Abstract
U7 small nuclear RNA (snRNA) is an essential component of the RNA-processing machinery which generates the 3' end of mature histone mRNA in the sea urchin. The U7 small nuclear ribonucleoprotein particle (snRNP) is classified as a member of the Sm-type U snRNP family by virtue of its recognition by both anti-trimethylguanosine and anti-Sm antibodies. We analyzed the function-structure relationship of the U7 snRNP by mutagenesis experiments. These suggested that the U7 snRNP of the sea urchin is composed of three important domains. The first domain encompasses the 5'-terminal sequences, up to about nucleotides 7, which are accessible to micrococcal nuclease, while the remainder of the RNA is highly protected and hence presumably bound by proteins. This region contains the sequence complementarities between the U7 snRNA and the histone pre-mRNA which have previously been shown to be required for 3' processing (F. Schaufele, G. M. Gilmartin, W. Bannwarth, and M. L. Birnstiel, Nature [London] 323:777-781, 1986). Nucleotides 9 to 20 constitute a second domain which includes sequences for Sm protein binding. The complementarities between the U7 snRNA sequences in this region and the terminal palindrome of the histone mRNA appear to be fortuitous and play only a secondary, if any, role in 3' processing. The third domain is composed of the terminal palindrome of U7 snRNA, the secondary structure of which must be maintained for the U7 snRNP to function, but its sequence can be drastically altered without any observable effect on snRNP assembly or 3' processing.
Full text
PDF


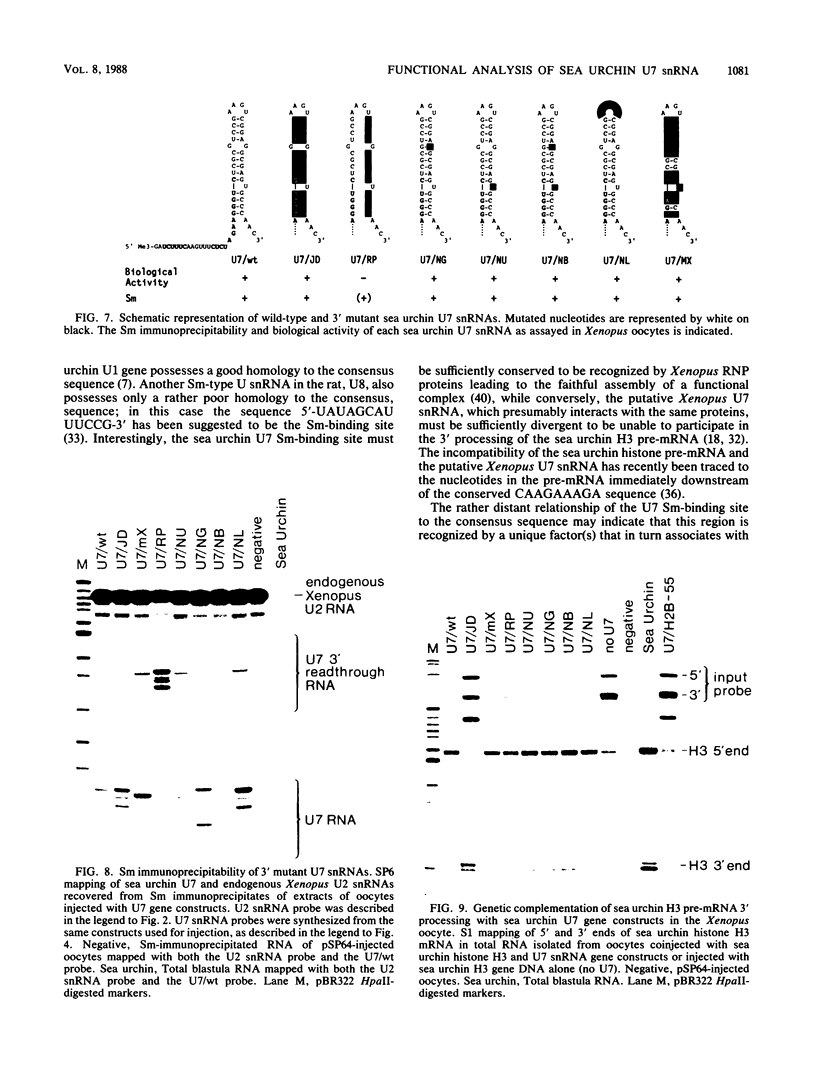



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ares M., Jr U2 RNA from yeast is unexpectedly large and contains homology to vertebrate U4, U5, and U6 small nuclear RNAs. Cell. 1986 Oct 10;47(1):49–59. doi: 10.1016/0092-8674(86)90365-x. [DOI] [PubMed] [Google Scholar]
- Berget S. M., Robberson B. L. U1, U2, and U4/U6 small nuclear ribonucleoproteins are required for in vitro splicing but not polyadenylation. Cell. 1986 Aug 29;46(5):691–696. doi: 10.1016/0092-8674(86)90344-2. [DOI] [PubMed] [Google Scholar]
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- Black D. L., Chabot B., Steitz J. A. U2 as well as U1 small nuclear ribonucleoproteins are involved in premessenger RNA splicing. Cell. 1985 Oct;42(3):737–750. doi: 10.1016/0092-8674(85)90270-3. [DOI] [PubMed] [Google Scholar]
- Black D. L., Steitz J. A. Pre-mRNA splicing in vitro requires intact U4/U6 small nuclear ribonucleoprotein. Cell. 1986 Aug 29;46(5):697–704. doi: 10.1016/0092-8674(86)90345-4. [DOI] [PubMed] [Google Scholar]
- Branlant C., Krol A., Ebel J. P., Lazar E., Haendler B., Jacob M. U2 RNA shares a structural domain with U1, U4, and U5 RNAs. EMBO J. 1982;1(10):1259–1265. doi: 10.1002/j.1460-2075.1982.tb00022.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown D. T., Morris G. F., Chodchoy N., Sprecher C., Marzluff W. F. Structure of the sea urchin U1 RNA repeat. Nucleic Acids Res. 1985 Jan 25;13(2):537–556. doi: 10.1093/nar/13.2.537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busch H., Reddy R., Rothblum L., Choi Y. C. SnRNAs, SnRNPs, and RNA processing. Annu Rev Biochem. 1982;51:617–654. doi: 10.1146/annurev.bi.51.070182.003153. [DOI] [PubMed] [Google Scholar]
