Abstract
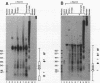
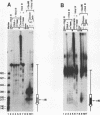
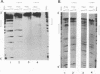
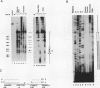
We describe the chromosomal organization of the major oocyte and somatic 5S RNA genes of Xenopus laevis in chromatin isolated from erythrocyte nuclei. Both major oocyte and somatic 5S DNA repeats are associated with nucleosomes; however, differences exist in the organization of chromatin over the oocyte and somatic 5S RNA genes. The repressed oocyte 5S RNA gene is protected from nuclease digestion by incorporation into a nucleosome, and the entire oocyte 5S DNA repeat is assembled into a loosely positioned array of nucleosomes. In contrast, the potentially active somatic 5S RNA gene is accessible to nuclease digestion, and the majority of somatic 5S RNA genes appear not to be incorporated into positioned nucleosomes. Evidence is presented supporting the stable association of transcription factors with the somatic 5S RNA genes. Histone H1 is shown to have a role both in determining the organization of nucleosomes over the oocyte 5S DNA repeat and in repressing transcription of the oocyte 5S RNA genes.
Full text
PDF










Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allan J., Cowling G. J., Harborne N., Cattini P., Craigie R., Gould H. Regulation of the higher-order structure of chromatin by histones H1 and H5. J Cell Biol. 1981 Aug;90(2):279–288. doi: 10.1083/jcb.90.2.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Allan J., Hartman P. G., Crane-Robinson C., Aviles F. X. The structure of histone H1 and its location in chromatin. Nature. 1980 Dec 25;288(5792):675–679. doi: 10.1038/288675a0. [DOI] [PubMed] [Google Scholar]
- Almouzni G., Méchali M., Wolffe A. P. Competition between transcription complex assembly and chromatin assembly on replicating DNA. EMBO J. 1990 Feb;9(2):573–582. doi: 10.1002/j.1460-2075.1990.tb08145.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Almouzni G., Méchali M., Wolffe A. P. Transcription complex disruption caused by a transition in chromatin structure. Mol Cell Biol. 1991 Feb;11(2):655–665. doi: 10.1128/mcb.11.2.655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Andrews M. T., Brown D. D. Transient activation of oocyte 5S RNA genes in Xenopus embryos by raising the level of the trans-acting factor TFIIIA. Cell. 1987 Nov 6;51(3):445–453. doi: 10.1016/0092-8674(87)90640-4. [DOI] [PubMed] [Google Scholar]
- Axelrod J. D., Majors J. An improved method for photofootprinting yeast genes in vivo using Taq polymerase. Nucleic Acids Res. 1989 Jan 11;17(1):171–183. doi: 10.1093/nar/17.1.171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birkenmeier E. H., Brown D. D., Jordan E. A nuclear extract of Xenopus laevis oocytes that accurately transcribes 5S RNA genes. Cell. 1978 Nov;15(3):1077–1086. doi: 10.1016/0092-8674(78)90291-x. [DOI] [PubMed] [Google Scholar]
- Blanco J., Millstein L., Razik M. A., Dilworth S., Cote C., Gottesfeld J. Two TFIIIA activities regulate expression of the Xenopus 5S RNA gene families. Genes Dev. 1989 Oct;3(10):1602–1612. doi: 10.1101/gad.3.10.1602. [DOI] [PubMed] [Google Scholar]
- Bryan P. N., Hofstetter H., Birnstiel M. L. Nucleosome arrangement on tRNA genes of Xenopus laevis. Cell. 1981 Dec;27(3 Pt 2):459–466. doi: 10.1016/0092-8674(81)90387-1. [DOI] [PubMed] [Google Scholar]
- Carroll D., Brown D. D. Repeating units of Xenopus laevis oocyte-type 5S DNA are heterogeneous in length. Cell. 1976 Apr;7(4):467–475. doi: 10.1016/0092-8674(76)90198-7. [DOI] [PubMed] [Google Scholar]
