Abstract
Liver regeneration is of major clinical importance in the setting of liver injury, resection, and transplantation. A20, a potent anti-inflammatory and NF-κB inhibitory protein, has established pro-proliferative properties in hepatocytes, in part through decreasing expression of the Cyclin Dependent Kinase Inhibitor, p21. Both C-terminal (7-Zinc fingers; 7Zn) and N-terminal (Nter) domains of A20 were required to decrease p21 and inhibit NF-κB. However, both independently increased hepatocyte proliferation, suggesting that additional mechanisms contributed to the pro-proliferative function of A20 in hepatocytes. We ascribed one of A20’s pro-proliferative mechanisms to increased and sustained IL-6 induced STAT3 phosphorylation, as a result of decreased hepatocyte expression of the negative regulator of IL-6 signaling, SOCS3. This novel A20 function segregates with its 7Zn not Nter domain. Conversely, total and partial loss of A20 in hepatocytes increased SOCS3 expression, hampering IL-6-induced STAT3 phosphorylation. Following liver resection in mice pro-proliferative targets downstream of IL-6/STAT3 signaling were increased by A20 overexpression and decreased by A20 knockdown. In contrast, IL-6/STAT3 pro-inflammatory targets were increased in A20 deficient livers, and decreased or unchanged in A20 overexpressing livers. Upstream of SOCS3, levels of its microRNA regulator miR203 were significantly decreased in A20-deficient livers. Altogether these results demonstrate that A20 enhances IL-6/STAT3 pro-proliferative signals in hepatocytes by down-regulating SOCS3, likely through a miR203-dependent manner. This finding together with A20 reducing the levels of the potent cell cycle brake p21 establishes its pro-proliferative properties in hepatocytes and prompts the pursuit of A20-based therapies to promote liver regeneration and repair.
Keywords: Hepatocyte proliferation, tnfaip3, NF-κB, miR203, mutational analysis
The liver has a unique regenerative capacity, restoring liver mass after surgical resection or toxic/viral hepatocyte damage (1). Liver regeneration (LR) is characterized by hepatocytes’ rapid, synchronized transition from quiescence into cell cycle. Rodent models of partial hepatectomy (PH) unveiled molecular drivers of successful LR, such as the priming cytokines TNF and IL-6, that drive hepatocytes entry into cell cycle (1–2). Shortly following hepatectomy, Kupffer cells release TNF, activating NF-κB in neighboring hepatocytes, thus increasing IL-6 secretion. In turn, IL-6 binds to its receptor, activating JAK/STAT3 pathway to promote hepatocyte proliferation (3). Accordingly, the superagonistic IL-6/soluble IL-6R fusion protein enhances LR (4). In contrast, mice with targeted disruption of IL-6 demonstrate impaired regeneration that could be reversed by overexpression or induction of STAT3 (5–8). Levels of IL-6 after PH determine its hepatotrophic effect. Indeed, inappropriately high IL-6 levels rather inhibit LR by increasing expression of the Cyclin Dependent Kinase Inhibitor (CDKI), p21 (9). IL-6 signaling in hepatocytes is finely regulated by a negative feedback loop provided by IL-6/STAT3 mediated induction of suppressor of cytokine signal-3 (SOCS3) (10). Accordingly, hepatocyte-specific SOCS3 knockout (hKO SOCS3) mice show improved LR following hepatectomy (11). In sum, the IL-6/STAT3/SOCS3 pathway is a promising target for pro-regenerative strategies.
We previously demonstrated that A20/tnfaip3, a key element of the cellular response to injury and inflammation (12, 13), exerts multiple hepatoprotective functions through combined anti-inflammatory, anti-apoptotic, and pro-proliferative effects in hepatocytes (14–16). A20 restrains inflammation by inhibiting NF-κB activation (17), blocks apoptosis by disrupting caspase-8 activation (18), limits ischemic damage by increasing PPARα expression (16), optimizes energy production through improved lipid/fatty acid metabolism, and promotes proliferation by decreasing p21 (15, 19). A20’s hepatoprotective functions improve LR and survival in mouse models of extreme liver injury, including extended (78%, EH) and radical/lethal (83%) hepatectomy, acute toxic hepatitis and lethal ischemia/reperfusion injury (14–16). A20 is a NF-κB dependent gene (14) that is part of the regenerative response of the liver, and accordingly, its expression levels increase in deceased and living donor liver grafts hours following reperfusion (20). A20 KO mice are born cachectic and die within 4–5 weeks of birth mainly from unfettered liver inflammation, which shows A20’s high rank in the hierarchy of the liver physiologic anti-inflammatory armamentarium (21).
Since transcription of key priming cytokine IL-6 is in part NF-κB-dependent (22), we questioned whether the NF-κB inhibitory protein A20 would decrease IL-6 levels, thereby attenuating its pro-proliferative advantage in hepatocytes. This work examines this question.
Material and Methods
Reagents
Human recombinant TNF and IL-6 were from R&D Systems (Minneapolis, MN). Lipopolysaccharide (LPS), insulin, fetal bovine serum (FBS), gelatin, and collagenase type IV were from Sigma-Aldrich Co. (St. Louis, MO).
Mice
Eight to 12-week-old BALB/c mice (Taconic, Germantown, NY) and A20 heterozygous (HT) and wild type littermate (WT) mice were used in models of hepatectomy (21). Four to 5-week-old A20 WT, HT and KO mice were used for hepatocyte isolation. All procedures were performed in accordance with the U.S. Department of Health and Human Services Guide for the Care and Use of Laboratory Animals, and approved by the Institutional Committee for Use and Care of Laboratory Animals.
Cell Culture
Mouse normal liver epithelial cell line (NMuLi, CRL-1638), human hepatocellular carcinoma cell line (HepG2, HB-8065) and human kidney embryonic cell line (HEK-293) were purchased from American Type Culture Collection (Manassas, VA) (16). Mouse primary hepatocytes (MPH) were isolated using a modified 2-step EDTA/collagenase protocol (23).
Hepatocyte Proliferation
NMuLi and HepG2 hepatocytes were synchronized in G0/G1 phase of the cell cycle by 24h serum starvation. Cell proliferation was determined by cell count using Trypan blue exclusion before and 24h after addition of 10% FBS.
Western Blot Analysis
HepG2 and MPH whole cell lysates were recovered before and following IL-6, TNF and/or LPS treatment, and protein concentration determined (16). Samples were analyzed by Western Blot (WB) using the following primary antibodies: rabbit anti-STAT3, rabbit anti-IκBα, mouse anti-βactin, (Santa Cruz Biotechnology, Inc., Santa Cruz, CA), rabbit anti-phospho-STAT3 (P-STAT3 Tyr705) (Cell Signaling Technology, Danvers, MA), chicken anti-TNFAIP3 (A20) (Abcam Inc., Cambridge, MA), mouse anti-glyceraldehyde 3-phosphate dehydrogenase (GAPDH) (Calbiochem/EMD Biosciences Inc., San Diego, CA), anti-hemagglutinin (HA) (Roche Applied Science, Indianapolis, IN); and secondary antibodies (Thermo Scientific, Rockford, IL). Immunoblots were scanned and band intensity quantified by densitometry using ImageJ 1.41 (NIH, Bethesda, MD).
