Abstract
Toxoplasma gondii is a ubiquitous protozoan parasite capable of infecting all warm-blooded animals, including humans. Its closest extant relative, Hammondia hammondi, has never been found to infect humans and, in contrast to T. gondii, is highly attenuated in mice. To better understand the genetic bases for these phenotypic differences, we sequenced the genome of a H. hammondi isolate (HhCatGer041) and found the genomic synteny between H. hammondi and T. gondii to be >95%. We used this genome to determine the H. hammondi primary sequence of two major T. gondii mouse virulence genes, TgROP5 and TgROP18. When we expressed these genes in T. gondii, we found that H. hammondi orthologs of TgROP5 and TgROP18 were functional. Similar to T. gondii, the HhROP5 locus is expanded, and two distinct HhROP5 paralogs increased the virulence of a T. gondii TgROP5 knockout strain. We also identified a 107 base pair promoter region, absent only in type III TgROP18, which is necessary for TgROP18 expression. This result indicates that the ROP18 promoter was active in the most recent common ancestor of these two species and that it was subsequently inactivated in progenitors of the type III lineage. Overall, these data suggest that the virulence differences between these species are not solely due to the functionality of these key virulence factors. This study provides evidence that other mechanisms, such as differences in gene expression or the lack of currently uncharacterized virulence factors, may underlie the phenotypic differences between these species.
Keywords: comparative genomics, gene content versus gene deployment, pathogenesis, host range
Approximately 20% of the US population is infected with the intracellular parasite Toxoplasma gondii (1). T. gondii is a member of the phylum Apicomplexa, which includes multiple human and animal pathogens, including the causative agents of malaria and cryptosporidiosis. A unique feature of this parasite compared with its apicomplexan relatives is the ability to infect, and cause disease in, nearly all warm-blooded animals studied to date, including birds (2). T. gondii is primarily asymptomatic in healthy humans but can cause severe disease in utero, in the immunocompromised, and in healthy adults (3–5). In the laboratory, the mouse model of toxoplasmosis has been extensively studied, a relevant model given that rodents are hosts for T. gondii in both rural and urban environments (6, 7). Highly virulent infections with T. gondii can be initiated by inoculation with fast-growing tachyzoites, slow-growing bradyzoite tissue stages (8), and oocysts (2). Felines are the definitive (i.e., sexual) host for T. gondii (9), and rodent to cat transmission via predation can expand the parasite population by up to 10 million-fold.
The closest extant relative of T. gondii is Hammondia hammondi (10, 11), a fellow apicomplexan that also has its sexual cycle in cats (10). This parasite was once thought to be a strain of T. gondii (12), but it is now accepted as a distinct species (2). Experimentally, a variety of intermediate hosts have been successfully infected with H. hammondi (including rabbits, pigs, rodents, and monkeys), but birds are refractory to infection (ref. 13, and reviewed in ref. 2). In contrast to T. gondii, in both wild-type and IFN-γ knockout mice, parenteral H. hammondi infections with all parasite life stages are nonlethal (2), although oral infections with high numbers of H. hammondi oocysts of some strains can cause disease (2) and mortality (13). In infected mice, H. hammondi tachyzoites and cysts can be found in several tissues, including lymph nodes and spleen (2). Importantly, H. hammondi tissue cysts (bradyzoites) from intermediate hosts are not infective to mice by any route, and therefore horizontal transmission between intermediate hosts is a unique feature of T. gondii (2, 13).