- Carroll D., Wright S. H., Ajioka R. S., Hussey C. E., Jr Genetic recombination of Xenopus laevis 5 S DNA in bacteria. J Mol Biol. 1984 Sep 15;178(2):155–172. doi: 10.1016/0022-2836(84)90137-2. [DOI] [PubMed] [Google Scholar]
- Church G. M., Gilbert W. Genomic sequencing. Proc Natl Acad Sci U S A. 1984 Apr;81(7):1991–1995. doi: 10.1073/pnas.81.7.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clarkson S. G., Kurer V., Smith H. O. Sequence organization of a cloned tDNA met fragment from Xenopus laevis. Cell. 1978 Jul;14(3):713–724. doi: 10.1016/0092-8674(78)90253-2. [DOI] [PubMed] [Google Scholar]
- Cozzarelli N. R., Gerrard S. P., Schlissel M., Brown D. D., Bogenhagen D. F. Purified RNA polymerase III accurately and efficiently terminates transcription of 5S RNA genes. Cell. 1983 Oct;34(3):829–835. doi: 10.1016/0092-8674(83)90540-8. [DOI] [PubMed] [Google Scholar]
- Croston G. E., Kerrigan L. A., Lira L. M., Marshak D. R., Kadonaga J. T. Sequence-specific antirepression of histone H1-mediated inhibition of basal RNA polymerase II transcription. Science. 1991 Feb 8;251(4994):643–649. doi: 10.1126/science.1899487. [DOI] [PubMed] [Google Scholar]
- Darby M. K., Andrews M. T., Brown D. D. Transcription complexes that program Xenopus 5S RNA genes are stable in vivo. Proc Natl Acad Sci U S A. 1988 Aug;85(15):5516–5520. doi: 10.1073/pnas.85.15.5516. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelke D. R., Gottesfeld J. M. Chromosomal footprinting of transcriptionally active and inactive oocyte-type 5S RNA genes of Xenopus laevis. Nucleic Acids Res. 1990 Oct 25;18(20):6031–6037. doi: 10.1093/nar/18.20.6031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelke D. R., Ng S. Y., Shastry B. S., Roeder R. G. Specific interaction of a purified transcription factor with an internal control region of 5S RNA genes. Cell. 1980 Mar;19(3):717–728. doi: 10.1016/s0092-8674(80)80048-1. [DOI] [PubMed] [Google Scholar]
- Fairall L., Rhodes D., Klug A. Mapping of the sites of protection on a 5 S RNA gene by the Xenopus transcription factor IIIA. A model for the interaction. J Mol Biol. 1986 Dec 5;192(3):577–591. doi: 10.1016/0022-2836(86)90278-0. [DOI] [PubMed] [Google Scholar]
- Fedoroff N. V., Brown D. D. The nucleotide sequence of oocyte 5S DNA in Xenopus laevis. I. The AT-rich spacer. Cell. 1978 Apr;13(4):701–716. doi: 10.1016/0092-8674(78)90220-9. [DOI] [PubMed] [Google Scholar]
- Gottesfeld J. M., Bloomer L. S. Nonrandom alignment of nucleosomes on 5S RNA genes of X. laevis. Cell. 1980 Oct;21(3):751–760. doi: 10.1016/0092-8674(80)90438-9. [DOI] [PubMed] [Google Scholar]
- Gurdon J. B., Dingwall C., Laskey R. A., Korn L. J. Developmental inactivity of 5S RNA genes persists when chromosomes are cut between genes. Nature. 1982 Oct 14;299(5884):652–653. doi: 10.1038/299652a0. [DOI] [PubMed] [Google Scholar]
- Hentschel C. C., Tata J. R. Template-engaged and free RNA polymerases during Xenopus erythroid cell maturation. Dev Biol. 1978 Aug;65(2):496–507. doi: 10.1016/0012-1606(78)90044-1. [DOI] [PubMed] [Google Scholar]
- Huibregtse J. M., Engelke D. R. Genomic footprinting of a yeast tRNA gene reveals stable complexes over the 5'-flanking region. Mol Cell Biol. 1989 Aug;9(8):3244–3252. doi: 10.1128/mcb.9.8.3244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huibregtse J. M., Evans C. F., Engelke D. R. Comparison of tRNA gene transcription complexes formed in vitro and in nuclei. Mol Cell Biol. 1987 Sep;7(9):3212–3220. doi: 10.1128/mcb.7.9.3212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Humphries S. E., Young D., Carroll D. Chromatin structure of the 5S ribonucleic acid genes of Xenopus laevis. Biochemistry. 1979 Jul 24;18(15):3223–3231. doi: 10.1021/bi00582a006. [DOI] [PubMed] [Google Scholar]