IL-6 Enzyme-linked Immunosorbent Assay
Supernatants from non-treated and LPS/TNF treated HepG2 and MPH cultures were assayed for IL-6 levels using human and mouse IL-6 ELISA Ready-SET-Go! (eBioscience, San Diego, CA). Results were normalized to total protein concentration.
Quantitative Reverse Transcriptase Polymerase Chain Reaction (qPCR)
RNA was extracted using Qiagen RNeasy Mini Kit (Qiagen, Valencia, CA) or TRIzol reagent (Sigma-Aldrich), and cDNA was synthesized using iScript cDNA synthesis kit (Bio-Rad). Gene expression was quantified using iTaq Fast SYBR Green Supermix with ROX (Bio-Rad) and gene specific primers (Invitrogen, Coralville, IA) listed in supplementary Table I, or TaqMan Mm00627280_m1 (tnfaip3), Mm00607939_s1 (βactin). Expression of target genes was normalized to that of the housekeeping genes βactin, TATA box binding protein (TBP) or 28S. miRNA was extracted using mirVana kit (Life Technologies, Grand Island, NY), and assayed for miR203, and the housekeeping miRNA, snoRNA202, using TaqMan (Applied Biosystems, Foster City, CA). qPCR were performed on a 7500 Fast Real-Time PCR System (Applied Biosystems).
Recombinant Adenoviruses
We generated recombinant adenovirus (rAd).A20 using a plasmid provided by Dr. V. Dixit (Genentench, San Francisco, CA) (24). The rAd.βgal was a gift of Dr. Robert Gerard (University of Texas SW, Dallas, Texas). By RT-PCR, we generated HA-tagged deletion mutants comprising the N-terminus (Nter) and seven Zinc (7Zn) domains of A20 and cloned them in pAC CMVpLpA SR(+) expression plasmid to generate rAd. (supplementary methods). We used HEK293 cells to generate, produce and titer rAd. that were purified by cesium chloride density gradient centrifugation for in vivo (24), or the AdenoPure LS Kit (Puresyn, Malvern, PA) for in vitro experiments. Hepatocyte cultures (60% confluent) were transduced with rAd. at a multiplicity of infection (MOI) of 50–200 plaque-forming units per cell (pfu/cell), leading to transgene expression in >95% of cells without toxicity (14, 15) (Fig. S1). In vivo, we injected 1×109 pfu of rAd. in 100 μL saline into the mouse penile vein. This dose and route of administration achieves maximal transgene expression in 30% of hepatocytes, 5 days after injection (15). Transgene expression was analyzed by WB (A20) and X-gal (5-bromo-4-chloro-3-indolyl-β-D-galactoside) staining (βgal).
Hepatectomy Model
78% hepatectomy (EH) was performed as described (15). Livers harvested before and after surgery were either frozen in liquid nitrogen for protein and RNA extraction, or fixed in 10% formalin for immunohistochemistry (IHC) and immunofluorescence (IF) analysis.
Immunochemistry and Immunofluorescence
For IHC and IF staining, we used the following primary antibodies: Goat anti-SOCS3, rabbit anti-P-STAT3 (Cell Signaling), rat anti-Ki67 (DAKO), chicken anti-albumin (Novus Biologicals, Littleton, CO), and goat anti-HNF4α (Santa Cruz), followed by HRP or Alexa Fluor 488 (green) and 594 (red) conjugated secondary antibodies (Invitrogen, Carlsbad, CA, USA). Ki67, P-STAT3 and SOCS3 positive cells per high power field (HPF) were counted using ImageJ automated or manual cell counting.
Statistical Analysis
Data were analyzed by one-way or two-way analysis of variance (ANOVA) followed by post-hoc Bonferroni multiple range test when F was significant, using Prism 5 (GraphPad Software, Inc., La Jolla, CA). Differences between groups were rated significant at a probability error (P) of less than 0.05.
Results
N-terminal and 7Zn C-terminal Domains of A20 Independently Promote Hepatocyte Proliferation in a p21-independent Fashion
We evaluated cell proliferation in non-transduced (C), rAd.A20, rAd.Nter, rAd.7n, and rAd.βgal-transduced NMuLi cells. This cell line responds in a physiologic manner to growth factor-induced cell cycle progression (15). Overexpression of A20 increased by 1.6 fold cell counts/well when compared to C and rAd.βgal transduced cells, 24h after addition of 10% FBS, (Fig. 1A, n=4–6; p<0.05). Similarly, rAd.Nter and rAd.7Zn-transduced cells showed a 1.8 and 1.9 fold increase in cell counts/well (Fig. 1A; n=3–4; p<0.05 vs. C and p<0.01 vs. rAd.βgal). This indicated that independent overexpression of Nter or 7Zn increases proliferation in NMuLi cells. We reproduced these results in HepG2 cells, validating their use in subsequent experiments (Fig. S2A; n=4; p<0.001).
Fig. 1. C-terminal and N-terminal domains of A20 independently promote hepatocyte proliferation but neither can independently decrease p21 expression or inhibit IκBα degradation.
A. NMuLi cells were transduced with rAd.A20, rAd.7Zn and rAd.Nter, serum starved for 24h to synchronize their cell cycle, then supplemented with 10% FBS enriched medium to drive cell proliferation. Cell count/well was evaluated 24h later by Trypan blue exclusion and plotted as mean ± SEM of 3–6 independent experiments. B. Relative p21 mRNA levels measured by qPCR in HepG2 cells transduced with rAd.A20, rAd.7Zn and rAd.Nter for 48h. Histograms represent mean± SEM of relative mRNA levels after normalization by βactin mRNA (n=3–5 independent experiments). C. Representative IκBα Western blot of cell lysates from rAd.A20, rAd.7Zn and rAd.Nter HepG2 cells treated with TNF (200 U/mL) for 15 min. βactin was used for loading control (n=3 independent experiments). Non-transduced (C) and rAd.βgal transduced cells were used as controls. *p<0.05, **p<0.01.
We had previously reported that A20’s pro-proliferative effect in hepatocytes related, at least in part, to decreased p21 expression (15). We confirmed in HepG2 that overexpression of full-length A20, but not Nter nor 7Zn, significantly decreased p21 mRNA levels as compared to βgal expressing cells (Fig 1B; n= 3–5; p<0.05). As for NF-κB inhibition (17) (Fig. 1C; n=3), cooperation between Nter and 7Zn domains was required to decrease p21, signifying that other mechanism(s) must account for their independent pro-proliferative effect in hepatocytes. Given potential discrepancies between cell lines and primary cells, we validated these results in MPH: full length A20 but neither 7Zn nor Nter decreased p21 mRNA levels (Fig. S2B; n=2; p<0.05), or inhibited TNF-induced IκBα degradation (Fig. S2C; n=3).
