Abstract
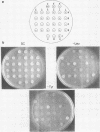
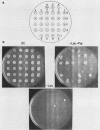
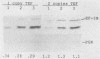
Changes in the dosage of genes encoding elongation factor EF-1 alpha were shown to cause parallel changes in the misreading of nonsense codons. Higher amounts of EF-1 alpha were correlated with increased nonsense suppression, suggesting that the level of EF-1 alpha is critically involved in translational fidelity.
Full text
PDF
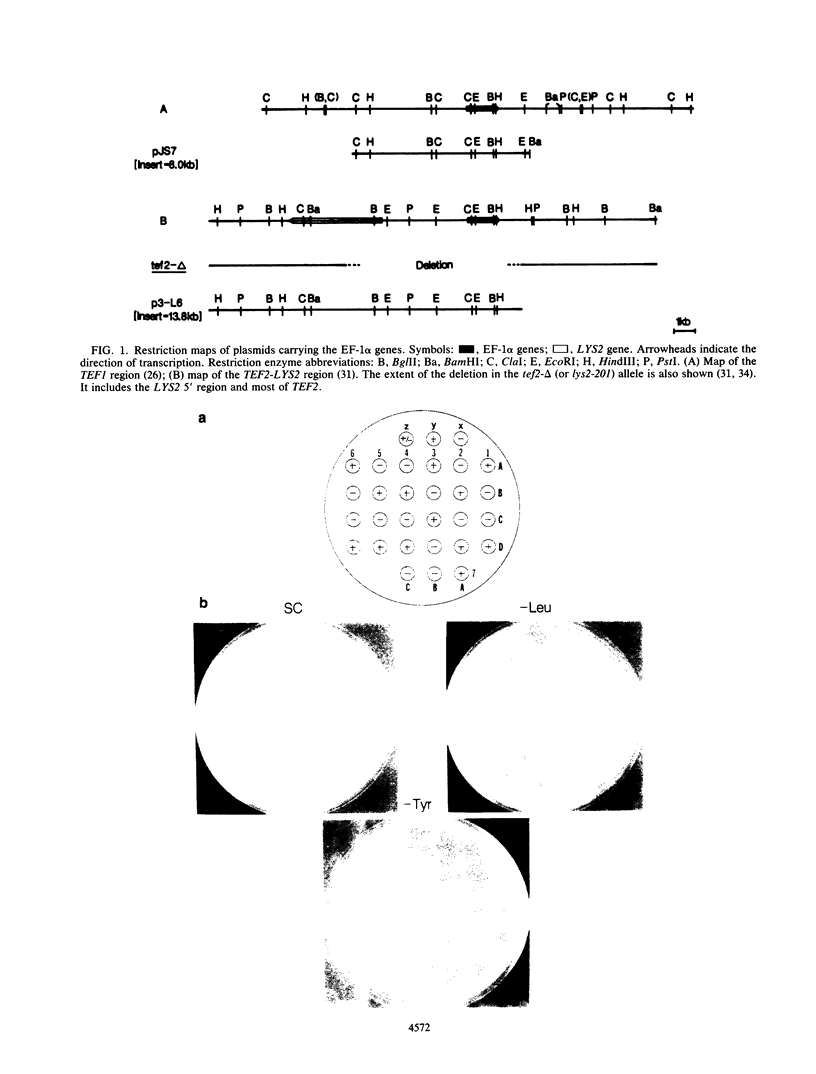
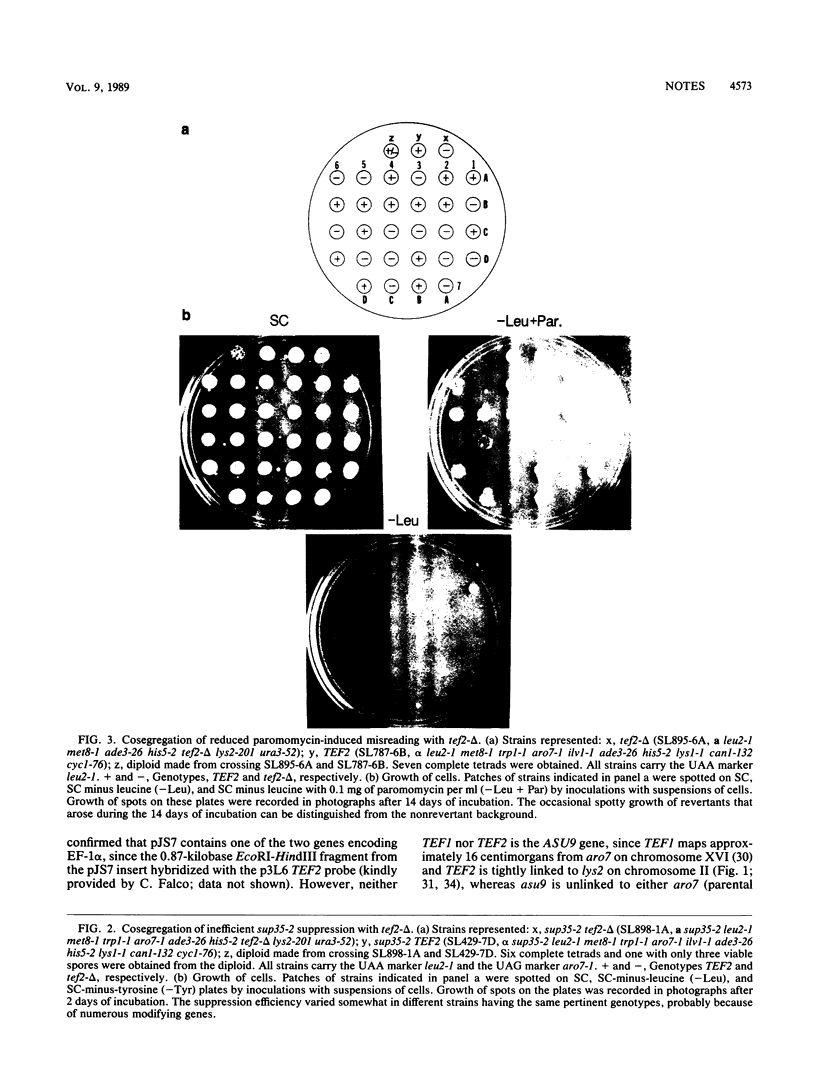




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bloom K. S., Fitzgerald-Hayes M., Carbon J. Structural analysis and sequence organization of yeast centromeres. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1175–1185. doi: 10.1101/sqb.1983.047.01.133. [DOI] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Breining P., Piepersberg W. Yeast omnipotent supressor SUP1 (SUP45): nucleotide sequence of the wildtype and a mutant gene. Nucleic Acids Res. 1986 Jul 11;14(13):5187–5197. doi: 10.1093/nar/14.13.5187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cottrelle P., Cool M., Thuriaux P., Price V. L., Thiele D., Buhler J. M., Fromageot P. Either one of the two yeast EF-1 alpha genes is required for cell viability. Curr Genet. 1985;9(8):693–697. doi: 10.1007/BF00449823. [DOI] [PubMed] [Google Scholar]
- Cottrelle P., Thiele D., Price V. L., Memet S., Micouin J. Y., Marck C., Buhler J. M., Sentenac A., Fromageot P. Cloning, nucleotide sequence, and expression of one of two genes coding for yeast elongation factor 1 alpha. J Biol Chem. 1985 Mar 10;260(5):3090–3096. [PubMed] [Google Scholar]
- Davydova E. K., Sitikov A. S., Ovchinnikov L. P. Phosphorylation of elongation factor 1 in polyribosome fraction of rabbit reticulocytes. FEBS Lett. 1984 Oct 29;176(2):401–405. doi: 10.1016/0014-5793(84)81206-5. [DOI] [PubMed] [Google Scholar]
- Erhart E., Hollenberg C. P. The presence of a defective LEU2 gene on 2 mu DNA recombinant plasmids of Saccharomyces cerevisiae is responsible for curing and high copy number. J Bacteriol. 1983 Nov;156(2):625–635. doi: 10.1128/jb.156.2.625-635.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grant C. M., Firoozan M., Tuite M. F. Mistranslation induces the heat-shock response in the yeast Saccharomyces cerevisiae. Mol Microbiol. 1989 Feb;3(2):215–220. doi: 10.1111/j.1365-2958.1989.tb01810.x. [DOI] [PubMed] [Google Scholar]
- Hawthorne D. C., Leupold U. Suppressors in yeast. Curr Top Microbiol Immunol. 1974;64(0):1–47. doi: 10.1007/978-3-642-65848-8_1. [DOI] [PubMed] [Google Scholar]