- Izaurralde E., Käs E., Laemmli U. K. Highly preferential nucleation of histone H1 assembly on scaffold-associated regions. J Mol Biol. 1989 Dec 5;210(3):573–585. doi: 10.1016/0022-2836(89)90133-2. [DOI] [PubMed] [Google Scholar]
- Jerzmanowski A., Cole R. D. Flanking sequences of Xenopus 5 S RNA genes determine differential inhibition of transcription by H1 histone in vitro. Mitotic phosphorylation of H1 decreases its inhibitory power. J Biol Chem. 1990 Jun 25;265(18):10726–10732. [PubMed] [Google Scholar]
- Kassavetis G. A., Braun B. R., Nguyen L. H., Geiduschek E. P. S. cerevisiae TFIIIB is the transcription initiation factor proper of RNA polymerase III, while TFIIIA and TFIIIC are assembly factors. Cell. 1990 Jan 26;60(2):235–245. doi: 10.1016/0092-8674(90)90739-2. [DOI] [PubMed] [Google Scholar]
- Losa R., Thoma F., Koller T. Involvement of the globular domain of histone H1 in the higher order structures of chromatin. J Mol Biol. 1984 Jun 5;175(4):529–551. doi: 10.1016/0022-2836(84)90183-9. [DOI] [PubMed] [Google Scholar]
- Marekov L. N., Beltchev B. Selective removal of histone H1 from chromatin at low salt concentration. Anal Biochem. 1981 Jul 15;115(1):93–96. doi: 10.1016/0003-2697(81)90529-7. [DOI] [PubMed] [Google Scholar]
- McGhee J. D., Rau D. C., Felsenfeld G. The high mobility group proteins HMG 14 and 17, do not prevent the formation of chromatin higher order structure. Nucleic Acids Res. 1982 Mar 25;10(6):2007–2016. doi: 10.1093/nar/10.6.2007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson R. C., Doering J. L., Brown D. D. Characterization of two xenopus somatic 5S DNAs and one minor oocyte-specific 5S DNA. Cell. 1980 May;20(1):131–141. doi: 10.1016/0092-8674(80)90241-x. [DOI] [PubMed] [Google Scholar]
- Renz M., Day L. A. Transition from noncooperative to cooperative and selective binding of histone H1 to DNA. Biochemistry. 1976 Jul 27;15(15):3220–3228. doi: 10.1021/bi00660a010. [DOI] [PubMed] [Google Scholar]
- Rhodes D. Structural analysis of a triple complex between the histone octamer, a Xenopus gene for 5S RNA and transcription factor IIIA. EMBO J. 1985 Dec 16;4(13A):3473–3482. doi: 10.1002/j.1460-2075.1985.tb04106.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Satchwell S. C., Travers A. A. Asymmetry and polarity of nucleosomes in chicken erythrocyte chromatin. EMBO J. 1989 Jan;8(1):229–238. doi: 10.1002/j.1460-2075.1989.tb03368.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schlissel M. S., Brown D. D. The transcriptional regulation of Xenopus 5s RNA genes in chromatin: the roles of active stable transcription complexes and histone H1. Cell. 1984 Jul;37(3):903–913. doi: 10.1016/0092-8674(84)90425-2. [DOI] [PubMed] [Google Scholar]
- Searle P. F., Tata J. R. Vitellogenin gene expression in male Xenopus hepatocytes during primary and secondary stimulation with estrogen in cell cultures. Cell. 1981 Mar;23(3):741–746. doi: 10.1016/0092-8674(81)90437-2. [DOI] [PubMed] [Google Scholar]