A20 Enhances IL-6/STAT3 Signaling Despite Overall Decreasing IL-6 production in Hepatocytes
Since IL-6 is central to hepatocyte proliferation, we measured IL-6 levels in supernatants of C, rAd.A20, rAd.Nter, rAd.7Zn, and rAd.βgal transduced HepG2 stimulated with TNF (200 U/mL) and LPS (10 μg/mL) for 6h, as to mimic the physiologic triggers of IL-6 secretion after hepatectomy (1). IL-6 levels significantly increased in all groups following TNF/LPS, as compared to corresponding non-stimulated cells (6.5 to 9.9 fold, Fig. 2A). However, IL-6 levels were significantly lower in supernatants of A20 overexpressing HepG2 compared to all other groups (Fig. 2A; n=4–7; p<0.01 vs. C; and p<0.001 vs. rAd.Nter, rAd.7Zn, and rAd.βgal). This result indicates that IL-6 transcription partially relies on NF-κB (22). Notably, neither Nter nor 7Zn decreased TNF/LPS-induced IL-6 secretion, with 7Zn overexpression moderately, yet significantly increasing it (Fig. 2A; p<0.01 vs. C).
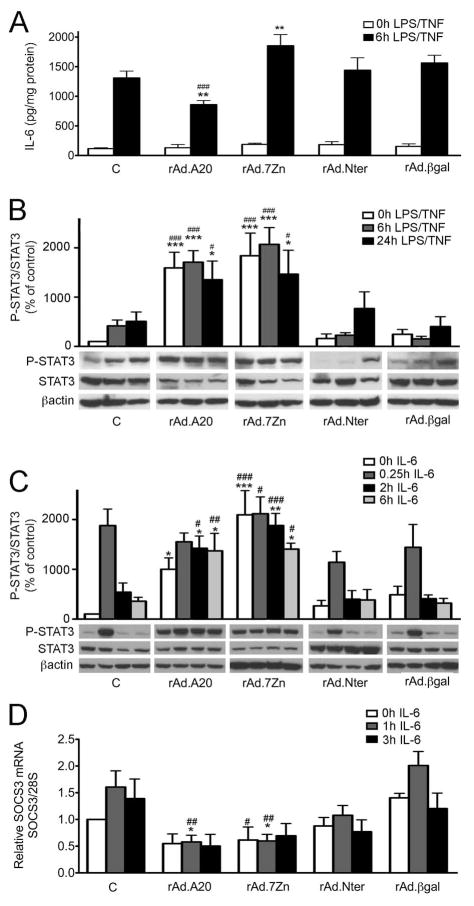
Fig. 2. Overexpression of A20 increases IL-6 induced STAT3 phosphorylation through down-regulation of SOCS3 expression, despite lower IL-6 production.
HepG2 were non-transduced (C), or transduced with rAd.A20, rAd.7Zn, rAd.Nter, and control rAd.βgal A. IL-6 levels were determined by ELISA in cell culture supernatants 6h following TNF (200 U/mL) and LPS (10ng/mL). Results are expressed as mean ± SEM of 4–7 independent experiments. Representative phospho (P-STAT3) and total STAT-3 Western blots following B. TNF (200 U/mL) and LPS (10ng/mL) for 6 to 24 h or C. IL-6 (50 ng/mL) for 15 min to 6 h. βactin was used as loading control. Corrected densitometry results, presented as percentage of control non-stimulated cells, are expressed as mean ± SEM of 3–5 independent experiments. D. Relative SOCS3 mRNA levels were measured by qPCR 1h to 3h following IL-6 (50 ng/mL). Histograms represent mean ± SEM of relative mRNA levels after normalization with 28S (n=4–5 independent experiments). *p<0.05, **p<0.01 and ***p< 0.001 vs. C, and #p<0.05, ##p<0.01 and ###p< 0.001 vs. rAd.βgal within each treatment group.
Despite lower IL-6 levels in supernatants of A20 overexpressing cells, STAT3 phosphorylation, downstream of IL-6, was enhanced at baseline (~150 fold; p<0.001), 6h (p<0.001) and 24h (p<0.05) after LPS/TNF treatment, when compared to C and rAd.βgal-transduced cells (Fig. 2B; n=3–4). Like A20, overexpression of 7Zn but not Nter, also increased STAT3 phosphorylation, reflecting either higher production of IL-6 by 7Zn expressing cells, or that A20’s 7Zn domain accounts for its ability to increase STAT3 phosphorylation.
To clarify this issue, we washed out the basal medium of C and rAd. transduced (A20, Nter, 7Zn, βgal) HepG2 cell cultures, then treated them with exogenous IL-6 (50 ng/mL) and checked for STAT3 phosphorylation 15min to 6h later. Control, rAd.Nter, and rAd.βgal transduced HepG2 showed low basal P-STAT3 levels, that transiently increased (peaking 15 min) after IL-6 stimulation. A20 and 7Zn overexpressing HepG2 had significantly higher baseline levels of P-STAT3 (comparable to IL-6 induced peak levels) that were slightly enhanced and sustained for at least 6h after IL-6 addition (Fig. 2C; n=4–5). These results indicate that this novel effect of A20 indeed maps to its 7Zn domain.
To investigate the molecular basis for A20-mediated increase in STAT3 phosphorylation, we assessed STAT3-dependent expression of the negative regulator of IL-6 signaling, SOCS3. Our results showed that both A20 and 7Zn, but not Nter, significantly decreased basal and IL-6-induced upregulation of SOCS3 mRNA in HepG2 cells, compared to controls (Fig. 2D; n=4–5; p<0.05 vs. C and p<0.01 vs. rAd.βgal). Altogether, these results uncover a novel mechanism by which A20 (7Zn domain) promotes hepatocyte proliferation through decreasing SOCS3 expression.
