.00–2.51 | .03 | 1.68 | 1.05–2.69 | .04 | 1.81 | 1.01–3.23 | |||
| Focal deletion at 1p36.21 | .80 | 1.07 | 0.64–1.78 | .04 | 1.74 | 1.03–2.93 | ||||||
| Focal deletion at 3p25.3 | .87 | 1.06 | 0.52–2.14 | .04 | 1.91 | 0.99–3.68 | ||||||
| Focal deletion at 9p24.1 | .04 | 2.42 | 1.03–5.70 | .02 | 3.73 | 1.30–10.73 | ||||||
| Focal deletion at 14q22.1 | .59 | 1.21 | 0.60–2.45 | .01 | 2.55 | 1.28–5.10 | .01 | 3.58 | 1.73–7.38 | |||
CI, confidence interval.
Missing data for recurrence (n = 6).
Recurrence Analysis
Patients in the proliferation class had significantly shorter time to recurrence (median, 15 vs 37 months in the inflammation class; P = .03; Figure 1C). At the same time, patients harboring focal deletion at 14q22.1 recurred earlier with a median time of 9 vs 21 months in the rest of the patients (P = .01; Supplementary Figure 12). Other variables also were associated with recurrence at univariate analysis such as tumor size, TNM stage, macrovascular invasion, invasion of visceral peritoneum, and satellite lesions (Table 3). Two molecular variables (ICC molecular classification and focal deletion at 14q22.1) and 2 pathologic variables (intraneural invasion and satellite lesions) retained independent predictive value on multivariate analysis (Table 3).
We next used the recently published data on CC gene expression prognostic classes25 to understand whether there was an enrichment of our signature in those patients previously reported with poor prognosis (Supplementary Materials and Methods section). Effectively, 42 of 48 patients from the poor outcomes class showed an enrichment of our proliferation class as compared with 21 of 56 of the good prognosis class (P = .006; Supplementary Table 12).
Furthermore, we created a specific signature with the sole purpose of discriminating patients according to outcome (recurrence and survival), regardless of tumor molecular traits, by using the Cox score for feature selection (Supplementary Tables 13 and 14, Supplementary Figure 13) and successfully validated their performance in a previously published CC data set25 (Supplementary Table 15). Interestingly, the poor prognosis signature identified a subgroup of patients with significantly worse survival (P < .001), which were accumulated significantly in the proliferation class (P < .001; Figure 3).
Discussion
ICC lacks an established molecular classification based on high-throughput genomic data. Our findings provide compelling evidence of the existence of 2 clear-cut molecular ICC subtypes based on integration of whole-genome expression data, chromosomal aberrations, and signaling pathway activation. The molecular landscape of ICC entails one class—the proliferation ICC class— characterized by induction of cellular signals involved in cell-cycle progression and proliferation. This class shows aggressive clinical behavior and holds genomic resemblance to poor-prognosis HCC, suggesting a shared ancestral line between both entities. In the second class—the inflammation ICC class—there is an enrichment of inflammation-related pathways, mainly IL-10/-6 and STAT3 (Figure 4).
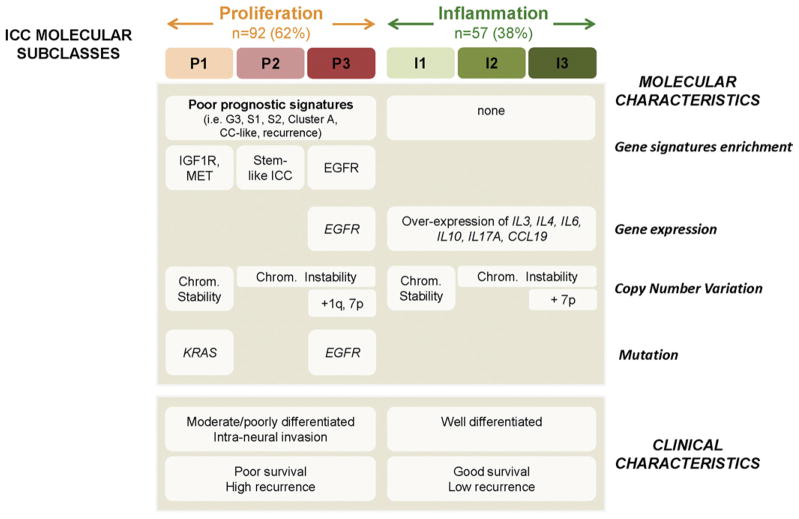
Figure 4.