- Shimamura A., Sapp M., Rodriguez-Campos A., Worcel A. Histone H1 represses transcription from minichromosomes assembled in vitro. Mol Cell Biol. 1989 Dec;9(12):5573–5584. doi: 10.1128/mcb.9.12.5573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimamura A., Tremethick D., Worcel A. Characterization of the repressed 5S DNA minichromosomes assembled in vitro with a high-speed supernatant of Xenopus laevis oocytes. Mol Cell Biol. 1988 Oct;8(10):4257–4269. doi: 10.1128/mcb.8.10.4257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith D. R., Jackson I. J., Brown D. D. Domains of the positive transcription factor specific for the Xenopus 5S RNA gene. Cell. 1984 Jun;37(2):645–652. doi: 10.1016/0092-8674(84)90396-9. [DOI] [PubMed] [Google Scholar]
- Staynov D. Z., Crane-Robinson C. Footprinting of linker histones H5 and H1 on the nucleosome. EMBO J. 1988 Dec 1;7(12):3685–3691. doi: 10.1002/j.1460-2075.1988.tb03250.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoma F., Koller T., Klug A. Involvement of histone H1 in the organization of the nucleosome and of the salt-dependent superstructures of chromatin. J Cell Biol. 1979 Nov;83(2 Pt 1):403–427. doi: 10.1083/jcb.83.2.403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoma F., Zatchej M. Chromatin folding modulates nucleosome positioning in yeast minichromosomes. Cell. 1988 Dec 23;55(6):945–953. doi: 10.1016/0092-8674(88)90240-1. [DOI] [PubMed] [Google Scholar]
- Wakefield L., Gurdon J. B. Cytoplasmic regulation of 5S RNA genes in nuclear-transplant embryos. EMBO J. 1983;2(9):1613–1619. doi: 10.1002/j.1460-2075.1983.tb01632.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Widmer R. M., Lezzi M., Koller T. Structural transition in inactive Balbiani ring chromatin of Chironomus during micrococcus nuclease digestion. EMBO J. 1987 Mar;6(3):743–748. doi: 10.1002/j.1460-2075.1987.tb04816.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Brown D. D. Developmental regulation of two 5S ribosomal RNA genes. Science. 1988 Sep 23;241(4873):1626–1632. doi: 10.1126/science.241.4873.1626. [DOI] [PubMed] [Google Scholar]
- Wolffe A. P., Brown D. D. Differential 5S RNA gene expression in vitro. Cell. 1987 Dec 4;51(5):733–740. doi: 10.1016/0092-8674(87)90096-1. [DOI] [PubMed] [Google Scholar]
- Wolffe A. P. Dominant and specific repression of Xenopus oocyte 5S RNA genes and satellite I DNA by histone H1. EMBO J. 1989 Feb;8(2):527–537. doi: 10.1002/j.1460-2075.1989.tb03407.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P., Jordan E., Brown D. D. A bacteriophage RNA polymerase transcribes through a Xenopus 5S RNA gene transcription complex without disrupting it. Cell. 1986 Feb 14;44(3):381–389. doi: 10.1016/0092-8674(86)90459-9. [DOI] [PubMed] [Google Scholar]
- Wolffe A. P., Morse R. H. The transcription complex of the Xenopus somatic 5 S RNA gene. A functional analysis of protein-DNA interactions outside of the internal control region. J Biol Chem. 1990 Mar 15;265(8):4592–4599. [PubMed] [Google Scholar]
- Wolffe A. P. Transcription fraction TFIIIC can regulate differential Xenopus 5S RNA gene transcription in vitro. EMBO J. 1988 Apr;7(4):1071–1079. doi: 10.1002/j.1460-2075.1988.tb02915.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolffe A. P. Transcriptional activation of Xenopus class III genes in chromatin isolated from sperm and somatic nuclei. Nucleic Acids Res. 1989 Jan 25;17(2):767–780. doi: 10.1093/nar/17.2.767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wormington W. M., Brown D. D. Onset of 5 S RNA gene regulation during Xenopus embryogenesis. Dev Biol. 1983 Sep;99(1):248–257. doi: 10.1016/0012-1606(83)90273-7. [DOI] [PubMed] [Google Scholar]
- Young D., Carroll D. Regular arrangement of nucleosomes on 5S rRNA genes in Xenopus laevis. Mol Cell Biol. 1983 Apr;3(4):720–730. doi: 10.1128/mcb.3.4.720. [DOI] [PMC free article] [PubMed] [Google Scholar]