A20 Knockdown Increases SOCS3 Expression and Attenuates IL-6/STAT3 Signaling, Despite Increasing IL-6 production in Hepatocytes
To investigate the physiologic role of A20 in regulating IL-6/STAT3/SOCS3 signaling, we performed loss of function experiments, using MPH isolated from A20 KO, A20 HT and WT littermate mice. We confirmed by qPCR that A20 mRNA was absent in A20 KO, and reduced by 50% in A20 HT MPH, as compared to WT (Fig. 3A; n=3; p<0.001). Total loss of A20 significantly increased basal (p<0.05) and TNF-induced (p<0.001) IL-6 production by MPH, when compared to HT and WT (Fig. 3B; n=4). Heterozygous MPH showed an intermediate result. Increased basal IL-6 levels in A20 KO and HT hepatocytes were paralleled by higher basal P-STAT3 levels, indicating a chronic state of IL-6-mediated activation of these hepatocytes (Fig. S3; n=2). However, when we washed away endogenously produced IL-6 prior to adding exogenous IL-6 (50ng/mL), STAT3 phosphorylation was almost abolished in A20 KO, and attenuated (but with similar kinetics) in A20 HT MPH, as compared to WT (Fig. 3C; n=3). Decreased STAT3 phosphorylation in A20 KO MPH correlated with significantly higher basal (p<0.05) and IL-6 induced (p<0.01 at 1h, p<0.05 at 3h) SOCS3 mRNA levels, compared to WT (Fig. 3D; n= 4–5). We obtained similar results in whole livers, when hepatocytes where still in their physiologic multicellular environment; SOCS3 mRNA levels were significantly higher in KO vs. HT (p<0.001) and WT (p<0.01), and slightly greater in HT vs. WT livers (Fig. 4A; n=7–9). Consequently, we noted significantly more SOCS3/albumin positive hepatocytes around the portal and central veins of A20 KO livers, as compared to HT and WT livers (Fig. 4B and 4C; n=3–4; p<0.01). In line with in vitro results, the number of P-STAT3/HNF4α positive hepatocytes, at baseline, was higher in A20 KO than in WT and HT livers (Fig. 4D and 4E; n=3–4; p<0.01, and <0.05, respectively). Despite increased P-STAT3 levels in A20 KO livers, the number of Ki67 proliferating hepatocytes was not significantly different among groups (Fig. S4C and S4D; n=3–4). We attribute this result to very high IL-6 levels in A20 KO mice, creating an IL-6 “hyperstimulation” outcome, i.e. impaired hepatocyte proliferation despite increased P-STAT3 levels (9). Indeed, intra-hepatic IL-6 levels were significantly higher in A20 KO vs. HT and WT livers (Fig. S4A; n=5; p<0.001), with a corresponding increase in mRNA levels of STAT3-dependent cell cycle inhibitor, p21 (Fig. S4B; n=5; p<0.05). We verified by qPCR that A20 mRNA levels were absent or reduced by 50% in A20 KO and HT livers respectively, as compared to WT (Fig. S4E; n=4–6). Since SOCS3 mRNA levels are also epigenetically regulated by miR203 (26), we checked by qPCR for miR203 levels and found them significantly lower in KO (p<0.01) and HT (p<0.05) vs. WT livers (Fig. 4F; n=5–7).
Fig. 3. A20 knockdown increases IL-6 production but decreases IL-6 induced STAT3 phosphorylation by up-regulating SOCS3.

A. A20 mRNA levels in mouse primary hepatocytes (MPH) isolated from WT, A20 HT and A20 KO livers, as measured by qPCR. Histograms represent mean ± SEM of relative A20 mRNA levels after normalization with βactin (n=3 mice per group). B. IL-6 levels determined by ELISA in cell culture supernatants of MPH 6h following TNF (200 U/mL). Results are expressed as mean ± SEM of 3–6 independent experiments. C. Representative phospho (P-STAT3) and total STAT-3 Western blots of MPH treated with IL-6 (50 ng/mL) for 30 min to 6 h. GAPDH was used as loading control. Corrected densitometry results, presented as percentage of WT non-stimulated cells are expressed as mean ± SEM of 3 independent experiments. D. Relative SOCS3 mRNA levels measured by qPCR in MPH cultures stimulated for 1h to 3h with IL-6 (50ng/mL). Histograms represent mean± SEM of relative mRNA levels after normalization with βactin (n= 4–5 independent experiments). *p<0.05, **p<0.01 and ***p< 0.001.
Fig. 4. A20 knockdown increases SOCS3 expression, which inversely correlates with lower miR-203 levels.
Livers of 4–5 weeks old WT, A20 HT and A20 KO mice were analyzed for A. SOCS3 mRNA levels by qPCR. Histograms represent mean ± SEM of relative SOCS3 mRNA levels after normalization with βactin (n=7–9 mice per group). B. Graph representing mean±SEM of SOCS3 positive hepatocytes per high power field (HPF), as analyzed C. by immunohistochemistry (brown). Double immunofluorescence staining, using fluorescence labeled SOCS3 and hepatocyte specific albumin antibodies (SOCS3: green; albumin: red; DAPI: blue), confirmed that SOCS3 positive cells were hepatocytes. D. Graph representing mean±SEM of P-STAT3 positive hepatocytes per HPF, as analyzed by E. immunohistochemistry (brown). Double immunofluorescence staining, using fluorescence labeled P-STAT3 and hepatocyte specific hepatocyte nuclear factor 4 α (HNF) (P-STAT3: red; HNF: green; DAPI: blue). Yellow arrows indicate P-STAT3/HNF/DAPI positive hepatocytes; pink arrows indicate P-STAT3/DAPI positive but HNF negative cells. Photomicrographs are representative of 3–4 mice per group. Original magnification X200 (light microscopy) and X400 (fluorescence microscopy). F. Histograms representing mean ± SEM of relative miR-203 mRNA levels measured by qPCR after normalization with the house keeping miRNA gene sno202 (n= 5–7 mice per group). *p<0.05 **p< 0.01 and ***p< 0.001.
A20 Overexpression in Mouse Livers Enhances and Accelerates Hepatocyte Proliferation Following Extended Hepatectomy by Inhibiting SOCS3 Upregulation
We chose to confirm the effect of A20 on IL-6/STAT3/SOCS3 in the mouse model of EH, characterized by delayed LR and high lethality rate (50%), as to magnify A20’s pro-proliferative mechanism(s) (15). We performed EH in BALB/C mice that did not receive any rAd. (C) or were intravenously injected with rAd.A20 or the control rAd.βgal. Overexpression of A20 inhibited EH-induced hepatic upregulation of SOCS3 mRNA, when compared to C or rAd.βgal treated livers, 24h following surgery (Fig. 5A; n=3–6). We confirmed these results by showing decreased SOCS3 immunostaining in A20 overexpressing livers, 36h following EH (Fig. 5B; n=5–6).
Fig. 5. Overexpression of A20 in livers down-regulates SOCS3 to promote hepatocyte proliferation following extended hepatectomy.