Summary of characteristics of ICC classes. Specific molecular and clinical characteristics differ between ICC classes. Molecular characteristics such as signatures of poor prognosis (ie, cluster A,22 CC-like,23 G3,24 S1,25 S2,25 and stem-cell like ICC31), oncogenic pathways (ie, IGF1R, MET, EGFR), gene expression (ie, EGFR, ILs), copy number variations, and oncogenes mutations (KRAS and EGFR) are differentially enriched in the proliferation and inflammation classes. Clinical characteristics such as moderate/poorly differentiated tumors and intraneural invasion are more frequent in the proliferation class. Differences in survival and recurrence were observed.
Previous reports have suggested that inflammation significantly contributes to ICC pathogenesis.7 In addition, it has been suggested that STAT3 constitutive activation, dependent on IL-6 and IL-17 signaling, could contribute to cholangiocarcinoma growth.29,30 Even so, the immune system plays a crucial role in tumor immunosurveillance by destroying cancer cells. Hence, the tumor-progression activities and tumor-suppression actions of the immune system are not mutually exclusive.31 Here, we show that an inflammatory background, defined by humoral response and imbalance of cytokines, mainly defines a class of ICC with a more favorable prognosis. Notably, we found that this inflammation-related class was associated with enrichment of inflammation and cytokine pathway signatures and with overexpression of IL-6, IL-10, IL-17, and STAT3 constitutive activation. On the other hand, the proliferation class showed more aggressive behavior, reflected by earlier recurrence and an enrichment of several oncogenic pathways, such as RTK signaling and angiogenesis-related pathways, and HCC gene signatures of poor prognosis, such as cluster A,17 G3 proliferation,19 and S1 signature20 (Figure 4). One can speculate that liver progenitor cells might be the origin both entities, considering that these cells have bipotential capacity to differentiate into hepatic or biliary progenitor cells. Interestingly, the progenitor-cell subclass S2 signature in HCC was clearly enriched in this ICC class.20 This common progenitor cell of origin has been suggested previously, and reinforced by the existence of combined hepatocellular-cholangiocarcinoma cases.32 In addition, 5 patients belonging to this class showed HLA at 11q13.2, coding for CCND1, FGF19, and ORAOV1. We previously reported the same alteration in 6 of 89 HCV-related early HCC patients.33 The expression levels of ORAOV1 were overexpressed significantly in both ICC and HCC patients with 11q13 HLAs, suggesting that this oncogene could be a common driver in liver tumors. Clinical and outcome data of this study also point toward a more aggressive phenotype in ICC belonging to the proliferation class. These findings give further support to our hypothesis that molecular features associated with an aggressive phenotype are shared between the ICC proliferation class and HCC. Validation in patients from different geographic areas would be needed to confirm our molecular classification. Given the rarity of this cancer, a multi-institutional effort might be required to ensure the recruitment of a large validation series of surgically resected ICC samples.
The identification of 2 classes raises the obvious question about whether all ICC patients should be treated similarly, or if a specific subset of patients would respond differently to multikinase inhibitors. Certainly these data indicate that preclinical studies recapitulating these classes and testing multiple treatments should be conducted. First, we need to understand if multikinase inhibitors with a broad spectrum of action might be active in all ICCs or in a specific subclass, namely the proliferation class. Sorafenib, the current standard of care in advanced HCC patients,14 has shown marginal activity in cholangiocarcinoma patients compared with systemic chemotherapies.15 Nonetheless, considering the molecular similarities between the ICC proliferation class and the HCC signatures of poor prognosis, and KRAS enrichment, it is reasonable to test a multikinase inhibitor such as regorafenib, sorafenib, or sunitinib for this subpopulation. It has to be kept in mind that the downside of otherwise liver-toxic drugs in cirrhotic patients (as was the case with sunitinib in HCC) will be less relevant in ICC patients. In fact, ICCs often arise in a noncirrhotic liver in the absence of a clear etiologic risk factor or underlying liver disease. In our dataset, only 17% of patients had cirrhosis, 9% had HBV infection, and 16% had HCV infection without significant difference between our ICC classes.