Why some humans become sick from T. gondii infection whereas most remain asymptomatic is largely unknown. Recently, attention has been focused on the genetic basis of parasite virulence, and mice have been used as a model for virulence in T. gondii. Among the virulence determinants, the most potent are rhoptry proteins 18 and 5 (TgROP18 and TgROP5), both members of a large superfamily of proteins secreted from parasite organelles called rhoptries (14–17). Subsequent work has shown that TgROP18 directly phosphorylates immune-related GTPases (IRGs) ,which prevents them from being efficiently loaded onto, and disrupting, the parasitophorous vacuole (PV) (18–24). For TgROP5, complete knockout of this locus in the hypervirulent type I strain renders the parasite completely avirulent in mice (15, 16), and this phenotype is due, at least in part, to ROP5 binding directly to IRG proteins, which presumably then permits TgROP18 IRG phosphorylation and inactivation (23, 24). Based on recent sequence analysis of the Neospora caninum genome [a parasite structurally similar to T. gondii but with a sexual cycle in dogs (25, 26)], it has been suggested that the orthologs of TgROP18 and TgROP5 are not functional as virulence genes in this species. Specifically, ROP18 is a pseudogene in N. caninum (25), and the NcROP5 locus has not significantly expanded in this species as it has in T. gondii, having only 2 copies compared with 4–10 in T. gondii (15, 16, 25). In addition, the N. caninum ROP18 ortholog has a similar promoter structure as the “avirulent” TgROP18 allele in T. gondii strains with low TgROP18 transcript abundance (27).
The genetic bases for the dramatic differences in mouse virulence between T. gondii and H. hammondi are unknown. The goal of the present study was to sequence the genome of H. hammondi strain HhCatGer041 and use it to determine the overall degree of genomic identity and synteny between HhCatGer041 and T. gondii. We then functionally characterized the H. hammondi orthologs of TgROP18 and TgROP5 (HhROP18 and HhROP5) to determine whether they could be linked, or not, to the phenotypic differences between these closely related species.
Results
H. hammondi Genome Has High Synteny and Similarity to That of T. gondii.
Based on previously published analyses of internal transcribed spacer sequences (11, 28), H. hammondi has been categorized as the nearest extant relative of T. gondii, and our rough draft of the H. hammondi genome confirms this similarity. Details about the sequence reads and assembly are in Tables S1and S2. The genome assembled into 56,472 scaffolds, with the largest being 81 kb. When reads were mapped to the v7.2 draft of the T. gondii genome (strain GT1; www.toxodb.org), average coverage for nucleotides with >3× coverage was ∼20×, and 68% of the GT1 nucleotides had ≥4× coverage (Fig. S1). Based on these alignments, the overall divergence genome-wide is 4.9%. The largest region without any sequence coverage was found on the first 175 kb of T. gondii Chr IX (Fig. S1). Despite its incomplete nature, the assembly provides strong evidence for a high degree of genomic synteny between these species. Specifically, strict reciprocal BLAST analysis of all predicted T. gondii coding regions (including introns) on 87 contigs >20 kb in length shows that 298 of 311 genes (∼96%) are in the same order and orientation as they are in T. gondii. Importantly, only 96 genes (out of 8,993 queried) had no significant reciprocal blast hits (E < 1e−5; Dataset S1). An identical comparison with N. caninum genes shows that 818 out of 7,226 predicted genes have no reciprocal blast hits [although more robust estimates of shared gene content between T. gondii and N. caninum have been published by others (25)]. Whereas the number of genes “missing” in H. hammondi compared with T. gondii will change as better assemblies of the H. hammondi genome emerge, this rough draft further establishes H. hammondi as the closest extant relative of T. gondii and sets the stage for gene-by-gene comparisons to better understand the evolution of virulence in T. gondii.
ROP5 Locus Is Expanded, Diversified, and Functionally Conserved In H. hammondi.
Recently we and others demonstrated a clear role for the TgROP5 locus in T. gondii virulence (15, 16). This locus was characterized by the presence of multiple paralogs that show clear evidence for selection-driven diversification (15). Using the H. hammondi genome sequence, we examined sequence read coverage on scaffolds with H. hammondi ROP5 sequence (Fig. 1A), and this analysis suggested that the ROP5 locus is also expanded in H. hammondi, with an estimated copy number of 8–10. To determine whether the locus had also diversified, we PCR-amplified and sequenced eight HhROP5 paralogs and identified eight distinct protein coding sequences (Fig. 1B, Fig. S2A), confirming that this locus is expanded and diversified in H. hammondi as it is in T. gondii (all primers used in this study are listed in Table S3). This finding is in contrast to what has been reported in N. caninum (25) although the role of NcROP5 in mouse virulence has not been directly tested. All eight isoforms harbored the same sequence features and domains as TgROP5 orthologs, including predicted signal sequences, an amphipathic helical domain in the N terminus that mediates interaction with the PV in TgROP5 (29), and a clear pseudokinase domain (Fig. S2A). Selection analyses also revealed that these individual copies showed strong evidence for selection. For example, isoform 2–4 has a non-synonymous/synonymous substitution ratio (dN/dS) of >2.8 compared with isoforms 1–3, 1–4, and 1–5, suggesting not only that the HhROP5 locus has expanded in H. hammondi but also that it has been under selection.