Livers from mice that did not receive recombinant adenovirus (C) or were injected with rAd.A20 or rAd.βgal were recovered before (pre) and 24h (mRNA) or 36h (immmunostaining) after extended hepatectomy (post) and evaluated for A. SOCS3 mRNA levels by qPCR. Histograms represent mean ± SEM relative SOCS3 mRNA levels (n= 3–6 mice per group). B. SOCS3 protein expression by immunohistochemistry (brown) (n=5–6 mice per group). C. Cyclin D1 (CCND1) and D. Cyclin A (CCNA) mRNA levels by qPCR. Histograms represent mean ± SEM relative CCND1 and CCNA mRNA levels (n= 3–6 mice per group). E. Hepatocyte proliferation was evaluated by Ki67 immunostaining (brown). Double immunofluorescence, using fluorescence labeled Ki67 and hepatocyte specific HNF (Ki67: red; HNF: green; DAPI: blue), confirm that ki67 positive cells are hepatocytes. Histograms represents mean±SE of Ki67 positive hepatocytes per HPF. Photomicrographs in B and E are representative of 5–6 (light microscopy) or 3–6 (fluorescence microscopy) mice per group. Original magnification ×200 (light microscopy) and ×400 (fluorescence microscopy). mRNA levels in A, C, D were normalized using TATA box binding protein (TBP) mRNA. *p<0.05, **p<0.01, ***p<0.001
EH resulted in a moderate increase in Cyclin D1 (CCND1) and Cyclin A (CCNA) mRNA in C livers. This is expected given impaired LR in this EH model. On the other hand, rAd-treated livers had higher basal mRNA levels of CCND1 and CCNA, when compared to C, possibly reflecting adenoviral toxicity. Whereas rAd. toxicity repressed further EH-induced upregulation of CCND1 and CCNA mRNA in rAd.βgal treated livers, overexpression of A20 rescued this outcome, allowing for a significant increase of both cyclins, 24h following EH (Fig. 5C & D; n=3–6). We ascribe this positive outcome in A20 treated livers to lower SOCS3 mRNA levels, hence enhanced IL-6/STAT3 signals, culminating in increased mRNA levels of downstream targets, CCND1 and CCNA. Consequently, hepatocyte proliferation was significantly enhanced, as shown by increased number of Ki67/HNF4α positive hepatocytes, in rAd.A20 vs C and rAd.βgal treated livers (Fig. 5E; n=5–6; p<0.001). Interestingly, greater hepatocyte proliferation in rAd.A20 vs. rAd.βgal livers occurred despite significantly higher basal hepatic IL-6 levels in the latter (Fig. S5; n=4–5; p<0.001). Higher IL-6 in rAd.βgal treated livers is caused by the inflammatory response to rAd. that is contained in rAd.A20 treated livers by the anti-inflammatory function of this protein.
Increased SOCS3 Expression in A20 Heterozygous Livers Correlates with Impaired Hepatocyte Proliferation During Liver Regeneration
We evaluated SOCS3 expression in livers of A20 HT and WT mice before and 24h following EH. A20 KO mice die within 3–6 weeks of age, and hence cannot be used for these experiments (21). HT livers showed a tendency towards higher SOCS3 mRNA and protein (IHC) (Fig 6A; n=5–6) and lower miR203 levels than WT, 24h after EH (Fig. 6B; n=5–6). Increased SOCS3 levels in HT livers associated with impaired hepatocyte proliferation and delayed LR, as evidenced by lower number of Ki67/HNF4α positive hepatocytes in HT vs. WT livers, 36h after EH (Fig. 6C; n=2 (WT) and 5 (HT); p<0.001). This also correlated with lower P-STAT3 levels in HT vs. WT livers, 4h following EH (Fig. 6D; n=2), and with inadequate upregulation of CCND1 (p<0.05) and CCNA (p<0.05) mRNA, 24h after EH (Fig. 6E and 6F; n=5–6).
Fig. 6. A20 knockdown increases SOCS3 expression, impairing hetapocyte proliferation following extended hepatectomy.
Mice livers from A20 WT and A20 HT mice were recovered and evaluated for A. SOCS3 mRNA levels by qPCR and protein expression by immunohistochemistry (brown), before (pre) and 24h after (post) EH. Histograms represent mean ± SEM relative SOCS3 mRNA levels (n= 5–6 mice per group). Photomicrographs are representative of 5–6 mice per group. B. miR-203 mRNA levels by qPCR, pre and 24h post EH. Hitograms represent mean ± SEM relative miR203 mRNA levels after normalization with the house keeping miRNA gene sno202 (n= 5–6 mice per group). C. Hepatocyte proliferation by Ki67 immunostaining (brown) and double immunofluorescence (Ki67: red; hepatocyte nuclear factor 4α (HNF): green; DAPI: blue), pre and 36h post EH. Histograms represent mean ± SEM of Ki67 positive cells per high power field (HPF) (n=2 (WT) and 5 (HT) mice). D. phospho (P-STAT3) and total STAT-3 by Western blot analysis, pre and 4h post EH. Immunoblots for GAPDH were used as loading control. Histograms represent mean ± SEM of corrected densitometry results, calculated as percentage of P-STAT3/STAT3 ratio in pre WT livers. E. Cyclin D1 (CCND1) and F. Cyclin A (CCNA) mRNA levels, by qPCR pre and 24h post EH. Histograms represent mean ± SEM of relative CCND1 and CCNA mRNA levels (n= 5–6 mice per group). Original magnification in A and C were ×200 (light microscopy) and ×400 (fluorescence microscopy). In A, E, and F mRNA levels were normalized using TATAbox binding protein (TBP) mRNA.*p<0.05, ***p<0.001.
A20 Overexpression Attenuates while A20 Knockdown Enhances Expression of STAT3-dependent Acute Phase Response Genes Following Liver Resection
IL-6/STAT3 signaling also triggers an acute inflammatory response in hepatocytes that, if uncontrolled, causes liver damage and counterproductively stuns cell cycle progression (9). Accordingly, we measured mRNA levels of STAT3-dependent liver-derived acute phase response proteins serum amyloid A 1 (SAA1) and fibrinogen (FGG) in A20-overexpressing and knockdown livers, at baseline and following EH. Overexpression of A20 significantly decreased EH-induced upregulation of FGG (Fig. 7A and 7B; n=6; p< 0.01), and reduced baseline and EH-induced upregulation of SAA1. Conversely, A20 HT livers expressed higher levels of SAA1 (Fig 7C; n=5–6; p<0.01), and FGG (Fig. 7D; n=6) following EH. These findings suggest that A20 does not aggravate and may even moderately decrease IL-6-induced pro-inflammatory signals.
Fig. 7. A20 expression regulates acute phase response gene expression following liver resection.

Serum Amyloid A 1 (SAA1) and fibrinogen gamma chain (FGG) mRNA levels were evaluated by qPCR in livers from: A and B. rAd.A20 and rAd.βgal transduced livers, and C and D WT and A20 HT livers before (pre) and 24h after (post) EH (n=5–6 mice per group). Histograms represent mean ± SEM of relative target gene mRNA levels after normalization with TATA box binding protein (TBP). **p<0.01.
Discussion
A20, a newly identified player in LR, is induced in mice and humans as part of the hepatocyte’s regenerative response following hepatectomy (15, 20). Overexpression of A20 dramatically improves LR and survival in mouse models of extended (78%) and lethal (87%) hepatectomy in part through decreasing the expression of CDKI, p21 (15). In contrast, A20 knockdown significantly impairs and delays LR (Studer et al, in preparation). We initiated this work to determine which A20 domain supports this function in hepatocytes. Our results showed that both Nter and 7Zn mutants independently promote hepatocyte proliferation, but neither recapitulates A20’s effect on p21. Akin to A20’s NF-κB inhibitory function, both domains were required to decrease p21. Inhibition of NF-κB by A20 has been ascribed to its sequential N-terminus mediated deubiquitination of signaling mediators such as RIP and NEMO, followed by C-terminus-mediated degradative polyubiquitination of the same substrates (17). Whether ubiquitin editing or other cooperative mechanisms account for A20 decreasing p21 mRNA is being explored. Regardless, these results suggested that mechanisms other than those involving p21 account for the pro-proliferative function of A20 in hepatocytes. In this work, we identified one of these mechanisms by demonstrating that A20 decreases expression of the negative regulator of IL-6 signaling, SOCS3.