Consistent with the class-specific therapeutic approach, JAK–STAT inhibitors might be specifically more active in the inflammation class. Targeting the IL-6/STAT3 pathway already has been proposed for the treatment of CC patients.34 Recently, in a phase I/II study, siltuximab, an anti–IL-6 monoclonal antibody, showed signs of efficacy in metastatic renal cell carcinoma.35 However, IL-6 is a pleiotropic cytokine and can generate functionally distinct or sometimes contradictory signals. Hence, therapies targeting IL-6 should be considered carefully and the development of more specific drugs remains an urgent need. Alternatively, the promising results obtained with STAT3 inhibitors in preclinical models of numerous cancers as well as early clinical trials in head and neck cancer suggest that STAT3 is an attractive therapeutic target. Similarly, dual JAK1–JAK 2 inhibitors have shown efficacy in myeloproliferative disorders.36 Thus, whether treatment of ICC should be tailored based on classes needs additional preclinical investigations.
To date, few genomic studies have been reported analyzing the whole-genome expression of ICC,25,37–39 and one study described a subgroup resembling stem-like HCCs.26 The most comprehensive study to date included both ICC and ECC and aimed to define an outcome-based classification of cholangiocarcinoma.25 The investigators suggested that patients belonging to the poor prognosis subclass, enriched with hilar-type tumors, had increased EGFR and HER-2/neu signaling and thus may benefit from treatment with lapatinib, a dual EGFR/HER2 tyrosine kinase inhibitor.25 It is important to keep in mind that a distinct pattern of genetic mutations as well as unique methylation and expression profiling has been identified differentiating ICC and ECCs.39 – 42 We found a significant enrichment of the signatures related to EGFR, HER2, and MET in our proliferation class, even though no amplification of these receptors was observed in our dataset, suggesting that mechanisms other than amplification (eg, epigenetic changes) could be involved in their activation. Particularly, signatures defining EGFR activation were specifically enriched in P3 of the proliferation subclass, along with up-regulation of EGFR and 2 receptor mutations. Furthermore, there is a significant enrichment of our proliferation signature in their reported class of poor prognosis,25 suggesting that our ICC signature might hold applicability to the general cholangiocarcinoma population.
A more compelling approach toward a more stratified treatment strategy would be to block specific oncogenic addiction loops.43 Cancer genetics has accelerated the time from driver mutation discovery to clinical proof-of-concept trials and regulatory approval. Two important examples have occurred recently: BRAF mutations were reported in melanoma44 and EML4-ALK translocations were reported in non–small-cell lung cancer.45 In both instances specific inhibitors such as PLX-4032 and crizotinib showed clinical efficacy.46,47 Our study provides some insight on potential driver mutation discovery that should be explored further by functional studies and early clinical testing. For instance, we described 4% BRAF mutations, mostly V600E, for which the specific inhibitor PLX-4032 has shown efficacy.46 Considering that approximately 70,000 new cases of ICC are diagnosed globally every year, this mutation might be involved in up to 3000 cases. Similarly, patients with FGF19 HLA (4%) could be treated with FGF inhibitors.
The feasibility of analyzing CNV in genomic DNA using FFPE tissues has been described.48 This may open new research frontiers by making large retrospective series of samples suitable for novel genomic studies. Here, we first tested a single-nucleotide polymorphism array platform to characterize broad and focal alterations in more than 100 ICC FFPE samples. We have to note that ICCs are desmoplastic cancers with marked cellular admixture and this is a fact that should be considered for the interpretation of results. Genome-wide CNV revealed a variety of chromosomal alterations heterogeneously distributed across our data set. Interestingly, CNV-based clustering identified further subgroups with specific molecular alterations within the gene expression-based classes (Figure 4). HLAs and focal deletions also were identified, providing the opportunity to pinpoint candidate regions harboring novel oncogenes and oncosuppressors. We found that 9% of patients harbored HLA at 1p13.1, spanning a number of transcriptional repressors such as VTCN1, TRIM45, and TTF2. VTCN1 recently was described as a B7 family co-regulator ligand, plays a crucial role in regulation of T-cell–mediated immunity, and has been associated with a more aggressive phenotype in several cancers.49,50 Six percent of patients harbored HLA at 1p31.3 covering only one candidate gene, ROR1. This gene is an orphan receptor that belongs to the evolutionarily conserved RTK family and, recently, it was found to be overexpressed in approximately 75% of cancer cell lines.22 Even if the role of ROR1 in disease is not yet clear, evidence suggesting its implication in MET signaling in human cancer has emerged.22 Focal deletions at cytoband covering already known oncosuppressors were found at 9p21.3 (CDKN2A, CDKN2B, and TUSC1). In addition, we found that 12% of patients showed a minimal common region of focal deletion at 14q22.1 covering only one gene, SAV1, the homolog of Drosophila Salvador. Recently, this gene has been proposed as the major suppressor of YAP, the main target of the Hippo pathway, in stem or progenitor cells, where it could act as an inhibitor of the inappropriate expansion of undifferentiated cells.24 Furthermore, SAV1 conditional mutant mice developed liver overgrowth and mixed hepatocholangiocarcinoma tumors.51 Functional loss of SAV1 results in the overexpression of the cell-cycle regulator cyclin E and anti-apoptotic proteins (eg, BIRC2 and BIRC5).51 In our dataset, focal deletion at 14q22.1 was associated significantly with SAV1 reduced gene expression, BIRC2 overexpression, and earlier recurrence. Thus, this gene may be relevant in a subset of ICC tumors and, taken together, these data point to SAV1 as a potentially pivotal alteration in the Hippo pathway. Functional validation of the role of these candidates is required before launching clinical trials.
