Abstract
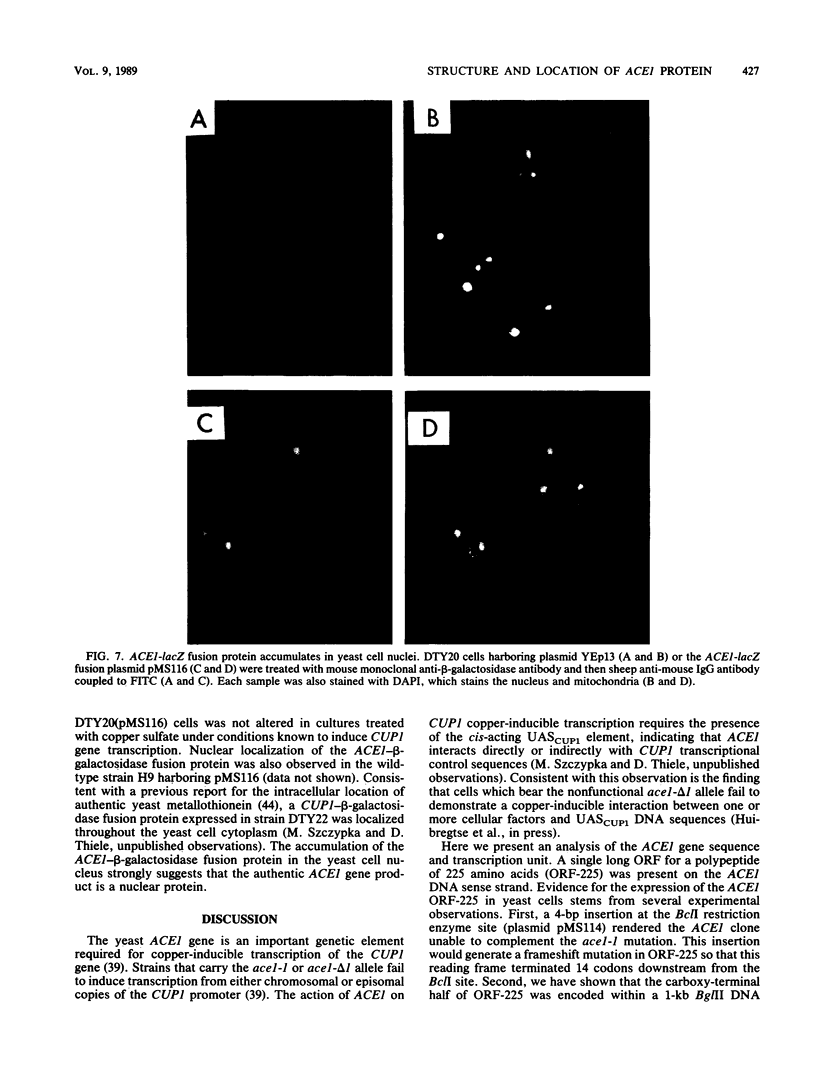
The ACE1 gene of the yeast Saccharomyces cerevisiae is required for copper-inducible transcription of the metallothionein gene (CUP1). The sequence of the cloned ACE1 gene predicted an open reading frame for translation of a 225-amino-acid polypeptide. This polypeptide was characterized by an amino-terminal half rich in cysteine residues and positively charged amino acids. The arrangement of many of the 12 cysteines in the configuration Cys-X-Cys or Cys-X-X-Cys suggested that the ACE1 protein may bind metal ions. The carboxyl-terminal half of the ACE1 protein was devoid of cysteines but was highly acidic in nature. The ability of a bifunctional ACE1-beta-galactosidase fusion protein to accumulate in yeast cell nuclei was consistent with the possibility that ACE1 plays a direct role in the regulation of copper-inducible transcription of the yeast metallothionein gene.