IL-6, produced following hepatectomy by Kupffer cells and hepatocytes (25), is the central trigger of LR via phosphorylation/activation of STAT3 (27). Successful LR depends on an intact TNF-NFκB-IL-6-STAT3 pathway. TNF-R1 KO, and hepatocyte specific IL-6 or STAT3 KO mice have impaired regeneration, causing sometimes lethality, following PH (5, 27–29). IL-6 administration decreases lethality rates post PH in TNF-R1 KO mice, indicating TNF promotes LR mostly by inducing IL-6 (28). The impact of A20 on IL-6/STAT3 signaling was unknown. We surmised that overexpression of A20, by blocking LPS/TNF-mediated NF-κB activation in hepatocytes, could reduce IL-6 production hence limit its own pro-proliferative advantage. We confirmed that overexpression of A20 (but neither Nter nor 7Zn that do not inhibit NF-κB) significantly decreased LPS/TNF induced upregulation of IL-6 in HepG2, without eliminating it, which indicated that IL-6 expression was not exclusively NF-κB dependent in hepatocytes (22). However, despite lower IL-6 levels, there was stronger baseline, and LPS/TNF-induced phosphorylation of STAT3 in A20 overexpressing hepatocytes. This effect was mimicked by 7Zn mutant. In fact, mere overexpression of A20 or 7Zn in HepG2 significantly increased STAT3 phosphorylation at baseline, and these levels were moderately increased or unchanged by IL-6 treatment, indicating that exogenous IL-6 was not necessary to produce this effect. We believe that high and sustained STAT3 phosphorylation in A20/7Zn overexpressing hepatocytes is key to their pro-proliferative advantage, regardless of whether these cells are treated with IL-6. Loss of function experiments supported A20’s physiologic impact upon IL-6/STAT3 signaling, as they showed significantly higher TNF-induced IL-6 secretion, with paradoxically lower STAT3 phosphorylation in A20 KO and HT hepatocytes. We attribute this paradox to A20 knockdown increasing SOCS3 expression. STAT3 inducible SOCS3 is part of a negative feedback loop that inhibits IL-6 signaling, i.e. STAT3 phosphorylation (30). Gain of function studies confirmed that SOCS3 was the prime target of A20 in modulating IL-6 signaling. Overexpression of A20 or 7Zn in hepatocytes significantly decreased SOCS3 expression, thereby increasing STAT3 phosphorylation. Increased and sustained STAT3 phosphorylation in A20 overexpressing hepatocytes is similar to that seen in IL-6 treated hKO SOCS3 hepatocytes (11, 30). Furthermore, hKO SOCS3 livers, similar to A20 overexpressing livers, demonstrate enhanced and accelerated regeneration following PH (11, 30).
In vivo, basal SOCS3 mRNA and protein levels were also significantly enhanced in A20 KO vs. WT livers, with A20 HT showing an intermediate phenotype. Basal immunostaining for P-STAT3 was higher in A20 KO vs. WT livers. We believe this result represents enhanced inflammation, as indicated by significantly higher IL-6 levels in A20 KO livers causing increased, but still inadequate, STAT3 phosphorylation. This is consistent with impaired hepatocyte proliferation following PH in mice either chronically exposed to high IL-6 levels (like A20 KO), or overexpressing the soluble IL-6-receptor gp80 and concomitantly treated with IL-6 (9). Impaired proliferation in these conditions results, at least in part, from IL-6 dependent upregulation of p21 (9, 31), as in A20 KO livers.
IL-6 levels were increased in A20 HT livers at baseline, yet these livers still showed a trend towards higher SOCS3 levels. We discovered that A20 knockdown (KO and HT) significantly decreased hepatic levels of miR-203. Since SOCS3 is an evolutionarily conserved target of miR-203 (26), A20-mediated modulation of SOCS3 expression in hepatocytes is, at least in part, epigenetically regulated by A20’s effect on miR-203.
We validated these findings in mouse models of EH. A20 overexpression significantly decreased SOCS3 mRNA and protein levels in mice livers following EH, while A20 knockdown had the opposite effect. Accordingly, STAT3 dependent CCNA and CCND1 levels increased in A20 overexpressing livers, enhancing hepatocyte proliferation following EH. These results are consistent with increased expression of Cyclins (D, E, A) and improved LR in SOCS3 heterozygous and hKO SOCS3 mice after PH (11, 31). In contrast, A20 HT livers failed to adequately upregulate CCND1 and CCNA, hence showed decreased hepatocyte proliferation following EH. We plan to overexpress A20 in IL-6 and SOCS3 KO mice undergoing EH in order to evaluate the contribution of A20’s impact on the IL-6/STAT3/SOCS3 pathway to its overall pro-proliferative function in hepatocytes. We recognize that SOCS3 knockdown/ STAT3 activation are linked to hepatocarcinogenesis (11, 32), a potential concern for A20 overexpressing livers. Our previous studies however indicate that short-term overexpression of A20 does not carry a significant carcinogenic risk. Indeed, no rAd.A20 treated mice developed liver carcinomas during the 6 months monitoring period (14–16). Longer follow-up periods may be required to completely rule out this risk.
In contrast to cell cycle targets of STAT3, overexpression of A20 slightly decreased and A20 knockdown increased STAT3-induced pro-inflammatory acute phase response genes, SAA1 and FGG, following EH (33). These data agree with NF-κB (inhibited by A20 overexpression) synergizing with STAT3 to induce acute phase response proteins (34), and with data demonstrating that increased SOCS3 (as in A20 KO) enhances NF-κB activation (35).
In summary, this is the first report demonstrating that A20 enhances IL-6 proliferative signals in hepatocytes through down-regulation of SOCS3 while decreasing IL-6 levels and reducing its pro-inflammatory signals, all of which optimize liver regeneration (Fig. 8). This key finding along with A20 reducing p21 levels underscore A20’s pro-proliferative properties in hepatocytes, and support pursuit of A20-based therapies to promote LR following extensive liver resections for living donation or large tumors.
Fig. 8. A20 promotes proliferative and reduces inflammatory IL-6/STAT3 signals.