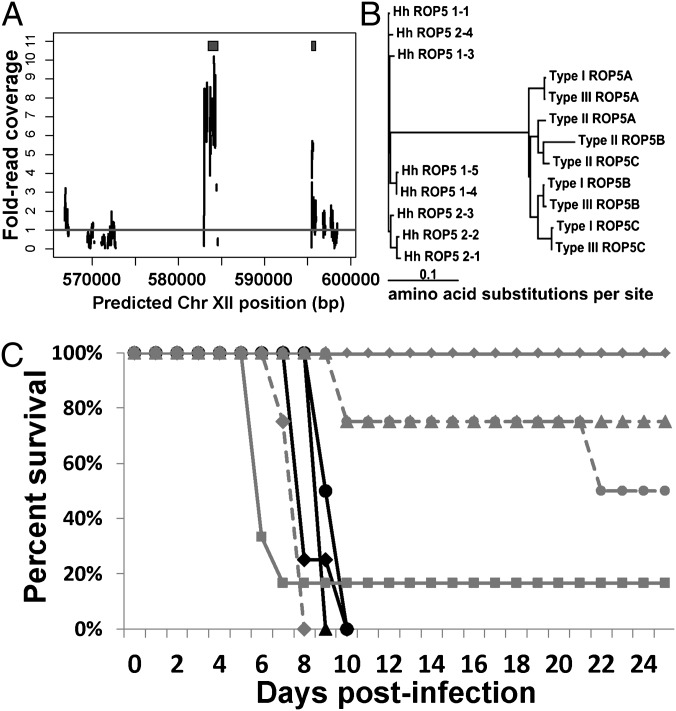
Fig. 1.
Expansion, diversification, and functionality of H. hammondi ROP5. (A) Sequence coverage plot for three H. hammondi genomic scaffolds arranged based on their putative location on H. hammondi Chr XII (determined by Nucmer alignment of H. hammondi scaffolds to the v7.2 GT1 genome). Black blocks above the plots indicate the location of significant (expect < 1 × 10−10) TBLASTN hits of TgROP5B against the H. hammondi genome. (B) Phylogram of amino acid sequences of eight HhROP5 paralogs and known TgROP5 paralogs. (C) Mice were infected with RH:WT (black lines) or RH∆ROP5 complemented with a single copy of either HhROP51–1 (gray solid lines) or HhROP52–1 (gray dashed lines). Four mice were used for each strain and dose with the exception of RH∆ROP5:HhROP51–1 at 106 (n = 6), and RH:WT at 104 (n = 3). Tachyzoite dose symbol key: ■, 106; ◆, 105; ▲, 104; ●, 103.