The increasing incidence and poor outcome of ICC and the lack of molecular targeted therapies for this neoplasm represent major scientific challenges. The molecular classification proposed herein supports the existence of 2 distinct classes of patients that differ in their transcriptional, biological, and outcome traits. In addition, the copy number alterations provide insight on potential oncogenes and tumor suppressors as novel targets for therapies such as CCND1-p16, FGF19, ROR1, and SAV1, for which entire aberrant networks or specific genes could be targeted in personalized therapy.
Supplementary Material
Acknowledgments
The authors would like to thank Kensuke Kojima for his generous help.
Funding
Helena Cornella is supported by a fellowship from the Instituto de Salud Carlos III (ISCIII/FIS FI10/00143); Clara Alsinet is supported by a fellowship from Instituto de Salud Carlos III (ISCIII/FIS FI09/00605); Rameen Beroukhim is supported by grants from the US National Cancer Institute (K08CA122833 and U54CA143798) and a V Foundation Scholarship; Jordi Bruix is supported by a grant from the Instituto Carlos III (ISCIII/FIS PI 11/01830); Vincenzo Mazzaferro is supported by grants from the Italian Association for Cancer Research and the Oncology Research Project of the Italian Ministry of Health; Josep Llovet is supported by grants from the Asociación Española Contra el Cáncer, the US National Institute of Diabetes and Digestive and Kidney Diseases (1R01DK076986-01), European Commission-FP7 Framework (HEPTROMIC, proposal no. 259744), the Samuel Waxman Cancer Research Foundation, and the Spanish National Health Institute (SAF-2010-16055).
Abbreviations used in this paper
- CC
●●●
- CI
confidence interval
- CNV
copy number variations
- ECC
extrahepatic cholangiocarcinoma
- EGFR
epidermal growth factor receptor
- FFPE
formalin-fixed paraffin-embedded
- HCC
hepatocellular carcinoma
- HLA
high-level amplification
- HR
hazard ratio
- ICC
intrahepatic cholangiocarcinoma
- IL
interleukin
- RTK
receptor tyrosine kinase
- STAT
●●●
Footnotes
Note: To access the supplementary material accompanying this article, visit the online version of Gastroenterology at www.gastrojournal.org, and at http://dx.doi.org/10.1053/j.gastro.2013.01.001.
Microarray data GEO accession number: GSE33327.
Conflicts of interest
The authors disclose no conflicts.