- Chabot B., Black D. L., LeMaster D. M., Steitz J. A. The 3' splice site of pre-messenger RNA is recognized by a small nuclear ribonucleoprotein. Science. 1985 Dec 20;230(4732):1344–1349. doi: 10.1126/science.2933810. [DOI] [PubMed] [Google Scholar]
- Ciliberto G., Dathan N., Frank R., Philipson L., Mattaj I. W. Formation of the 3' end on U snRNAs requires at least three sequence elements. EMBO J. 1986 Nov;5(11):2931–2937. doi: 10.1002/j.1460-2075.1986.tb04589.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Lorenzi M., Rohrer U., Birnstiel M. L. Analysis of a sea urchin gene cluster coding for the small nuclear U7 RNA, a rare RNA species implicated in the 3' editing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1986 May;83(10):3243–3247. doi: 10.1073/pnas.83.10.3243. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Robertis E. M., Lienhard S., Parisot R. F. Intracellular transport of microinjected 5S and small nuclear RNAs. Nature. 1982 Feb 18;295(5850):572–577. doi: 10.1038/295572a0. [DOI] [PubMed] [Google Scholar]
- Galli G., Hofstetter H., Stunnenberg H. G., Birnstiel M. L. Biochemical complementation with RNA in the Xenopus oocyte: a small RNA is required for the generation of 3' histone mRNA termini. Cell. 1983 Oct;34(3):823–828. doi: 10.1016/0092-8674(83)90539-1. [DOI] [PubMed] [Google Scholar]
- Gick O., Krämer A., Keller W., Birnstiel M. L. Generation of histone mRNA 3' ends by endonucleolytic cleavage of the pre-mRNA in a snRNP-dependent in vitro reaction. EMBO J. 1986 Jun;5(6):1319–1326. doi: 10.1002/j.1460-2075.1986.tb04362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gold H. A., Altman S. Reconstitution of RNAase P activity using inactive subunits from E. coli and HeLa cells. Cell. 1986 Jan 31;44(2):243–249. doi: 10.1016/0092-8674(86)90758-0. [DOI] [PubMed] [Google Scholar]
- Guerrier-Takada C., Gardiner K., Marsh T., Pace N., Altman S. The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell. 1983 Dec;35(3 Pt 2):849–857. doi: 10.1016/0092-8674(83)90117-4. [DOI] [PubMed] [Google Scholar]
- Hashimoto C., Steitz J. A. A small nuclear ribonucleoprotein associates with the AAUAAA polyadenylation signal in vitro. Cell. 1986 May 23;45(4):581–591. doi: 10.1016/0092-8674(86)90290-4. [DOI] [PubMed] [Google Scholar]
- Hentschel C., Probst E., Birnstiel M. L. Transcriptional fidelity of histone genes injected into Xenopus oocyte nuclei. Nature. 1980 Nov 6;288(5786):100–102. doi: 10.1038/288100a0. [DOI] [PubMed] [Google Scholar]
- Hernandez N. Formation of the 3' end of U1 snRNA is directed by a conserved sequence located downstream of the coding region. EMBO J. 1985 Jul;4(7):1827–1837. doi: 10.1002/j.1460-2075.1985.tb03857.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krainer A. R., Maniatis T. Multiple factors including the small nuclear ribonucleoproteins U1 and U2 are necessary for pre-mRNA splicing in vitro. Cell. 1985 Oct;42(3):725–736. doi: 10.1016/0092-8674(85)90269-7. [DOI] [PubMed] [Google Scholar]