In response to decreased liver mFfigass and relative increase in hepatic LPS concentration, activated kupffer cells secrete tumor necrosis factor alpha (TNF). When bound to TNF, the TNF receptor transduces mitogenic but also pro-inflammatory signals in neighboring hepatocytes, which are mediated by the transcription factor NF-κB. Activation of NF-κB occurs via phosphorylation of the NF-κB inhibitor IκBα, resulting in the dissociation and subsequent nuclear localization of phosphorylated NF-κB (p65/p50), initiating transcription of NF-κB dependent genes such as IL-6 and the NF-κB regulatory protein A20/tnfaip3. Upon IL-6 binding, the IL-6R/gp130 dimer induces phosphorylation of JAK1 and 3, which in turn phosphorylates STAT3. Phosphorylated STAT3 dimerizes and translocates to the nucleus, where it binds STAT3 binding element (SBE) and initiates transcription of cyclins, acute phase proteins, and the negative regulator of IL-6 signaling, socs3. A20 regulates IL-6/STAT3 signaling by increasing miR203 levels, which downregulates SOCS3 mRNA thus unleashes STAT3 signaling and transcription of STAT3-dependent mitogenic genes such as cyclin d1 and cyclin a. On the other hand, A20 blocks NF-κB signaling, thus reducing transcription of STAT3-dependent proinflammatory genes, such as saa1 and fgg, whose transcription relies on synergy between STAT3 and NF-κB. In sum overexpression of A20 in hepatocytes promotes their proliferation by favoring IL-6 proliferative over inflammatory signals.
Supplementary Material
Acknowledgments
This study was supported NIH grant RO1s DK063275 and HL080130, and R21 grant DK091822 to CF. CRP, and VIP were supported by NIH T32 HL07734. PS was supported by a grant from the National Swiss Foundation. DM was supported by a fellowship from the Centro Nacional de Desenvolvimento Cientifíco e Tecnológico CNPq.
We thank Dr. Vishva Dixit and Robert Gerard for providing the A20 plasmid and the recombinant β-galactosidase adenovirus. We also acknowledge the help of Mr. Alon Neidich in editing this manuscript.
List of Abbreviations
- 7Zn
A20’s 7 zinc C-terminal domain
- ANOVA
analysis of variance
- βgal
βgalactosidase
- CCNA
Cyclin A
- CCND1
Cyclin D1
- CDKI
Cyclin Dependent Kinase Inhibitor
- EDTA
ethylenediaminetetraacetic acid
- EH
extended hepatectomy
- ELISA
enzyme-linked immunosorbent assay
- FGG
fibrinogen gamma chain
- FBS
Fetal bovine serum
- GAPDH
glyceraldehyde 3-phosphate dehydrogenase
- HA
hemagglutinin
- HT
heterozygous
- IHC
immunohistochemistry
- IL-6
interleukin-6
- LR
liver regeneration
- KO
knockout
- LPS
lipopolysaccharide
- miR
microRNA
- MPH
mouse primary hepatocytes
- NF-κB
Nuclear Factor Kappa B
- Nter
A20’s N-terminal domain
- PCNA
proliferating cell nuclear antigen
- PH
partial hepatectomy
- qPCR
quantitative real time polymerase chain reaction
- rAd
recombinant adenovirus
- SOCS3
suppressor of cytokine signaling 3
- SAA1
serum amyloid A 1
- STAT3
signal transducer and activator of transcription 3
- TBP
TATA box binding protein
- TNF
tumor necrosis factor alpha
- WT
wild type
References
- 1.Fausto N, Campbell JS, Riehle KJ. Liver regeneration. Hepatology. 2006;43:S45–53. doi: 10.1002/hep.20969. [DOI] [PubMed] [Google Scholar]
- 2.Michalopoulos GK. Liver regeneration. J Cell Physiol. 2007;213:286–300. doi: 10.1002/jcp.21172. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Cressman DE, Diamond RH, Taub R. Rapid activation of the Stat3 transcription complex in liver regeneration. Hepatology. 1995;21:1443–1449. [PubMed] [Google Scholar]
- 4.Galun E, Zeira E, Pappo O, Peters M, Rose-John S. Liver regeneration induced by a designer human IL-6/sIL-6R protein reverses severe hepatocellular injury. FASEB J. 2000;14:1979–1987. doi: 10.1096/fj.99-0913com. [DOI] [PubMed] [Google Scholar]
- 5.Cressman DE, Greenbaum LE, DeAngelis RA, Ciliberto G, Furth EE, Poli V, Taub R. Liver failure and defective hepatocyte regeneration in interleukin-6-deficient mice. Science. 1996;274:1379–1383. doi: 10.1126/science.274.5291.1379. [DOI] [PubMed] [Google Scholar]
- 6.Blindenbacher A, Wang X, Langer I, Savino R, Terracciano L, Heim MH. Interleukin 6 is important for survival after partial hepatectomy in mice. Hepatology. 2003;38:674–682. doi: 10.1053/jhep.2003.50378. [DOI] [PubMed] [Google Scholar]
- 7.Ren X, Hogaboam C, Carpenter A, Colletti L. Stem cell factor restores hepatocyte proliferation in IL-6 knockout mice following 70% hepatectomy. J Clin Invest. 2003;112:1407–1418. doi: 10.1172/JCI17391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Huda KA, Guo L, Haga S, Murata H, Ogino T, Fukai M, Yagi T, et al. Ex vivo adenoviral gene transfer of constitutively activated STAT3 reduces post-transplant liver injury and promotes regeneration in a 20% rat partial liver transplant model. Transpl Int. 2006;19:415–423. doi: 10.1111/j.1432-2277.2006.00285.x. [DOI] [PubMed] [Google Scholar]
- 9.Wustefeld T, Rakemann T, Kubicka S, Manns MP, Trautwein C. Hyperstimulation with interleukin 6 inhibits cell cycle progression after hepatectomy in mice. Hepatology. 2000;32:514–522. doi: 10.1053/jhep.2000.16604. [DOI] [PubMed] [Google Scholar]
- 10.Campbell JS, Prichard L, Schaper F, Schmitz J, Stephenson-Famy A, Rosenfeld ME, Argast GM, et al. Expression of suppressors of cytokine signaling during liver regeneration. J Clin Invest. 2001;107:1285–1292. doi: 10.1172/JCI11867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Riehle KJ, Campbell JS, McMahan RS, Johnson MM, Beyer RP, Bammler TK, Fausto N. Regulation of liver regeneration and hepatocarcinogenesis by suppressor of cytokine signaling 3. J Exp Med. 2008;205:91–103. doi: 10.1084/jem.20070820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Cooper JT, Stroka DM, Brostjan C, Palmetshofer A, Bach FH, Ferran C. A20 blocks endothelial cell activation through a NF-kB-dependent mechanism. J Biol Chem. 1996;271:18068–18073. doi: 10.1074/jbc.271.30.18068. [DOI] [PubMed] [Google Scholar]
- 13.Bach FH, Hancock WW, Ferran C. Protective genes expressed in endothelial cells: a regulatory response to injury. Immunol Today. 1997;18:483–486. doi: 10.1016/s0167-5699(97)01129-8. [DOI] [PubMed] [Google Scholar]