References
- 1.Bleachacz BR, Gores GJ. Cholangiocarcinoma. Clin Liver Dis. 2008;12:131–150. doi: 10.1016/j.cld.2007.11.003. [DOI] [PubMed] [Google Scholar]
- 2.Sempoux C, Jibara G, Ward S, et al. Intrahepatic cholangiocarcinoma: new insights in pathology. Semin Liver Dis. 2011;31:104–110. doi: 10.1055/s-0031-1272839. [DOI] [PubMed] [Google Scholar]
- 3.Khan SA, Taylor-Robinson SD, Toledano MB, et al. Changing international trends in mortality rates for liver, biliary and pancreatic tumours. J Hepatol. 2002;37:806–813. doi: 10.1016/s0168-8278(02)00297-0. [DOI] [PubMed] [Google Scholar]
- 4.Singh P, Patel T. Advances in the diagnosis, evaluation and management of cholangiocarcinoma. Curr Opin Gastroenterol. 2006;22:294–299. doi: 10.1097/01.mog.0000218967.60633.64. [DOI] [PubMed] [Google Scholar]
- 5.Valle J, Wasan H, Palmer DH, et al. Cisplatin plus gemcitabine versus gemcitabine for biliary tract cancer. N Engl J Med. 2010;362:1273–1281. doi: 10.1056/NEJMoa0908721. [DOI] [PubMed] [Google Scholar]
- 6.Tan JC, Coburn NG, Baxter NN, et al. Surgical management of intrahepatic cholangiocarcinoma–a population-based study. Ann Surg Oncol. 2008;15:600–608. doi: 10.1245/s10434-007-9627-x. [DOI] [PubMed] [Google Scholar]
- 7.Khan SA, Thomas HC, Davidson BR, et al. Cholangiocarcinoma. Lancet. 2005;366:1303–1314. doi: 10.1016/S0140-6736(05)67530-7. [DOI] [PubMed] [Google Scholar]
- 8.Wehbe H, Henson R, Meng F, et al. Interleukin-6 contributes to growth in cholangiocarcinoma cells by aberrant promoter methylation and gene expression. Cancer Res. 2006;66:10517–10524. doi: 10.1158/0008-5472.CAN-06-2130. [DOI] [PubMed] [Google Scholar]
- 9.Isomoto H, Kobayashi S, Werneburg NW, et al. Interleukin 6 upregulates myeloid cell leukemia-1 expression through a STAT3 pathway in cholangiocarcinoma cells. Hepatology. 2005;42:1329–1338. doi: 10.1002/hep.20966. [DOI] [PubMed] [Google Scholar]
- 10.Tannapfel A, Benicke M, Katalinic A, et al. Frequency of p16 (INK4A) alterations and K-ras mutations in intrahepatic cholangiocarcinoma of the liver. Gut. 2000;47:721–727. doi: 10.1136/gut.47.5.721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Momoi H, Itoh T, Nozaki Y, et al. Microsatellite instability and alternative genetic pathway in intrahepatic chol angiocarcinoma. J Hepatol. 2001;35:235–244. doi: 10.1016/s0168-8278(01)00106-4. [DOI] [PubMed] [Google Scholar]
- 12.Terada T, Nakanuma Y, Sirica AE. Immunohistochemical demonstration of MET overexpression in human intrahepatic cholangiocarcinoma and in hepatolithiasis. Hum Pathol. 1998;29:175–180. doi: 10.1016/s0046-8177(98)90229-5. [DOI] [PubMed] [Google Scholar]
- 13.Terada T, Ashida K, Endo K, et al. c-erbB-2 protein is expressed in hepatolithiasis and cholangiocarcinoma. Histopathology. 1998;33:325–331. doi: 10.1046/j.1365-2559.1998.00496.x. [DOI] [PubMed] [Google Scholar]
- 14.Llovet JM, Ricci S, Mazzaferro V, et al. SHARP Investigators Study Group. Sorafenib in advanced hepatocellular carcinoma. N Engl J Med. 2008;359:378–390. doi: 10.1056/NEJMoa0708857. [DOI] [PubMed] [Google Scholar]
- 15.Bengala C, Bertolini F, Malavasi N, et al. Sorafenib in patients with advanced biliary tract carcinoma: a phase II trial. Br J Cancer. 2010;102:68–72. doi: 10.1038/sj.bjc.6605458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Brunet JP, Tamayo P, Golub TR, et al. Metagenes and molecular pattern discovery using matrix factorization. Proc Natl Acad Sci U S A. 2004;101:4164–4169. doi: 10.1073/pnas.0308531101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Lee JS, Chu IS, Heo J, et al. Classification and prediction of survival in hepatocellular carcinoma by gene expression profiling. Hepatology. 2004;40:667–676. doi: 10.1002/hep.20375. [DOI] [PubMed] [Google Scholar]
