Abstract
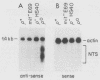
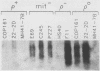
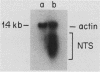
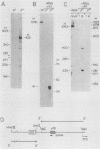
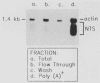
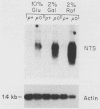
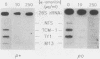
We have identified stable transcripts from the so-called nontranscribed spacer region (NTS) of the nuclear ribosomal DNA repeat in certain respiration-deficient strains of Saccharomyces cerevisiae. These RNAs, which are transcribed from the same strand as is the 37S rRNA precursor, are 500 to 800 nucleotides long and extend from the 5' end of the 5S rRNA gene to three major termination sites about 1,780, 1,830, and 1,870 nucleotides from the 3' end of the 26S rRNA gene. A survey of various wild-type and respiration-deficient strains showed that NTS transcript abundance depended on the mitochondrial genotype and a single codominant nuclear locus. In strains with that nuclear determinant, NTS transcripts were barely detected in [rho+] cells, were slightly more abundant in various mit- derivatives, and were most abundant in petites. However, in one petite that was hypersuppressive and contained a putative origin of replication (ori5) within its 757-base-pair mitochondrial genome, NTS transcripts were no more abundant than in [rho+] cells. The property of low NTS transcript abundance in the hypersuppressive petite was unstable, and spontaneous segregants that contained NTS transcripts as abundant as in the other petites examined could be obtained. Thus, respiration deficiency per se is not the major factor contributing to the accumulation of these unusual RNAs. Unlike RNA polymerase I transcripts, the abundant NTS RNAs were glucose repressible, fractionated as poly(A)+ RNAs, and were sensitive to inhibition by 10 micrograms of alpha-amanitin per ml, a concentration that had no effect on rRNA synthesis. Abundant NTS RNAs are therefore most likely derived by polymerase II transcription.
Full text
PDF


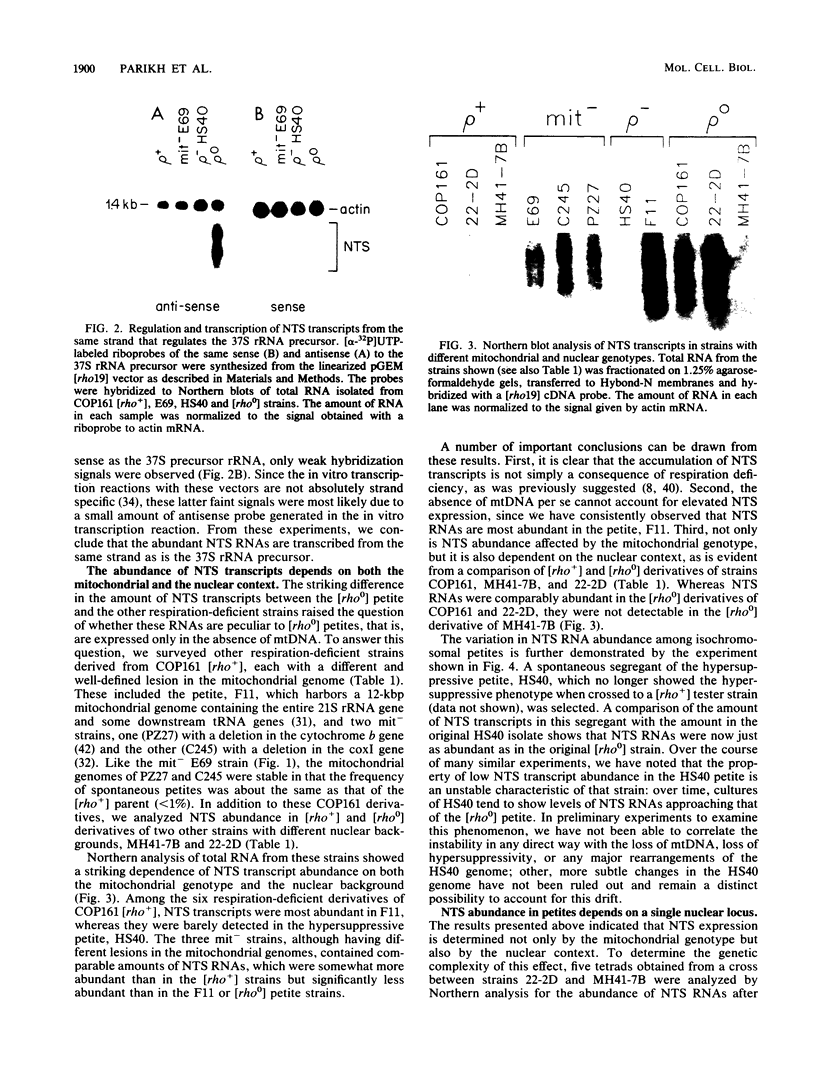
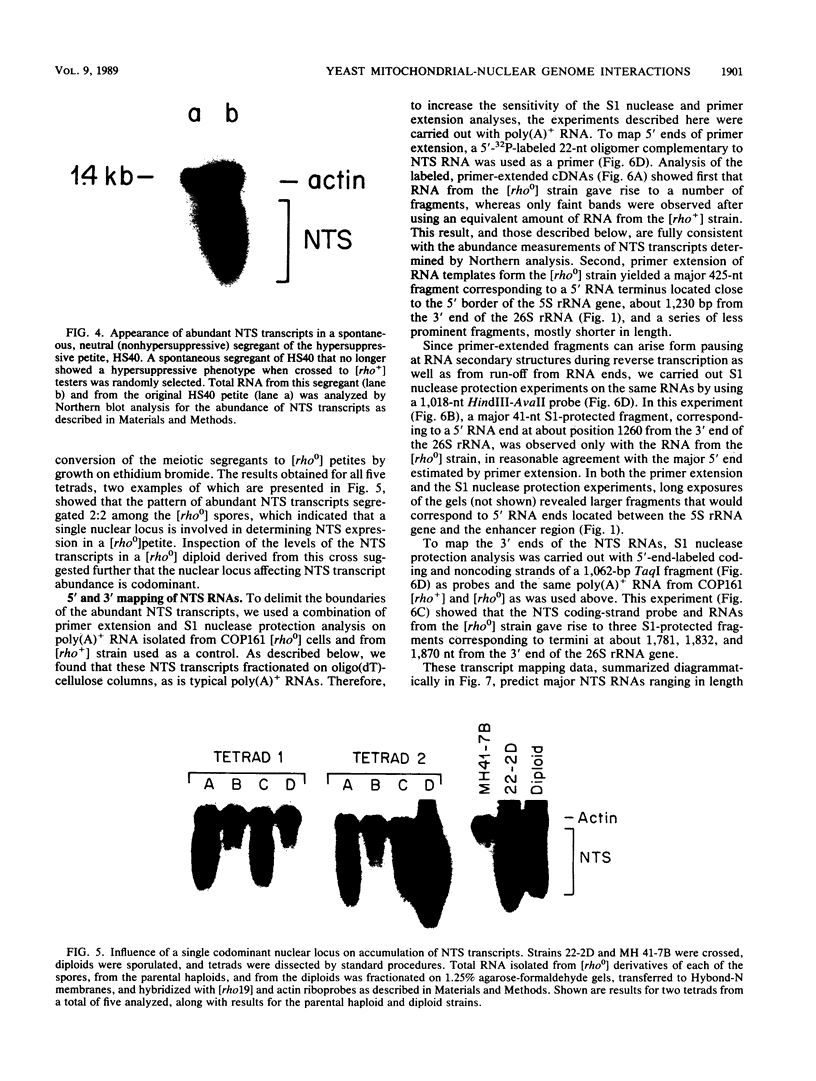






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aviv H., Leder P. Purification of biologically active globin messenger RNA by chromatography on oligothymidylic acid-cellulose. Proc Natl Acad Sci U S A. 1972 Jun;69(6):1408–1412. doi: 10.1073/pnas.69.6.1408. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barath Z., Küntzel H. Cooperation of mitochondrial and nuclear genes specifying the mitochondrial genetic apparatus in Neurospora crassa. Proc Natl Acad Sci U S A. 1972 Jun;69(6):1371–1374. doi: 10.1073/pnas.69.6.1371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bayev A. A., Georgiev O. I., Hadjiolov A. A., Kermekchiev M. B., Nikolaev N., Skryabin K. G., Zakharyev V. M. The structure of the yeast ribosomal RNA genes. 