Abstract
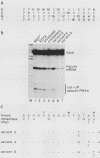
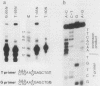
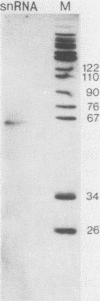
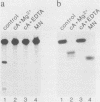
Oligonucleotides derived from the spacer element of the histone RNA 3' processing signal were used to characterize mouse U7 small nuclear RNA (snRNA), i.e., the snRNA component active in 3' processing of histone pre-mRNA. Under RNase H conditions, such oligonucleotides inhibited the processing reaction, indicating the formation of a DNA-RNA hybrid with a functional ribonucleoprotein component. Moreover, these oligonucleotides hybridized to a single nuclear RNA species of approximately 65 nucleotides. The sequence of this RNA was determined by primer extension experiments and was found to bear several structural similarities with sea urchin U7 snRNA. The comparison of mouse and sea urchin U7 snRNA structures yields some further insight into the mechanism of histone RNA 3' processing.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Billeter M. A., Baczko K., Schmid A., Ter Meulen V. Cloning of DNA corresponding to four different measles virus genomic regions. Virology. 1984 Jan 15;132(1):147–159. doi: 10.1016/0042-6822(84)90099-0. [DOI] [PubMed] [Google Scholar]
- Birchmeier C., Folk W., Birnstiel M. L. The terminal RNA stem-loop structure and 80 bp of spacer DNA are required for the formation of 3' termini of sea urchin H2A mRNA. Cell. 1983 Dec;35(2 Pt 1):433–440. doi: 10.1016/0092-8674(83)90176-9. [DOI] [PubMed] [Google Scholar]
- Birchmeier C., Schümperli D., Sconzo G., Birnstiel M. L. 3' editing of mRNAs: sequence requirements and involvement of a 60-nucleotide RNA in maturation of histone mRNA precursors. Proc Natl Acad Sci U S A. 1984 Feb;81(4):1057–1061. doi: 10.1073/pnas.81.4.1057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnstiel M. L., Busslinger M., Strub K. Transcription termination and 3' processing: the end is in site! Cell. 1985 Jun;41(2):349–359. doi: 10.1016/s0092-8674(85)80007-6. [DOI] [PubMed] [Google Scholar]
- Coles L. S., Wells J. R. An H1 histone gene-specific 5' element and evolution of H1 and H5 genes. Nucleic Acids Res. 1985 Jan 25;13(2):585–594. doi: 10.1093/nar/13.2.585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D'Andrea R., Harvey R., Wells J. R. Vertebrate histone genes: nucleotide sequence of a chicken H2A gene and regulatory flanking sequences. Nucleic Acids Res. 1981 Jul 10;9(13):3119–3128. doi: 10.1093/nar/9.13.3119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- England T. E., Uhlenbeck O. C. 3'-terminal labelling of RNA with T4 RNA ligase. Nature. 1978 Oct 12;275(5680):560–561. doi: 10.1038/275560a0. [DOI] [PubMed] [Google Scholar]
- Georgiev O., Birnstiel M. L. The conserved CAAGAAAGA spacer sequence is an essential element for the formation of 3' termini of the sea urchin H3 histone mRNA by RNA processing. EMBO J. 1985 Feb;4(2):481–489. doi: 10.1002/j.1460-2075.1985.tb03654.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Keller W., Birnstiel M. L. Generation of histone mRNA 3' ends by endonucleolytic cleavage of the pre-mRNA in a snRNP-dependent in vitro reaction. EMBO J. 1986 Jun;5(6):1319–1326. doi: 10.1002/j.1460-2075.1986.tb04362.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gick O., Krämer A., Vasserot A., Birnstiel M. L. Heat-labile regulatory factor is required for 3' processing of histone precursor mRNAs. Proc Natl Acad Sci U S A. 1987 Dec;84(24):8937–8940. doi: 10.1073/pnas.84.24.8937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gilmartin G. M., Schaufele F., Schaffner G., Birnstiel M. L. Functional analysis of the sea urchin U7 small nuclear RNA. Mol Cell Biol. 1988 Mar;8(3):1076–1084. doi: 10.1128/mcb.8.3.1076. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harvey R. P., Robins A. J., Wells J. R. Independently evolving chicken histone H2B genes: identification of a ubiquitous H2B-specific 5' element. Nucleic Acids Res. 1982 Dec 11;10(23):7851–7863. doi: 10.1093/nar/10.23.7851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heintz N., Zernik M., Roeder R. G. The structure of the human histone genes: clustered but not tandemly repeated. Cell. 1981 Jun;24(3):661–668. doi: 10.1016/0092-8674(81)90092-1. [DOI] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Krainer A. R., Maniatis T. Multiple factors including the small nuclear ribonucleoproteins U1 and U2 are necessary for pre-mRNA splicing in vitro. Cell. 1985 Oct;42(3):725–736. doi: 10.1016/0092-8674(85)90269-7. [DOI] [PubMed] [Google Scholar]