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adams A. E., Pringle J. R. Relationship of actin and tubulin distribution to bud growth in wild-type and morphogenetic-mutant Saccharomyces cerevisiae. J Cell Biol. 1984 Mar;98(3):934–945. doi: 10.1083/jcb.98.3.934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- BRENES-POMALES A., LINDEGREN G., LINDEGREN C. C. Gene control of coppersensitivity in Saccharomyces. Nature. 1955 Oct 29;176(4487):841–842. doi: 10.1038/176841a0. [DOI] [PubMed] [Google Scholar]
- Berg J. M. Potential metal-binding domains in nucleic acid binding proteins. Science. 1986 Apr 25;232(4749):485–487. doi: 10.1126/science.2421409. [DOI] [PubMed] [Google Scholar]
- Berg J. M. Proposed structure for the zinc-binding domains from transcription factor IIIA and related proteins. Proc Natl Acad Sci U S A. 1988 Jan;85(1):99–102. doi: 10.1073/pnas.85.1.99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biggin M. D., Gibson T. J., Hong G. F. Buffer gradient gels and 35S label as an aid to rapid DNA sequence determination. Proc Natl Acad Sci U S A. 1983 Jul;80(13):3963–3965. doi: 10.1073/pnas.80.13.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blumberg H., Hartshorne T. A., Young E. T. Regulation of expression and activity of the yeast transcription factor ADR1. Mol Cell Biol. 1988 May;8(5):1868–1876. doi: 10.1128/mcb.8.5.1868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Broach J. R. Construction of high copy yeast vectors using 2-microns circle sequences. Methods Enzymol. 1983;101:307–325. doi: 10.1016/0076-6879(83)01024-1. [DOI] [PubMed] [Google Scholar]
- Butt T. R., Ecker D. J. Yeast metallothionein and applications in biotechnology. Microbiol Rev. 1987 Sep;51(3):351–364. doi: 10.1128/mr.51.3.351-364.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butt T. R., Sternberg E. J., Gorman J. A., Clark P., Hamer D., Rosenberg M., Crooke S. T. Copper metallothionein of yeast, structure of the gene, and regulation of expression. Proc Natl Acad Sci U S A. 1984 Jun;81(11):3332–3336. doi: 10.1073/pnas.81.11.3332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Butt T. R., Sternberg E., Herd J., Crooke S. T. Cloning and expression of a yeast copper metallothionein gene. Gene. 1984 Jan;27(1):23–33. doi: 10.1016/0378-1119(84)90235-x. [DOI] [PubMed] [Google Scholar]
- Casadaban M. J., Martinez-Arias A., Shapira S. K., Chou J. Beta-galactosidase gene fusions for analyzing gene expression in escherichia coli and yeast. Methods Enzymol. 1983;100:293–308. doi: 10.1016/0076-6879(83)00063-4. [DOI] [PubMed] [Google Scholar]
- Dingwall C., Laskey R. A. Protein import into the cell nucleus. Annu Rev Cell Biol. 1986;2:367–390. doi: 10.1146/annurev.cb.02.110186.002055. [DOI] [PubMed] [Google Scholar]
- Ecker D. J., Butt T. R., Sternberg E. J., Neeper M. P., Debouck C., Gorman J. A., Crooke S. T. Yeast metallothionein function in metal ion detoxification. J Biol Chem. 1986 Dec 25;261(36):16895–16900. [PubMed] [Google Scholar]
- Fogel S., Welch J. W. Tandem gene amplification mediates copper resistance in yeast. Proc Natl Acad Sci U S A. 1982 Sep;79(17):5342–5346. doi: 10.1073/pnas.79.17.5342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guarente L. Yeast promoters and lacZ fusions designed to study expression of cloned genes in yeast. Methods Enzymol. 1983;101:181–191. doi: 10.1016/0076-6879(83)01013-7. [DOI] [PubMed] [Google Scholar]
- Hamer D. H., Thiele D. J., Lemontt J. E. Function and autoregulation of yeast copperthionein. Science. 1985 May 10;228(4700):685–690. doi: 10.1126/science.3887570. [DOI] [PubMed] [Google Scholar]
- Hanas J. S., Hazuda D. J., Bogenhagen D. F., Wu F. Y., Wu C. W. Xenopus transcription factor A requires zinc for binding to the 5 S RNA gene. J Biol Chem. 1983 Dec 10;258(23):14120–14125. [PubMed] [Google Scholar]