2. The nucleotide sequence of the initiation site for ribosomal RNA transcription. Nucleic Acids Res. 1980 Nov 11;8(21):4919–4926. doi: 10.1093/nar/8.21.4919. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blanc H., Dujon B. Replicator regions of the yeast mitochondrial DNA responsible for suppressiveness. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3942–3946. doi: 10.1073/pnas.77.7.3942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boeke J. D., LaCroute F., Fink G. R. A positive selection for mutants lacking orotidine-5'-phosphate decarboxylase activity in yeast: 5-fluoro-orotic acid resistance. Mol Gen Genet. 1984;197(2):345–346. doi: 10.1007/BF00330984. [DOI] [PubMed] [Google Scholar]
- Butow R. A., Docherty R., Parikh V. S. A path from mitochondria to the yeast nucleus. Philos Trans R Soc Lond B Biol Sci. 1988 May 31;319(1193):127–133. doi: 10.1098/rstb.1988.0037. [DOI] [PubMed] [Google Scholar]
- Cassidy B. G., Yang-Yen H. F., Rothblum L. I. Transcriptional role for the nontranscribed spacer of rat ribosomal DNA. Mol Cell Biol. 1986 Aug;6(8):2766–2773. doi: 10.1128/mcb.6.8.2766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clare J., Farabaugh P. Nucleotide sequence of a yeast Ty element: evidence for an unusual mechanism of gene expression. Proc Natl Acad Sci U S A. 1985 May;82(9):2829–2833. doi: 10.1073/pnas.82.9.2829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Costanzo M. C., Fox T. D. Specific translational activation by nuclear gene products occurs in the 5' untranslated leader of a yeast mitochondrial mRNA. Proc Natl Acad Sci U S A. 1988 Apr;85(8):2677–2681. doi: 10.1073/pnas.85.8.2677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dieckmann C. L., Homison G., Tzagoloff A. Assembly of the mitochondrial membrane system. Nucleotide sequence of a yeast nuclear gene (CBP1) involved in 5' end processing of cytochrome b pre-mRNA. J Biol Chem. 1984 Apr 25;259(8):4732–4738. [PubMed] [Google Scholar]
- Dieckmann C. L., Koerner T. J., Tzagoloff A. Assembly of the mitochondrial membrane system. CBP1, a yeast nuclear gene involved in 5' end processing of cytochrome b pre-mRNA. J Biol Chem. 1984 Apr 25;259(8):4722–4731. [PubMed] [Google Scholar]
- Dieckmann C. L., Mittelmeier T. M. Nuclearly-encoded CBP1 interacts with the 5' end of mitochondrial cytochrome b pre-mRNA. Curr Genet. 1987;12(6):391–397. doi: 10.1007/BF00434815. [DOI] [PubMed] [Google Scholar]