- Himmelfarb H. J., Maicas E., Friesen J. D. Isolation of the SUP45 omnipotent suppressor gene of Saccharomyces cerevisiae and characterization of its gene product. Mol Cell Biol. 1985 Apr;5(4):816–822. doi: 10.1128/mcb.5.4.816. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes D., Atkins J. F., Thompson S. Mutants of elongation factor Tu promote ribosomal frameshifting and nonsense readthrough. EMBO J. 1987 Dec 20;6(13):4235–4239. doi: 10.1002/j.1460-2075.1987.tb02772.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes D. Mutant forms of tufA and tufB independently suppress nonsense mutations. J Mol Biol. 1987 Oct 20;197(4):611–615. doi: 10.1016/0022-2836(87)90467-0. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaziro Y. The role of guanosine 5'-triphosphate in polypeptide chain elongation. Biochim Biophys Acta. 1978 Sep 21;505(1):95–127. doi: 10.1016/0304-4173(78)90009-5. [DOI] [PubMed] [Google Scholar]
- Kushnirov V. V., Ter-Avanesyan M. D., Telckov M. V., Surguchov A. P., Smirnov V. N., Inge-Vechtomov S. G. Nucleotide sequence of the SUP2 (SUP35) gene of Saccharomyces cerevisiae. Gene. 1988 Jun 15;66(1):45–54. doi: 10.1016/0378-1119(88)90223-5. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Liebman S. W., Cavenagh M. An antisuppressor that acts on omnipotent suppressors in yeast. Genetics. 1980 May;95(1):49–61. doi: 10.1093/genetics/95.1.49. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liebman S. W., Sherman F., Stewart J. W. Isolation and characterization of amber suppressors in yeast. Genetics. 1976 Feb;82(2):251–272. doi: 10.1093/genetics/82.2.251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mellor J., Dobson M. J., Roberts N. A., Tuite M. F., Emtage J. S., White S., Lowe P. A., Patel T., Kingsman A. J., Kingsman S. M. Efficient synthesis of enzymatically active calf chymosin in Saccharomyces cerevisiae. Gene. 1983 Sep;24(1):1–14. doi: 10.1016/0378-1119(83)90126-9. [DOI] [PubMed] [Google Scholar]
- Nagashima K., Kasai M., Nagata S., Kaziro Y. Structure of the two genes coding for polypeptide chain elongation factor 1 alpha (EF-1 alpha) from Saccharomyces cerevisiae. Gene. 1986;45(3):265–273. doi: 10.1016/0378-1119(86)90024-7. [DOI] [PubMed] [Google Scholar]
- Nagata S., Nagashima K., Tsunetsugu-Yokota Y., Fujimura K., Miyazaki M., Kaziro Y. Polypeptide chain elongation factor 1 alpha (EF-1 alpha) from yeast: nucleotide sequence of one of the two genes for EF-1 alpha from Saccharomyces cerevisiae. EMBO J. 1984 Aug;3(8):1825–1830. doi: 10.1002/j.1460-2075.1984.tb02053.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose M. D., Novick P., Thomas J. H., Botstein D., Fink G. R. A Saccharomyces cerevisiae genomic plasmid bank based on a centromere-containing shuttle vector. Gene. 1987;60(2-3):237–243. doi: 10.1016/0378-1119(87)90232-0. [DOI] [PubMed] [Google Scholar]