- Hannig E. M., Thiele D. J., Leibowitz M. J. Saccharomyces cerevisiae killer virus transcripts contain template-coded polyadenylate tracts. Mol Cell Biol. 1984 Jan;4(1):101–109. doi: 10.1128/mcb.4.1.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Huibregtse J. M., Engelke D. R., Thiele D. J. Copper-induced binding of cellular factors to yeast metallothionein upstream activation sequences. Proc Natl Acad Sci U S A. 1989 Jan;86(1):65–69. doi: 10.1073/pnas.86.1.65. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston M. A model fungal gene regulatory mechanism: the GAL genes of Saccharomyces cerevisiae. Microbiol Rev. 1987 Dec;51(4):458–476. doi: 10.1128/mr.51.4.458-476.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston M., Davis R. W. Sequences that regulate the divergent GAL1-GAL10 promoter in Saccharomyces cerevisiae. Mol Cell Biol. 1984 Aug;4(8):1440–1448. doi: 10.1128/mcb.4.8.1440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston M. Genetic evidence that zinc is an essential co-factor in the DNA binding domain of GAL4 protein. Nature. 1987 Jul 23;328(6128):353–355. doi: 10.1038/328353a0. [DOI] [PubMed] [Google Scholar]
- Kadonaga J. T., Carner K. R., Masiarz F. R., Tjian R. Isolation of cDNA encoding transcription factor Sp1 and functional analysis of the DNA binding domain. Cell. 1987 Dec 24;51(6):1079–1090. doi: 10.1016/0092-8674(87)90594-0. [DOI] [PubMed] [Google Scholar]
- Karin M., Najarian R., Haslinger A., Valenzuela P., Welch J., Fogel S. Primary structure and transcription of an amplified genetic locus: the CUP1 locus of yeast. Proc Natl Acad Sci U S A. 1984 Jan;81(2):337–341. doi: 10.1073/pnas.81.2.337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kilmartin J. V., Adams A. E. Structural rearrangements of tubulin and actin during the cell cycle of the yeast Saccharomyces. J Cell Biol. 1984 Mar;98(3):922–933. doi: 10.1083/jcb.98.3.922. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kozak M. Comparison of initiation of protein synthesis in procaryotes, eucaryotes, and organelles. Microbiol Rev. 1983 Mar;47(1):1–45. doi: 10.1128/mr.47.1.1-45.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kägi J. H., Kojima Y. Chemistry and biochemistry of metallothionein. Experientia Suppl. 1987;52:25–61. doi: 10.1007/978-3-0348-6784-9_3. [DOI] [PubMed] [Google Scholar]
- Laughon A., Gesteland R. F. Isolation and preliminary characterization of the GAL4 gene, a positive regulator of transcription in yeast. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6827–6831. doi: 10.1073/pnas.79.22.6827. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Messing J. New M13 vectors for cloning. Methods Enzymol. 1983;101:20–78. doi: 10.1016/0076-6879(83)01005-8. [DOI] [PubMed] [Google Scholar]
- Nagai K., Nakaseko Y., Nasmyth K., Rhodes D. Zinc-finger motifs expressed in E. coli and folded in vitro direct specific binding to DNA. Nature. 1988 Mar 17;332(6161):284–286. doi: 10.1038/332284a0. [DOI] [PubMed] [Google Scholar]
- Pinkham J. L., Olesen J. T., Guarente L. P. Sequence and nuclear localization of the Saccharomyces cerevisiae HAP2 protein, a transcriptional activator. Mol Cell Biol. 1987 Feb;7(2):578–585. doi: 10.1128/mcb.7.2.578. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silver P. A., Keegan L. P., Ptashne M. Amino terminus of the yeast GAL4 gene product is sufficient for nuclear localization. Proc Natl Acad Sci U S A. 1984 Oct;81(19):5951–5955. doi: 10.1073/pnas.81.19.5951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thiele D. J. ACE1 regulates expression of the Saccharomyces cerevisiae metallothionein gene. Mol Cell Biol. 1988 Jul;8(7):2745–2752. doi: 10.1128/mcb.8.7.2745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thiele D. J., Hamer D. H. Tandemly duplicated upstream control sequences mediate copper-induced transcription of the Saccharomyces cerevisiae copper-metallothionein gene. Mol Cell Biol. 1986 Apr;6(4):1158–1163. doi: 10.1128/mcb.6.4.1158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thiele D. J., Walling M. J., Hamer D. H. Mammalian metallothionein is functional in yeast. Science. 1986 Feb 21;231(4740):854–856. doi: 10.1126/science.3080806. [DOI] [PubMed] [Google Scholar]
- Welch J. W., Fogel S., Cathala G., Karin M. Industrial yeasts display tandem gene iteration at the CUP1 region. Mol Cell Biol. 1983 Aug;3(8):1353–1361. doi: 10.1128/mcb.3.8.1353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winge D. R., Nielson K. B., Gray W. R., Hamer D. H. Yeast metallothionein. Sequence and metal-binding properties. J Biol Chem. 1985 Nov 25;260(27):14464–14470. [PubMed] [Google Scholar]
- Wright C. F., McKenney K., Hamer D. H., Byrd J., Winge D. R. Structural and functional studies of the amino terminus of yeast metallothionein. J Biol Chem. 1987 Sep 25;262(27):12912–12919. [PubMed] [Google Scholar]