- Elion E. A., Warner J. R. An RNA polymerase I enhancer in Saccharomyces cerevisiae. Mol Cell Biol. 1986 Jun;6(6):2089–2097. doi: 10.1128/mcb.6.6.2089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elion E. A., Warner J. R. The major promoter element of rRNA transcription in yeast lies 2 kb upstream. Cell. 1984 Dec;39(3 Pt 2):663–673. doi: 10.1016/0092-8674(84)90473-2. [DOI] [PubMed] [Google Scholar]
- Fitzgerald M., Shenk T. The sequence 5'-AAUAAA-3'forms parts of the recognition site for polyadenylation of late SV40 mRNAs. Cell. 1981 Apr;24(1):251–260. doi: 10.1016/0092-8674(81)90521-3. [DOI] [PubMed] [Google Scholar]
- Fried H. M., Warner J. R. Cloning of yeast gene for trichodermin resistance and ribosomal protein L3. Proc Natl Acad Sci U S A. 1981 Jan;78(1):238–242. doi: 10.1073/pnas.78.1.238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grummt I., Kuhn A., Bartsch I., Rosenbauer H. A transcription terminator located upstream of the mouse rDNA initiation site affects rRNA synthesis. Cell. 1986 Dec 26;47(6):901–911. doi: 10.1016/0092-8674(86)90805-6. [DOI] [PubMed] [Google Scholar]
- Guarente L., Mason T. Heme regulates transcription of the CYC1 gene of S. cerevisiae via an upstream activation site. Cell. 1983 Apr;32(4):1279–1286. doi: 10.1016/0092-8674(83)90309-4. [DOI] [PubMed] [Google Scholar]
- Hahn S., Pinkham J., Wei R., Miller R., Guarente L. The HAP3 regulatory locus of Saccharomyces cerevisiae encodes divergent overlapping transcripts. Mol Cell Biol. 1988 Feb;8(2):655–663. doi: 10.1128/mcb.8.2.655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington C. A., Chikaraishi D. M. Transcription of spacer sequences flanking the rat 45S ribosomal DNA gene. Mol Cell Biol. 1987 Jan;7(1):314–325. doi: 10.1128/mcb.7.1.314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayward D. C., Glover D. M. Analysis of the Drosophila rDNA promoter by transient expression. Nucleic Acids Res. 1988 May 25;16(10):4253–4268. doi: 10.1093/nar/16.10.4253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Henderson S., Sollner-Webb B. A transcriptional terminator is a novel element of the promoter of the mouse ribosomal RNA gene. Cell. 1986 Dec 26;47(6):891–900. doi: 10.1016/0092-8674(86)90804-4. [DOI] [PubMed] [Google Scholar]
- Kaneko T., Watanabe T., Oishi M. Effect of mitochondrial protein synthesis inhibitors on erythroid differentiation of mouse erythroleukemia (Friend) cells. Mol Cell Biol. 1988 Aug;8(8):3311–3315. doi: 10.1128/mcb.8.8.3311. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kempers-Veenstra A. E., Oliemans J., Offenberg H., Dekker A. F., Piper P. W., Planta R. J., Klootwijk J. 3'-End formation of transcripts from the yeast rRNA operon. EMBO J. 1986 Oct;5(10):2703–2710. doi: 10.1002/j.1460-2075.1986.tb04554.