Based on our sequence analyses of HhROP5, we chose two paralogs, 1–1 and 2–1 (Fig. 1B, Fig. S2A), for complementation into a type I T. gondii strain lacking the entire TgROP5 locus [RH∆TgROP5 (15)]. In contrast to the parental strain (RH), which has an effective LD100 of a single parasite (30), this strain does not cause mortality at doses as high as 106 tachyzoites (15, 16). When expressed in T. gondii, HhROP5 paralogs 1–1 and 2–1 are both effectively trafficked to the T. gondii rhoptries (Fig. S3B) and can be found associated with the PV (Fig. S3E) as expected. Fusing 713 and 702 bp of upstream sequence from HhROP51–1 and HhROP52–1, respectively (Figs. S2B and S3A), with luciferase is capable of driving reporter expression at levels similar to one another and to a homologous upstream sequence for T. gondii type II ROP5B (Fig. S3A). In multiple experiments, it was found that the promoter activity of HhROP52–1 is slightly lower than HhROP51–1. Anti-HA Western blotting on parasites expressing these paralogs is consistent with this slight difference in promoter activity: normalized protein levels for HhROP52–1 are slightly lower than for HhROP51–1 (Fig. S3 C and D). To determine whether these HhROP5 orthologs were functionally conserved as virulence genes in H. hammondi, we infected mice with two genetically distinct clone sets of RH∆ROP5:ROP5Hh1–1 and RH∆ROP5:ROP5Hh2–1, along with RH∆ROP5 and wild-type RH (Fig. S4A). For clone set 1, RH∆ROP5:ROP5Hh2–1 was 100% lethal at a dose of 105, and 50% lethal at a dose of 103 (Fig. 1C). In contrast, RH∆ROP5:ROP5Hh1–1 was significantly less virulent than RH∆ROP5:ROP5Hh2–1, causing 80% mortality at a dose of 106 but none at lower doses (Fig. 1C). However, mice infected with RH∆ROP5:ROP5Hh1–1 did show signs of morbidity, in that mice infected with all doses (102 to 106) lost weight over the course of the infection (Fig. S4B). When a genetically distinct clone set was used to infect mice at doses of 105 and 104 tachyzoites, we observed similar differences in the efficacy of the HhROP5 alleles (with the 2–1 allele being more effective at increasing the virulence of RH∆ROP5 than the 1–1 allele) although in these experiments both alleles conferred some lethality at doses as low as 104 tachyzoites (Fig. S4C). Regardless of the reasons for the differences in the results of these experiments with distinct clones, in both experiments, both the crude LD50 as well as the time to death at each dose was always shorter in mice infected with RH∆ROP5:ROP5Hh2–1 compared with RH∆ROP5:ROP5Hh1–1. Overall, these data show that ROP5Hh2–1 and ROP5Hh1–1 are functional virulence genes that are not equivalent in terms of their effects on mouse virulence when heterologously expressed in T. gondii.
H. hammondi ROP18 Upstream Sequence Contains a 107-bp Core Promoter That Is Found Only in Active TgROP18 Alleles.
The H. hammondi ROP18 gene is structurally most similar to the avirulent type III TgROP18, both in its promoter and coding region (Fig. S5A). It has a promoter similar in length to type III TgROP18, whose promoter is inactive and four-to-five times longer than T. gondii type I and II ROP18 promoters (27, 31). This difference in promoter structure is due to a 2.1-kb insertion in the type III gene relative to types I and II, and a corresponding 201-bp deletion (27, 31). Surprisingly, when we placed the H. hammondi ROP18 promoter sequence upstream of luciferase, we found that it is highly active in these assays, in direct contrast to the corresponding upstream sequence from type III TgROP18 (Fig. 2B). This surprising result indicates that the 2.1-kb insertion itself is not the cause of reduced transcription of TgROP18, but rather some other promoter element. We have identified this element, which is a 107-bp deletion in the type III promoter compared with the HhROP18 promoter (Fig. 2A). This region is present in both the type I and type II TgROP18 alleles, and H. hammondi, just upstream of the predicted transcriptional start site (Fig. 2A). When this 107-bp region is deleted in the HhROP18 promoter, it completely inactivates its activity in luciferase reporter assays (Fig. 2B, Left; raw values shown in Table S4). Additionally, when this 107-bp fragment is inserted into the type III strain TgROP18 promoter, the presence of this sequence dramatically increases the activity of this promoter in reporter assays, increasing its activity by over eightfold compared with the type II TgROP18 promoter (Fig. 2B, Right). Therefore, we conclude that this small upstream sequence plays a key role in HhROP18 expression in H. hammondi and suggests that it is similarly important in type I and type II strains of T. gondii.
Fig. 2.
A 107-bp region of the H. hammondi ROP18 promoter is necessary for gene expression. (A) Alignment of the HhROP18 and TgROP18III promoters just upstream of the transcriptional start site (gray arrow). Numbers represent nucleotide position relative to the start codon. (B) The HhROP18 upstream sequence had promoter activity that was similar to TgROP18II but significantly greater than TgROP18III. Deletion of the 107-bp region present in HhROP18 and types I and II T. gondii dramatically reduced promoter activity when assayed in T. gondii. Data for two luciferase assays are shown as the ratios of firefly luciferase signal to Renilla luciferase signal (Left). When this 107-bp fragment was introduced into the TgROP18 type III upstream sequence, it dramatically increased the normalized luciferase signal compared with the type II and type III ROP18 promoter sequences (Right). (C) Anti-HA immunofluorescence showing the localization of TgROP18II, TgROP18III, and HhROP18 and lack of staining in the strains harboring TgROP18III. (Scale bar: 10 µm.)