- Rose M., Winston F. Identification of a Ty insertion within the coding sequence of the S. cerevisiae URA3 gene. Mol Gen Genet. 1984;193(3):557–560. doi: 10.1007/BF00382100. [DOI] [PubMed] [Google Scholar]
- Ryazanov A. G., Shestakova E. A., Natapov P. G. Phosphorylation of elongation factor 2 by EF-2 kinase affects rate of translation. Nature. 1988 Jul 14;334(6178):170–173. doi: 10.1038/334170a0. [DOI] [PubMed] [Google Scholar]
- Sandbaken M. G., Culbertson M. R. Mutations in elongation factor EF-1 alpha affect the frequency of frameshifting and amino acid misincorporation in Saccharomyces cerevisiae. Genetics. 1988 Dec;120(4):923–934. doi: 10.1093/genetics/120.4.923. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schirmaier F., Philippsen P. Identification of two genes coding for the translation elongation factor EF-1 alpha of S. cerevisiae. EMBO J. 1984 Dec 20;3(13):3311–3315. doi: 10.1002/j.1460-2075.1984.tb02295.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simchen G., Winston F., Styles C. A., Fink G. R. Ty-mediated gene expression of the LYS2 and HIS4 genes of Saccharomyces cerevisiae is controlled by the same SPT genes. Proc Natl Acad Sci U S A. 1984 Apr;81(8):2431–2434. doi: 10.1073/pnas.81.8.2431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Spalding A., Tuite M. F. Host-plasmid interactions in Saccharomyces cerevisiae: effect of host ploidy on plasmid stability and copy number. J Gen Microbiol. 1989 Apr;135(4):1037–1045. doi: 10.1099/00221287-135-4-1037. [DOI] [PubMed] [Google Scholar]
- Tapio S., Kurland C. G. Mutant EF-Tu increases missense error in vitro. Mol Gen Genet. 1986 Oct;205(1):186–188. doi: 10.1007/BF02428051. [DOI] [PubMed] [Google Scholar]
- Thompson R. C. EFTu provides an internal kinetic standard for translational accuracy. Trends Biochem Sci. 1988 Mar;13(3):91–93. doi: 10.1016/0968-0004(88)90047-3. [DOI] [PubMed] [Google Scholar]
- Towbin H., Staehelin T., Gordon J. Electrophoretic transfer of proteins from polyacrylamide gels to nitrocellulose sheets: procedure and some applications. Proc Natl Acad Sci U S A. 1979 Sep;76(9):4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vijgenboom E., Vink T., Kraal B., Bosch L. Mutants of the elongation factor EF-Tu, a new class of nonsense suppressors. EMBO J. 1985 Apr;4(4):1049–1052. doi: 10.1002/j.1460-2075.1985.tb03737.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson P. G., Culbertson M. R. SUF12 suppressor protein of yeast. A fusion protein related to the EF-1 family of elongation factors. J Mol Biol. 1988 Feb 20;199(4):559–573. doi: 10.1016/0022-2836(88)90301-4. [DOI] [PubMed] [Google Scholar]