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kempers-Veenstra A. E., van Heerikhuizen H., Musters W., Klootwijk J., Planta R. J. Transcription of an artificial ribosomal RNA gene in yeast. EMBO J. 1984 Jun;3(6):1377–1382. doi: 10.1002/j.1460-2075.1984.tb01980.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kief D. R., Warner J. R. Coordinate control of syntheses of ribosomal ribonucleic acid and ribosomal proteins during nutritional shift-up in Saccharomyces cerevisiae. Mol Cell Biol. 1981 Nov;1(11):1007–1015. doi: 10.1128/mcb.1.11.1007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuiper M. T., Akins R. A., Holtrop M., de Vries H., Lambowitz A. M. Isolation and analysis of the Neurospora crassa Cyt-21 gene. A nuclear gene encoding a mitochondrial ribosomal protein. J Biol Chem. 1988 Feb 25;263(6):2840–2847. [PubMed] [Google Scholar]
- Locker J., Rabinowitz M. Transcription in yeast mitochondria: analysis of the 21 S rRNA region and its transcripts. Plasmid. 1981 Nov;6(3):302–314. doi: 10.1016/0147-619x(81)90038-x. [DOI] [PubMed] [Google Scholar]
- McStay B., Reeder R. H. A termination site for Xenopus RNA polymerase I also acts as an element of an adjacent promoter. Cell. 1986 Dec 26;47(6):913–920. doi: 10.1016/0092-8674(86)90806-8. [DOI] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss T. A transcriptional function for the repetitive ribosomal spacer in Xenopus laevis. Nature. 1983 Mar 17;302(5905):223–228. doi: 10.1038/302223a0. [DOI] [PubMed] [Google Scholar]
- Murtif V. L., Rae P. M. In vivo transcription of rDNA spacers in Drosophila. Nucleic Acids Res. 1985 May 10;13(9):3221–3239. doi: 10.1093/nar/13.9.3221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagley P., Linnane A. W. Expression of mitochondrial DNA in Saccharomyces cerevisiae: the construction of sets of isonuclear haploid strains containing different specified mitochondrial genomes. Biochem Biophys Res Commun. 1978 Nov 29;85(2):585–592. doi: 10.1016/0006-291x(78)91203-2. [DOI] [PubMed] [Google Scholar]
- Nasmyth K. Molecular analysis of a cell lineage. Nature. 1983 Apr 21;302(5910):670–676. doi: 10.1038/302670a0. [DOI] [PubMed] [Google Scholar]
- Ng R., Abelson J. Isolation and sequence of the gene for actin in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A. 1980 Jul;77(7):3912–3916. doi: 10.1073/pnas.77.7.3912. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parikh V. S., Morgan M. M., Scott R., Clements L. S., Butow R. A. The mitochondrial genotype can influence nuclear gene expression in yeast. Science. 1987 Jan 30;235(4788):576–580. doi: 10.1126/science.3027892. [DOI] [PubMed] [Google Scholar]