H. hammondi ROP18 Is a Highly Functional Virulence Gene in T. gondii.
To see whether H. hammondi ROP18 is functional as a virulence gene, we complemented a type III strain with HhROP18 or TgROP18 from each of the three major T. gondii lineages (types I, II and III). As expected, HA-tagged TgROP18II and HhROP18 (Fig. 2C) all trafficked to the T. gondii rhoptries, and HA-tagged HhROP18 could also be found on the parasitophorous vacuolar membrane (PVM) (Fig. S5D). In contrast, and consistent with our promoter activity experiments (Fig. 2B), there was no visible HA staining in TgROP18III-transfected parasites, despite the fact that the construct was present (Fig. S5E). To compare the impact of expression of these 4 ROP18 alleles on parasite virulence, mice were infected intraperitoneally with 1,000 tachyzoites of III:EV (empty vector), III:TgROP18I, III:TgROP18II, III:TgROP18III, and III:HhROP18, and parasite burden was assessed daily using in vivo bioluminescence imaging (BLI). As expected, mice infected with III:TgROP18III and III:EV did not succumb to the infection and showed comparatively low levels of parasite burden over the course of the infection (Fig. 3 A–C). All mice infected with III:TgROP18I and III:TgROP18II died by day 16 (Fig. 3B). Remarkably, >80% of mice infected with III:HhROP18 died before mice infected with any of the other strains, including those infected with III:TgROP18I (Fig. 3B). This difference in time to mortality is consistent with the in vivo BLI data. Specifically, III:HhROP18 increased in number in infected mice much more rapidly than all other strains, being over 10-fold higher than all other strains on days 6 and 7 postinfection (Fig. 3A). These data show that, despite being derived from a parasite species that is attenuated in mice, the H. hammondi ROP18 gene, when expressed in T. gondii, may actually be a more effective virulence gene than the alleles from T. gondii types I and II. Based on quantitative Western blotting, HhROP18 is expressed at similar levels as TgROP18I (Fig. S5 B and C), suggesting that the HhROP18 allele, rather than differences in expression level, are responsible for the increased impact of HhROP18 on virulence compared with the T. gondii ROP18 alleles from type I and type II strains.
Fig. 3.
Expressing HhROP18 in a TgType III background increases parasite lethality and in vivo proliferation. Mice were infected with 103 parasites of a type III strain complemented with TgROP18I, TgROP18II, TgROP18III, HhROP18, or empty vector (n = 3, 8, 5, 7, and 5, respectively). Results shown are pooled from two experiments. (A) Quantitative in vivo BLI of infected mice. Asterisks indicate significantly higher luciferase signal in HhROP18-complemented parasites compared with TgROP18I-complemented parasites (*P < 0.05; **P < 0.01). (B) Mouse mortality. (C) Representative images of parasite burden in mice on day 6 postinfection.
Discussion
The differences in virulence between H. hammondi and T. gondii are belied by their close phylogenetic relationship. In contrast to T. gondii, when oocysts are injected subcutaneously or intraperitoneally, H. hammondi is completely avirulent in wild-type and IFN-γ knockout mice (2) and has never been found to infect humans. A number of mechanisms could explain the dramatic virulence differences between two species that share such a recent common ancestry. One is that virulence is driven by genes that are uniquely present and/or divergent in T. gondii compared with H. hammondi. Another is that both species harbor the same arsenal of effectors, but they are differentially regulated by distinct transcriptional mechanisms or transcription factors. However, another is that the H. hammondi genome contains “avirulence” genes that are only present and/or expressed in H. hammondi. Genome sequence comparison between these organisms is a crucial first step in addressing these hypotheses and to ultimately determine how mouse virulence, as well as other traits, evolved uniquely in T. gondii.