- Kretzner L., Rymond B. C., Rosbash M. S. cerevisiae U1 RNA is large and has limited primary sequence homology to metazoan U1 snRNA. Cell. 1987 Aug 14;50(4):593–602. doi: 10.1016/0092-8674(87)90032-8. [DOI] [PubMed] [Google Scholar]
- Krämer A., Keller W., Appel B., Lührmann R. The 5' terminus of the RNA moiety of U1 small nuclear ribonucleoprotein particles is required for the splicing of messenger RNA precursors. Cell. 1984 Aug;38(1):299–307. doi: 10.1016/0092-8674(84)90551-8. [DOI] [PubMed] [Google Scholar]
- Lerner M. R., Steitz J. A. Antibodies to small nuclear RNAs complexed with proteins are produced by patients with systemic lupus erythematosus. Proc Natl Acad Sci U S A. 1979 Nov;76(11):5495–5499. doi: 10.1073/pnas.76.11.5495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liautard J. P., Sri-Widada J., Brunel C., Jeanteur P. Structural organization of ribonucleoproteins containing small nuclear RNAs from HeLa cells. Proteins interact closely with a similar structural domain of U1, U2, U4 and U5 small nuclear RNAs. J Mol Biol. 1982 Dec 15;162(3):623–643. doi: 10.1016/0022-2836(82)90392-8. [DOI] [PubMed] [Google Scholar]
- Mattaj I. W. Cap trimethylation of U snRNA is cytoplasmic and dependent on U snRNP protein binding. Cell. 1986 Sep 12;46(6):905–911. doi: 10.1016/0092-8674(86)90072-3. [DOI] [PubMed] [Google Scholar]
- Mattaj I. W., De Robertis E. M. Nuclear segregation of U2 snRNA requires binding of specific snRNP proteins. Cell. 1985 Jan;40(1):111–118. doi: 10.1016/0092-8674(85)90314-9. [DOI] [PubMed] [Google Scholar]
- Moore C. L., Sharp P. A. Accurate cleavage and polyadenylation of exogenous RNA substrate. Cell. 1985 Jul;41(3):845–855. doi: 10.1016/s0092-8674(85)80065-9. [DOI] [PubMed] [Google Scholar]
- Moore C. L., Sharp P. A. Site-specific polyadenylation in a cell-free reaction. Cell. 1984 Mar;36(3):581–591. doi: 10.1016/0092-8674(84)90337-4. [DOI] [PubMed] [Google Scholar]
- Mowry K. L., Steitz J. A. Both conserved signals on mammalian histone pre-mRNAs associate with small nuclear ribonucleoproteins during 3' end formation in vitro. Mol Cell Biol. 1987 May;7(5):1663–1672. doi: 10.1128/mcb.7.5.1663. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patterson B., Guthrie C. An essential yeast snRNA with a U5-like domain is required for splicing in vivo. Cell. 1987 Jun 5;49(5):613–624. doi: 10.1016/0092-8674(87)90537-x. [DOI] [PubMed] [Google Scholar]
- Probst E., Kressmann A., Birnstiel M. L. Expression of sea urchin histone genes in the oocyte of Xenopus laevis. J Mol Biol. 1979 Dec 15;135(3):709–732. doi: 10.1016/0022-2836(79)90173-6. [DOI] [PubMed] [Google Scholar]
- Reddy R., Henning D., Busch H. Primary and secondary structure of U8 small nuclear RNA. J Biol Chem. 1985 Sep 15;260(20):10930–10935. [PubMed] [Google Scholar]
- Riedel N., Wolin S., Guthrie C. A subset of yeast snRNA's contains functional binding sites for the highly conserved Sm antigen. Science. 1987 Jan 16;235(4786):328–331. doi: 10.1126/science.2948278. [DOI] [PubMed] [Google Scholar]