- Partaledis J. A., Mason T. L. Structure and regulation of a nuclear gene in Saccharomyces cerevisiae that specifies MRP13, a protein of the small subunit of the mitochondrial ribosome. Mol Cell Biol. 1988 Sep;8(9):3647–3660. doi: 10.1128/mcb.8.9.3647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Petes T. D. Yeast ribosomal DNA genes are located on chromosome XII. Proc Natl Acad Sci U S A. 1979 Jan;76(1):410–414. doi: 10.1073/pnas.76.1.410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfeifer K., Arcangioli B., Guarente L. Yeast HAP1 activator competes with the factor RC2 for binding to the upstream activation site UAS1 of the CYC1 gene. Cell. 1987 Apr 10;49(1):9–18. doi: 10.1016/0092-8674(87)90750-1. [DOI] [PubMed] [Google Scholar]
- Pfeifer K., Prezant T., Guarente L. Yeast HAP1 activator binds to two upstream activation sites of different sequence. Cell. 1987 Apr 10;49(1):19–27. doi: 10.1016/0092-8674(87)90751-3. [DOI] [PubMed] [Google Scholar]
- Pinkham J. L., Guarente L. Cloning and molecular analysis of the HAP2 locus: a global regulator of respiratory genes in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Dec;5(12):3410–3416. doi: 10.1128/mcb.5.12.3410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poutre C. G., Fox T. D. PET111, a Saccharomyces cerevisiae nuclear gene required for translation of the mitochondrial mRNA encoding cytochrome c oxidase subunit II. Genetics. 1987 Apr;115(4):637–647. doi: 10.1093/genetics/115.4.637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ruohola H., Baker S. M., Parker R., Platt T. Orientation-dependent function of a short CYC1 DNA fragment in directing mRNA 3' end formation in yeast. Proc Natl Acad Sci U S A. 1988 Jul;85(14):5041–5045. doi: 10.1073/pnas.85.14.5041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Santangelo G. M., Tornow J., McLaughlin C. S., Moldave K. Properties of promoters cloned randomly from the Saccharomyces cerevisiae genome. Mol Cell Biol. 1988 Oct;8(10):4217–4224. doi: 10.1128/mcb.8.10.4217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smale S. T., Tjian R. Transcription of herpes simplex virus tk sequences under the control of wild-type and mutant human RNA polymerase I promoters. Mol Cell Biol. 1985 Feb;5(2):352–362. doi: 10.1128/mcb.5.2.352. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sollner-Webb B., Tower J. Transcription of cloned eukaryotic ribosomal RNA genes. Annu Rev Biochem. 1986;55:801–830. doi: 10.1146/annurev.bi.55.070186.004101. [DOI] [PubMed] [Google Scholar]