- Krieg P. A., Melton D. A. Formation of the 3' end of histone mRNA by post-transcriptional processing. Nature. 1984 Mar 8;308(5955):203–206. doi: 10.1038/308203a0. [DOI] [PubMed] [Google Scholar]
- Krämer A., Keller W., Appel B., Lührmann R. The 5' terminus of the RNA moiety of U1 small nuclear ribonucleoprotein particles is required for the splicing of messenger RNA precursors. Cell. 1984 Aug;38(1):299–307. doi: 10.1016/0092-8674(84)90551-8. [DOI] [PubMed] [Google Scholar]
- Lüscher B., Schümperli D. RNA 3' processing regulates histone mRNA levels in a mammalian cell cycle mutant. A processing factor becomes limiting in G1-arrested cells. EMBO J. 1987 Jun;6(6):1721–1726. doi: 10.1002/j.1460-2075.1987.tb02423.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Reddy R., Henning D., Busch H. Primary and secondary structure of U8 small nuclear RNA. J Biol Chem. 1985 Sep 15;260(20):10930–10935. [PubMed] [Google Scholar]
- Schaufele F., Gilmartin G. M., Bannwarth W., Birnstiel M. L. Compensatory mutations suggest that base-pairing with a small nuclear RNA is required to form the 3' end of H3 messenger RNA. 1986 Oct 30-Nov 5Nature. 323(6091):777–781. doi: 10.1038/323777a0. [DOI] [PubMed] [Google Scholar]
- Seiler-Tuyns A., Birnstiel M. L. Structure and expression in L-cells of a cloned H4 histone gene of the mouse. J Mol Biol. 1981 Oct 5;151(4):607–625. doi: 10.1016/0022-2836(81)90426-5. [DOI] [PubMed] [Google Scholar]
- Sierra F., Stein G., Stein J. Structure and in vitro transcription of a human H4 histone gene. Nucleic Acids Res. 1983 Oct 25;11(20):7069–7086. doi: 10.1093/nar/11.20.7069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sippel A. E. Purification and characterization of adenosine triphosphate: ribonucleic acid adenyltransferase from Escherichia coli. Eur J Biochem. 1973 Aug 1;37(1):31–40. doi: 10.1111/j.1432-1033.1973.tb02953.x. [DOI] [PubMed] [Google Scholar]
- Sittman D. B., Graves R. A., Marzluff W. F. Structure of a cluster of mouse histone genes. Nucleic Acids Res. 1983 Oct 11;11(19):6679–6697. doi: 10.1093/nar/11.19.6679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stauber C., Lüscher B., Eckner R., Lötscher E., Schümperli D. A signal regulating mouse histone H4 mRNA levels in a mammalian cell cycle mutant and sequences controlling RNA 3' processing are both contained within the same 80-bp fragment. EMBO J. 1986 Dec 1;5(12):3297–3303. doi: 10.1002/j.1460-2075.1986.tb04643.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Birnstiel M. L. Genetic complementation in the Xenopus oocyte: co-expression of sea urchin histone and U7 RNAs restores 3' processing of H3 pre-mRNA in the oocyte. EMBO J. 1986 Jul;5(7):1675–1682. doi: 10.1002/j.1460-2075.1986.tb04411.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strub K., Galli G., Busslinger M., Birnstiel M. L. The cDNA sequences of the sea urchin U7 small nuclear RNA suggest specific contacts between histone mRNA precursor and U7 RNA during RNA processing. EMBO J. 1984 Dec 1;3(12):2801–2807. doi: 10.1002/j.1460-2075.1984.tb02212.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stunnenberg H. G., Birnstiel M. L. Bioassay for components regulating eukaryotic gene expression: a chromosomal factor involved in the generation of histone mRNA 3' termini. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6201–6204. doi: 10.1073/pnas.79.20.6201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugarman B. J., Dodgson J. B., Engel J. D. Genomic organization, DNA sequence, and expression of chicken embryonic histone genes. J Biol Chem. 1983 Jul 25;258(14):9005–9016. [PubMed] [Google Scholar]
- Taylor J. D., Wellman S. E., Marzluff W. F. Sequences of four mouse histone H3 genes: implications for evolution of mouse histone genes. J Mol Evol. 1986;23(3):242–249. doi: 10.1007/BF02115580. [DOI] [PubMed] [Google Scholar]
- Turner P. C., Woodland H. R. H3 and H4 histone cDNA sequences from Xenopus: a sequence comparison of H4 genes. Nucleic Acids Res. 1982 Jun 25;10(12):3769–3780. doi: 10.1093/nar/10.12.3769. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhong R., Roeder R. G., Heintz N. The primary structure and expression of four cloned human histone genes. Nucleic Acids Res. 1983 Nov 11;11(21):7409–7425. doi: 10.1093/nar/11.21.7409. [DOI] [PMC free article] [PubMed] [Google Scholar]