In the present study, we found that H. hammondi orthologs of ROP5 and ROP18, which are the most potent mouse virulence factors known in T. gondii (14–17), are functional when expressed heterologously in T. gondii. As in T. gondii, the HhROP5 locus has expanded, and individual paralogs appear to be under diversifying selection in H. hammondi. Specifically, H. hammondi ROP52–1 increases the virulence of a T. gondii ROP5 knockout strain 100-fold, and this effect is more potent than complementation of the same ROP5 knockout strain with a single copy of T. gondii ROP5 [the ROP5 “A”isoform (15)]. Whereas RHΔTgROP5:TgROP5A was lethal to only 20% at a dose of 105 tachzyoites, we observed 100% mortality with this dose of RHΔTgROP5:ROP5Hh2–1, and 50% mortality at a dose of 1,000 RHΔTgROP5:Hh2–1 tachyzoites. Interestingly, in terms of mouse mortality, HhROP51–1 was less potent than HhROP52–1. Overall these data indicate that the ROP5 locus in H. hammondi bears all of the hallmarks of the orthologous locus from T. gondii: (i) it contains multiple copies that have undergone selection-driven diversification in sequence, (ii) individual copies can increase the pathogenesis of a T. gondii ROP5 knockout, and (iii) individual copies vary in terms of their effects on virulence. Interestingly, across the H. hammondi ROP5 paralogs, the most polymorphic region is found within the pseudokinase domain in residues 490–501, and a similar level of polymorphism is found in this same region across the T. gondii ROP5 paralogs (Fig. S2A) (15). Future work on the other six H. hammondi ROP5 paralogs will provide more information on the impact of locus diversification in this species. Moreover, we have not yet tested the impact of expressing multiple distinct copies of H. hammondi ROP5 orthologs, nor the impact of H. hammondi ROP5 expression on the loading of IRG GTPases (23, 24). These studies should help to inform current models of TgROP5 structure-activity relationships in T. gondii.
In T. gondii, virulence gene expression, rather than individual amino acid polymorphisms, has been linked to differences in virulence between strains. Specifically, TgROP18 transcript abundance in type III strains is significantly lower than in members of the type I and II lineages, and, within the T. gondii species, this lower transcript abundance is clearly due to a 2.1-kb sequence inserted 85 bp upstream of the ROP18 start codon compared to strain types I and II (27, 31). We found that a similarly homologous insertion was present in the H. hammondi ROP18 upstream sequence. Given the presence of this insertion, we were surprised to find that, in contrast to the corresponding sequence from T. gondii type III strains, the H. hammondi ROP18 promoter drove significantly high levels of luciferase in promoter assays conducted in T. gondii. We identified a 107-bp region in H. hammondi and type I and II T. gondii ROP18 upstream of the predicted transcriptional start site that is not present in T. gondii type III ROP18. Deletion of this sequence in H. hammondi ROP18 destroys promoter activity, suggesting that this sequence contains a core ROP18 promoter or enhancer element in both species (at least when heterologously expressed in T. gondii). Even more exciting, inserting this H. hammondi sequence into the comparatively ineffective type III promoter at the proper location effectively resurrects this promoter to levels that are even higher than the TgROP18II promoter. Importantly, based on BLASTN, this 107-bp sequence cannot be identified in the entire N. caninum genome (expect >10), suggesting that the ROP18 promoter became “active” after the split between the N. caninum and T. gondii/H. hammondi lineages. This increased promoter activity has significant phenotypic consequence: HhROP18 appears to be at least as potent as, and possibly MORE potent than, TgROP18I.