- 14.Arvelo MB, Cooper JT, Longo C, Daniel S, Grey ST, Mahiou J, Czismadia E, et al. A20 protects mice from D-galactosamine/lipopolysaccharide acute toxic lethal hepatitis. Hepatology. 2002;35:535–543. doi: 10.1053/jhep.2002.31309. [DOI] [PubMed] [Google Scholar]
- 15.Longo CR, Patel VI, Shrikhande GV, Scali ST, Csizmadia E, Daniel S, Sun DW, et al. A20 protects mice from lethal radical hepatectomy by promoting hepatocyte proliferation via a p21waf1-dependent mechanism. Hepatology. 2005;42:156–164. doi: 10.1002/hep.20741. [DOI] [PubMed] [Google Scholar]
- 16.Ramsey HE, Da Silva CG, Longo CR, Csizmadia E, Studer P, Patel VI, Damrauer SM, et al. A20 protects mice from lethal liver ischemia/reperfusion injury by increasing peroxisome proliferator-activated receptor-alpha expression. Liver Transpl. 2009;15:1613–1621. doi: 10.1002/lt.21879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wertz IE, O’Rourke KM, Zhou H, Eby M, Aravind L, Seshagiri S, Wu P, et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-kappaB signalling. Nature. 2004;430:694–699. doi: 10.1038/nature02794. [DOI] [PubMed] [Google Scholar]
- 18.Daniel S, Arvelo MB, Patel VI, Longo CR, Shrikhande G, Shukri T, Mahiou J, et al. A20 protects endothelial cells from TNF-, Fas-, and NK-mediated cell death by inhibiting caspase 8 activation. Blood. 2004;104:2376–2384. doi: 10.1182/blood-2003-02-0635. [DOI] [PubMed] [Google Scholar]
- 19.Damrauer SM, Studer P, da Silva CG, Longo CR, Ramsey HE, Csizmadia E, Shrikhande GV, et al. A20 modulates lipid metabolism and energy production to promote liver regeneration. PLoS ONE. 2011;6:e17715. doi: 10.1371/journal.pone.0017715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.de Jonge J, Kurian S, Shaked A, Reddy KR, Hancock W, Salomon DR, Olthoff KM. Unique early gene expression patterns in human adult-to-adult living donor liver grafts compared to deceased donor grafts. Am J Transplant. 2009;9:758–772. doi: 10.1111/j.1600-6143.2009.02557.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lee EG, Boone DL, Chai S, Libby SL, Chien M, Lodolce JP, Ma A. Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science. 2000;289:2350–2354. doi: 10.1126/science.289.5488.2350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Matsusaka T, Fujikawa K, Nishio Y, Mukaida N, Matsushima K, Kishimoto T, Akira S. Transcription factors NF-IL6 and NF-kappa B synergistically activate transcription of the inflammatory cytokines, interleukin 6 and interleukin 8. Proceedings of the National Academy of Sciences of the United States of America. 1993;90:10193–10197. doi: 10.1073/pnas.90.21.10193. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Harman AW, McCamish LE, Henry CA. Isolation of hepatocytes from postnatal mice. J Pharmacol Methods. 1987;17:157–163. doi: 10.1016/0160-5402(87)90026-x. [DOI] [PubMed] [Google Scholar]
- 24.Ferran C, Stroka DM, Badrichani AZ, Cooper JT, Wrighton CJ, Soares M, Grey ST, et al. A20 inhibits NF-kappaB activation in endothelial cells without sensitizing to tumor necrosis factor-mediated apoptosis. Blood. 1998;91:2249–2258. [PubMed] [Google Scholar]
- 25.Panesar N, Tolman K, Mazuski JE. Endotoxin stimulates hepatocyte interleukin-6 production. J Surg Res. 1999;85:251–258. doi: 10.1006/jsre.1999.5648. [DOI] [PubMed] [Google Scholar]
- 26.Wei T, Orfanidis K, Xu N, Janson P, Stahle M, Pivarcsi A, Sonkoly E. The expression of microRNA-203 during human skin morphogenesis. Exp Dermatol. 2010;19:854–856. doi: 10.1111/j.1600-0625.2010.01118.x. [DOI] [PubMed] [Google Scholar]
- 27.Li W, Liang X, Kellendonk C, Poli V, Taub R. STAT3 contributes to the mitogenic response of hepatocytes during liver regeneration. J Biol Chem. 2002;277:28411–28417. doi: 10.1074/jbc.M202807200. [DOI] [PubMed] [Google Scholar]
- 28.Yamada Y, Kirilova I, Peschon JJ, Fausto N. Initiation of liver growth by tumor necrosis factor: deficient liver regeneration in mice lacking type I tumor necrosis factor receptor. Proc Natl Acad Sci USA. 1997;94:1441–1446. doi: 10.1073/pnas.94.4.1441. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Moh A, Iwamoto Y, Chai GX, Zhang SS, Kano A, Yang DD, Zhang W, et al. Role of STAT3 in liver regeneration: survival, DNA synthesis, inflammatory reaction and liver mass recovery. Lab Invest. 2007;87:1018–1028. doi: 10.1038/labinvest.3700630. [DOI] [PubMed] [Google Scholar]
- 30.Croker BA, Krebs DL, Zhang JG, Wormald S, Willson TA, Stanley EG, Robb L, et al. SOCS3 negatively regulates IL-6 signaling in vivo. Nat Immunol. 2003;4:540–545. doi: 10.1038/ni931. [DOI] [PubMed] [Google Scholar]
- 31.Sun R, Jaruga B, Kulkarni S, Sun H, Gao B. IL-6 modulates hepatocyte proliferation via induction of HGF/p21cip1: regulation by SOCS3. Biochem Biophys Res Commun. 2005;338:1943–1949. doi: 10.1016/j.bbrc.2005.10.171. [DOI] [PubMed] [Google Scholar]
- 32.Ogata H, Kobayashi T, Chinen T, Takaki H, Sanada T, Minoda Y, Koga K, et al. Deletion of the SOCS3 gene in liver parenchymal cells promotes hepatitis-induced hepatocarcinogenesis. Gastroenterology. 2006;131:179–193. doi: 10.1053/j.gastro.2006.04.025. [DOI] [PubMed] [Google Scholar]
- 33.Leu JI, Crissey MA, Leu JP, Ciliberto G, Taub R. Interleukin-6-induced STAT3 and AP-1 amplify hepatocyte nuclear factor 1-mediated transactivation of hepatic genes, an adaptive response to liver injury. Mol Cell Biol. 2001;21:414–424. doi: 10.1128/MCB.21.2.414-424.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Hagihara K, Nishikawa T, Sugamata Y, Song J, Isobe T, Taga T, Yoshizaki K. Essential role of STAT3 in cytokine-driven NF-kappaB-mediated serum amyloid A gene expression. Genes Cells. 2005;10:1051–1063. doi: 10.1111/j.1365-2443.2005.00900.x. [DOI] [PubMed] [Google Scholar]
- 35.Park SH, Kim KE, Hwang HY, Kim TY. Regulatory effect of SOCS on NF-kappaB activity in murine monocytes/macrophages. DNA Cell Biol. 2003;22:131–139. doi: 10.1089/104454903321515931. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.