- 18.Woo HG, Lee JH, Yoon JH, et al. Identification of a cholangiocarcinoma-like gene expression trait in hepatocellular carcinoma. Cancer Res. 2010;70:3034–3041. doi: 10.1158/0008-5472.CAN-09-2823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Boyault S, Rickman DS, de Reyniès A, et al. Transcriptome classification of HCC is related to gene alterations and to new therapeutic targets. Hepatology. 2007;45:42–52. doi: 10.1002/hep.21467. [DOI] [PubMed] [Google Scholar]
- 20.Hoshida Y, Nijman SM, Kobayashi M, et al. Integrative transcriptome analysis reveals common molecular subclasses of human hepatocellular carcinoma. Cancer Res. 2009;69:7385–7392. doi: 10.1158/0008-5472.CAN-09-1089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Bromberg JF, Wrzeszczynska MH, Devgan G, et al. Stat3 as an oncogene. Cell. 1999;98:295–303. doi: 10.1016/s0092-8674(00)81959-5. [DOI] [PubMed] [Google Scholar]
- 22.Gentile A, Lazzari L, Benvenuti S, et al. Ror1 is a pseudokinase that is crucial for Met-driven tumorigenesis. Cancer Res. 2011;71:3132–3141. doi: 10.1158/0008-5472.CAN-10-2662. [DOI] [PubMed] [Google Scholar]
- 23.Montpetit A, Boily G, Sinnett D. A detailed transcriptional map of the chromosome 12p12 tumour suppressor locus. Eur J Hum Genet. 2002;10:62–71. doi: 10.1038/sj.ejhg.5200766. [DOI] [PubMed] [Google Scholar]
- 24.Lee KP, Lee JH, Kim TS, et al. The Hippo-Salvador pathway restrains hepatic oval cell proliferation, liver size, and liver tumorigenesis. Proc Natl Acad Sci U S A. 2010;107:8248–8253. doi: 10.1073/pnas.0912203107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Andersen JB, Spee B, Blechacz BR, et al. Genomic and genetic characterization of cholangiocarcinoma identifies therapeutic targets for tyrosine kinase inhibitors. Gastroenterology. 2012;142:1021–1031.e15. doi: 10.1053/j.gastro.2011.12.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Oishi N, Kumar MR, Roessler S, et al. Transcriptomic profiling reveals hepatic stem-like gene signatures and interplay of mir-200c and EMT in intrahepatic cholangiocarcinoma. Hepatology. 2012;56:1792–1803. doi: 10.1002/hep.25890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Zhang HT, Gorn M, Smith K, et al. Transcriptional profiling of human microvascular endothelial cells in the proliferative and quiescent state using cDNA arrays. Angiogenesis. 1999;3:211–219. doi: 10.1023/a:1009085422384. [DOI] [PubMed] [Google Scholar]
- 28.de Jong MC, Nathan H, Sotiropoulos GC, et al. Intrahepatic cholangiocarcinoma: an international multi-institutional analysis of prognostic factors and lymph node assessment. J Clin Oncol. 2011;29:3140–3145. doi: 10.1200/JCO.2011.35.6519. [DOI] [PubMed] [Google Scholar]
- 29.Wang L, Yi T, Kortylewski M, et al. IL-17 can promote tumor growth through an IL-6-Stat3 signaling pathway. J Exp Med. 2009;206:1457–1464. doi: 10.1084/jem.20090207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Isomoto H, Mott JL, Kobayashi S, et al. Sustained IL-6/STAT-3 signaling in cholangiocarcinoma cells due to SOCS-3 epigenetic silencing. Gastroenterology. 2007;132:384–396. doi: 10.1053/j.gastro.2006.10.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Schreiber RD, Old LJ, Smyth MJ. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science. 2011;331:1565–1570. doi: 10.1126/science.1203486. [DOI] [PubMed] [Google Scholar]
- 32.Roskams T. Liver stem cells and their implication in hepatocellular and cholangiocarcinoma. Oncogene. 2006;25:3818–3822. doi: 10.1038/sj.onc.1209558. [DOI] [PubMed] [Google Scholar]
- 33.Chiang DY, Villanueva A, Hoshida Y, et al. Focal gains of VEGFA and molecular classification of hepatocellular carcinoma. Cancer Res. 2008;68:6779–6788. doi: 10.1158/0008-5472.CAN-08-0742. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Blechacz B, Gores GJ. Cholangiocarcinoma: advances in pathogenesis, diagnosis, and treatment. Hepatology. 2008;48:308–321. doi: 10.1002/hep.22310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Rossi JF, Négrier S, James ND, et al. A phase I/II study of siltuximab (CNTO 328), an anti-interleukin-6 monoclonal antibody, in metastatic renal cell cancer. Br J Cancer. 2010;103:1154–1162. doi: 10.1038/sj.bjc.6605872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Verstovsek S, Kantarjian H, Mesa RA, et al. Safety and efficacy of INCB018424, a JAK1 and JAK2 inhibitor, in myelofibrosis. N Engl J Med. 2010;363:1117–1127. doi: 10.1056/NEJMoa1002028. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Wang AG, Yoon SY, Oh JH, et al. Identification of intrahepatic cholangiocarcinoma related genes by comparison with normal liver tissues using expressed sequence tags. Biochem Biophys Res Commun. 2006;345:1022–1032. doi: 10.1016/j.bbrc.2006.04.175. [DOI] [PubMed] [Google Scholar]