Through a preliminary sequence analysis of an isolate of H. hammondi (HhCatGer041), we have found that orthologs of key T. gondii mouse virulence genes are functionally conserved in this species. Whereas virulence commonly evolves in pathogenic bacteria by changes in gene content via acquisition of so-called pathogenicity islands [e.g., in pathogenic Escherichia coli (32)], in the case of the eukaryotic pathogens T. gondii and H. hammondi, at least, gene content and functionality may not be sufficient to explain their phenotypic differences. One hypothesis is that the inability of H. hammondi to infect and be transmitted by mice compared with T. gondii is due to differences in gene deployment. This explanation for the phenotypic differences between T. gondii and H. hammondi would be somewhat analogous to what has been found in transcriptome comparisons between free-living and “parasitic” forms of Listeria monocytogenes where highly distinct transcriptional profiles, driven by activation/deactivation of key transcription factors, are the hallmark of the pathogenic lifestyle (33). High quality data from the H. hammondi transcriptome will be necessary to address this hypothesis. Transcriptome analyses in H. hammondi are made more difficult by the fact that H. hammondi does not replicate for more than 5 d in cell culture and therefore must be generated directly from cat-derived oocyst stages. However, the depth of next generation sequencing provides a potential solution to this problem. These analyses will be facilitated by the crude H. hammondi genome draft presented here as well as the recently released draft of a different isolate (H.H.34) by the J. Craig Venter Institute (http://jcvi.org).
Methods
Parasite Strains and Maintenance.
T. gondii parasites were allowed to invade monolayers of human foreskin fibroblasts (HFFs) that were grown at 37 °C in 5% CO2. HFFs were maintained in Dulbecco’s modified Eagle’s medium with 10% (vol/vol) FCS, 2 mM glutamine, and 50 μg/mL each of penicillin and streptomycin (cDMEM). RH, ME49, and CEP were used as representative type I, II, and III strains, respectively. The H. hammondi strain (HhCatGer041) was isolated from cat feces on June 29, 2009 during a survey of Toxoplasma gondii in Germany. The isolate was confirmed as Hammondia hammondi as previously described (34) using specific Hammondia hammondi primers (35) and by the complete nonvirulence of the parasite in IFN-γ knockout mice. For H. hammondi oocyst production, IFN-γ knockout mice were fed 104 H. hammondi oocysts and killed 59 d later. Muscles from infected mice were fed to 10- to 20-wk-old cats, and feces were collected during days 5–11 postinfection. Unsporulated oocysts were isolated by sucrose floatation and allowed to sporulate at room temperature in 2% sulfuric acid (2).
DNA Isolation from H. hammondi Oocysts.
Sporulated oocysts (40-80 million) were washed four times in Hanks’ balanced salt solution (HBSS) and treated with 10% bleach in PBS for 30 min. Washed pellets were resuspended in 4 mL of HBSS in a 15-mL falcon tube, and 1 g of sterile glass beads (710-1,180 μM; Sigma) were added. Parasites were vortexed on high speed for 30 s, cooled for 30 s, and then vortexed for 30 s more. The supernatant was removed and pelleted by centrifugation. DNA was isolated from this preparation, which contained sporocysts that had been freed from the oocyst using the DNAzol reagent according to the manufacturer (Life Technologies). For two different preparations, 8 ng of DNA was linearly amplified using the GenomiPhi DNA Amplification Kit (GE Healthcare Life Sciences) and ethanol precipitated. These preparations were used for Illumina and PacBio RS (Pacific Biosciences) library preparation and sequencing and for PCR amplification (described in Gene Identification, Sequencing, and Generation of Transgenic T. gondii).
Sequencing, de Novo Assembly, and Analysis of the H. hammondi Genome.
For Illumina sequencing, two different preparations of linearly amplified H. hammondi DNA were combined, and a single library (insert size ∼200 bp) was made from the preparation using the DNA TrueSeq kit. Thirty million 36-bp paired-end reads were generated in a single lane of an Illumina Genome Analyzer IIx at the Tufts Genomics Core Facility. Mean insert size and SD were calculated via de novo assembly in Ray (36). Two additional linearly amplified DNA samples were combined and submitted to the Duke Genomics Core Facility, and sequence data from four single molecule, real-time (SMRT) cells were obtained from a 2-kb insert library on a PacBio RS system. Consensus PacBio RS reads were assembled along with the Illumina reads using Ray with the following settings: K-mer size of 21, a mean insert size of 206, and a SD of 33. The K-mer size was determined iteratively by running the assembly using K-mer sizes ranging from 12 to 23 to maximize the N50 value. Synteny analyses were performed by reciprocal BLASTN between the genomic DNA coding regions for all v7.2 ME49 predicted genes versus the H. hammondi genomic contigs with an expected value of 1e−5. Genes on contigs larger than 20 kb were then scored manually for reciprocal hit presence, order, and orientation. Raw Illumina sequence reads were aligned to the v7.2 GT1 T. gondii genome using Bowtie2 (http://bowtie-bio.sourceforge.net/bowtie2/index.shtml) using the following parameters: bowtie2 –local -L 10 -k 10 -i S,1,0.5. Coverage plots were generated using Samtools mpileup on the bowtie2 output, and coverage was determined at each base using a custom perl script.