- Schaffner W., Kunz G., Daetwyler H., Telford J., Smith H. O., Birnstiel M. L. Genes and spacers of cloned sea urchin histone DNA analyzed by sequencing. Cell. 1978 Jul;14(3):655–671. doi: 10.1016/0092-8674(78)90249-0. [DOI] [PubMed] [Google Scholar]
- Schaufele F., Birnstiel M. L. The inability of the Psammechinus miliaris H3 RNA to be processed in the Xenopus oocyte is associated with sequences distinct from those highly conserved amongst sea urchin histone RNAs. Nucleic Acids Res. 1987 Oct 26;15(20):8305–8317. doi: 10.1093/nar/15.20.8305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schaufele F., Gilmartin G. M., Bannwarth W., Birnstiel M. L. Compensatory mutations suggest that base-pairing with a small nuclear RNA is required to form the 3' end of H3 messenger RNA. 1986 Oct 30-Nov 5Nature. 323(6091):777–781. doi: 10.1038/323777a0. [DOI] [PubMed] [Google Scholar]
- Siliciano P. G., Brow D. A., Roiha H., Guthrie C. An essential snRNA from S. cerevisiae has properties predicted for U4, including interaction with a U6-like snRNA. Cell. 1987 Aug 14;50(4):585–592. doi: 10.1016/0092-8674(87)90031-6. [DOI] [PubMed] [Google Scholar]
- Sperry A. O., Berget S. M. In vitro cleavage of the simian virus 40 early polyadenylation site adjacent to a required downstream TG sequence. Mol Cell Biol. 1986 Dec;6(12):4734–4741. doi: 10.1128/mcb.6.12.4734. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Birnstiel M. L. Genetic complementation in the Xenopus oocyte: co-expression of sea urchin histone and U7 RNAs restores 3' processing of H3 pre-mRNA in the oocyte. EMBO J. 1986 Jul;5(7):1675–1682. doi: 10.1002/j.1460-2075.1986.tb04411.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Galli G., Busslinger M., Birnstiel M. L. The cDNA sequences of the sea urchin U7 small nuclear RNA suggest specific contacts between histone mRNA precursor and U7 RNA during RNA processing. EMBO J. 1984 Dec 1;3(12):2801–2807. doi: 10.1002/j.1460-2075.1984.tb02212.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weaver R. F., Weissmann C. Mapping of RNA by a modification of the Berk-Sharp procedure: the 5' termini of 15 S beta-globin mRNA precursor and mature 10 s beta-globin mRNA have identical map coordinates. Nucleic Acids Res. 1979 Nov 10;7(5):1175–1193. doi: 10.1093/nar/7.5.1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yuo C. Y., Ares M., Jr, Weiner A. M. Sequences required for 3' end formation of human U2 small nuclear RNA. Cell. 1985 Aug;42(1):193–202. doi: 10.1016/s0092-8674(85)80115-x. [DOI] [PubMed] [Google Scholar]
- Zhuang Y., Weiner A. M. A compensatory base change in U1 snRNA suppresses a 5' splice site mutation. Cell. 1986 Sep 12;46(6):827–835. doi: 10.1016/0092-8674(86)90064-4. [DOI] [PubMed] [Google Scholar]
- Zinn K., DiMaio D., Maniatis T. Identification of two distinct regulatory regions adjacent to the human beta-interferon gene. Cell. 1983 Oct;34(3):865–879. doi: 10.1016/0092-8674(83)90544-5. [DOI] [PubMed] [Google Scholar]