- Strick C. A., Fox T. D. Saccharomyces cerevisiae positive regulatory gene PET111 encodes a mitochondrial protein that is translated from an mRNA with a long 5' leader. Mol Cell Biol. 1987 Aug;7(8):2728–2734. doi: 10.1128/mcb.7.8.2728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sutton A., Broach J. R. Signals for transcription initiation and termination in the Saccharomyces cerevisiae plasmid 2 micron circle. Mol Cell Biol. 1985 Oct;5(10):2770–2780. doi: 10.1128/mcb.5.10.2770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swanson M. E., Yip M., Holland M. J. Characterization of an RNA polymerase I-dependent promoter within the spacer region of yeast ribosomal cistrons. J Biol Chem. 1985 Aug 15;260(17):9905–9915. [PubMed] [Google Scholar]
- Szekely E., Montgomery D. L. Glucose represses transcription of Saccharomyces cerevisiae nuclear genes that encode mitochondrial components. Mol Cell Biol. 1984 May;4(5):939–946. doi: 10.1128/mcb.4.5.939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waldron C., Lacroute F. Effect of growth rate on the amounts of ribosomal and transfer ribonucleic acids in yeast. J Bacteriol. 1975 Jun;122(3):855–865. doi: 10.1128/jb.122.3.855-865.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winkler H., Adam G., Mattes E., Schanz M., Hartig A., Ruis H. Co-ordinate control of synthesis of mitochondrial and non-mitochondrial hemoproteins: a binding site for the HAP1 (CYP1) protein in the UAS region of the yeast catalase T gene (CTT1). EMBO J. 1988 Jun;7(6):1799–1804. doi: 10.1002/j.1460-2075.1988.tb03011.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]
- Zarkower D., Stephenson P., Sheets M., Wickens M. The AAUAAA sequence is required both for cleavage and for polyadenylation of simian virus 40 pre-mRNA in vitro. Mol Cell Biol. 1986 Jul;6(7):2317–2323. doi: 10.1128/mcb.6.7.2317. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zassenhaus H. P., Butow R. A., Hannon Y. P. Rapid electroelution of nucleic acids from agarose and acrylamide gels. Anal Biochem. 1982 Sep 1;125(1):125–130. doi: 10.1016/0003-2697(82)90392-x. [DOI] [PubMed] [Google Scholar]
- Zitomer R. S., Montgomery D. L., Nichols D. L., Hall B. D. Transcriptional regulation of the yeast cytochrome c gene. Proc Natl Acad Sci U S A. 1979 Aug;76(8):3627–3631. doi: 10.1073/pnas.76.8.3627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Zamaroczy M., Faugeron-Fonty G., Baldacci G., Goursot R., Bernardi G. The ori sequences of the mitochondrial genome of a wild-type yeast strain: number, location, orientation and structure. Gene. 1984 Dec;32(3):439–457. doi: 10.1016/0378-1119(84)90019-2. [DOI] [PubMed] [Google Scholar]