- 38.Jinawath N, Chamgramol Y, Furukawa Y, et al. Comparison of gene expression profiles between Opisthorchis viverrini and non-Opis-thorchis viverrini associated human intrahepatic cholangiocarcinoma. Hepatology. 2006;44:1025–1038. doi: 10.1002/hep.21330. [DOI] [PubMed] [Google Scholar]
- 39.Miller G, Socci ND, Dhall D, et al. Genome wide analysis and clinical correlation of chromosomal and transcriptional mutations in cancers of the biliary tract. J Exp Clin Cancer Res. 2009;28:62. doi: 10.1186/1756-9966-28-62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Yang B, House MG, Guo M, et al. Promoter methylation profiles of tumor suppressor genes in intrahepatic and extrahepatic cholangiocarcinoma. Mod Pathol. 2005;18:412–420. doi: 10.1038/modpathol.3800287. [DOI] [PubMed] [Google Scholar]
- 41.Karamitopoulou E, Tornillo L, Zlobec I, et al. Clinical significance of cell cycle- and apoptosis-related markers in biliary tract cancer: a tissue microarray-based approach revealing a distinctive immuno-phenotype for intrahepatic and extrahepatic cholangiocarcinomas. Am J Clin Pathol. 2008;130:780–786. doi: 10.1309/AJCP35FDCAVANWMM. [DOI] [PubMed] [Google Scholar]
- 42.Hezel AF, Deshpande V, Zhu AX. Genetics of biliary tract cancers and emerging targeted therapies. J Clin Oncol. 2010;28:3531–3540. doi: 10.1200/JCO.2009.27.4787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chin L, Andersen JN, Futreal PA. Cancer genomics: from discovery science to personalized medicine. Nat Med. 2011;17:297–303. doi: 10.1038/nm.2323. [DOI] [PubMed] [Google Scholar]
- 44.Davies H, Bignell GR, Cox C, et al. Mutations of the BRAF gene in human cancer. Nature. 2002;417:949–954. doi: 10.1038/nature00766. [DOI] [PubMed] [Google Scholar]
- 45.Soda M, Choi YL, Enomoto M, et al. Identification of the transforming EML4-ALK fusion gene in non-small-cell lung cancer. Nature. 2007;448:561–566. doi: 10.1038/nature05945. [DOI] [PubMed] [Google Scholar]
- 46.Flaherty KT, Puzanov I, Kim KB, et al. Inhibition of mutated, activated BRAF in metastatic melanoma. N Engl J Med. 2010;363:809–819. doi: 10.1056/NEJMoa1002011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Chabner BA. Early accelerated approval for highly targeted cancer drugs. N Engl J Med. 2011;364:1087–1089. doi: 10.1056/NEJMp1100548. [DOI] [PubMed] [Google Scholar]
- 48.Stokes A, Drozdov I, Guerra E, et al. Copy number and loss of heterozygosity detected by SNP array of formalin-fixed tissues using whole-genome amplification. PLoS One. 2011;6:e24503. doi: 10.1371/journal.pone.0024503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Quandt D, Fiedler E, Boettcher D, et al. B7-h4 expression in human melanoma: its association with patients’ survival and antitumor immune response. Clin Cancer Res. 2011;17:3100–3111. doi: 10.1158/1078-0432.CCR-10-2268. [DOI] [PubMed] [Google Scholar]
- 50.Krambeck AE, Thompson RH, Dong H, et al. B7-H4 expression in renal cell carcinoma and tumor vasculature: associations with cancer progression and survival. Proc Natl Acad Sci U S A. 2006;103:10391–10396. doi: 10.1073/pnas.0600937103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Lu L, Li Y, Kim SM, et al. Hippo signaling is a potent in vivo growth and tumor suppressor pathway in the mammalian liver. Proc Natl Acad Sci U S A. 2010;107:1437–1442. doi: 10.1073/pnas.0911427107. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.