Gene Identification, Sequencing, and Generation of Transgenic T. gondii.
H. hammondi contigs with ROP5 and ROP18 coding and upstream sequences were identified using TBLASTN with T. gondii ROP5 and ROP18 protein sequences. HhROP5 isoforms were PCR amplified using short-cycle, long-extension PCR to amplify ∼1,000 bp upstream of the predicted start codon to the predicted stop codon (Invitrogen). Eight distinct clones were sequenced and protein sequences aligned using CLUSTALW (37). Two paralogs (1-1 and 2-1) were amplified directly from the cloning plasmid (PCR2.1 TOPO) using primers with AscI (Forward) and XmaI (Reverse) sites and cloned into the pUPRT-HA vector (15) in frame with an HA tag. Selection for plasmid integration into RHΔTgROP5 was performed using 5-fluorodeoxyuridine (38), and resistant parasites were cloned by limiting dilution. Sequences for Hh, TgI, TgII, and TgIII ROP18 were directionally cloned into pENTR-D-Topo (Invitrogen) and recombined into a derivative of the pGRA-HA-HPT vector (14) in which the Gra1 upstream sequence was replaced with the attR1-ccdB-attR2 cassette for Gateway recombination cloning (Invitrogen) (39, 40). The constructs were transfected into a click beetle luciferase-expressing T. gondii strain null for HXGPRT (CEP) (14). Parasites were selected for resistance to mycophenolic acid and xanthine and cloned by limiting dilution (41). PCR was used to verify the presence of the full construct in III:TgROP18III parasites. All clones were validated using anti-HA immunofluorescence and Western blotting. Primers are listed in Table S3.
Supplementary Material
Acknowledgments
We thank Michael Reese for helpful comments on the manuscript. This work was supported by a Pew Scholarship in the Biomedical Sciences and by National Institute of Allergy and Infectious Diseases K22 Research Scholar Award 5K22AI080977 (to J.P.B.), and by Howard Hughes Medical Institute-sponsored undergraduate research fellowships (to K.A.W. and R.A.D.). D.C.H. and G.S. were supported by German Federal Ministry of Education and Research Grants 01 Kl 0765, Toxonet01 and 01 KI 1002 F, Toxonet02.
Footnotes
The authors declare no conflict of interest.
Data deposition: The HhCatGer041 genomic assembly and raw sequence reads have been deposited in GenBank under BioProject PRJNA191036 and BioSample 1974510.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1304322110/-/DCSupplemental.
References
- 1.Jones JL, et al. Toxoplasma gondii infection in the United States: Seroprevalence and risk factors. Am J Epidemiol. 2001;154(4):357–365. doi: 10.1093/aje/154.4.357. [DOI] [PubMed] [Google Scholar]
- 2.Dubey JP, Sreekumar C. Redescription of Hammondia hammondi and its differentiation from Toxoplasma gondii. Int J Parasitol. 2003;33(13):1437–1453. doi: 10.1016/s0020-7519(03)00141-3. [DOI] [PubMed] [Google Scholar]
- 3.Luft BJ, et al. Toxoplasmic encephalitis in patients with the acquired immunodeficiency syndrome. N Engl J Med. 1993;329(14):995–1000. doi: 10.1056/NEJM199309303291403. [DOI] [PubMed] [Google Scholar]
- 4.Remington JS, Klein JO. Infectious Diseases of the Fetus and Newborn Infant. 5th Ed. Philadelphia: Saunders; 2001. [Google Scholar]
- 5.Carme B, Demar M, Ajzenberg D, Dardé ML. Severe acquired toxoplasmosis caused by wild cycle of Toxoplasma gondii, French Guiana. Emerg Infect Dis. 2009;15(4):656–658. doi: 10.3201/eid1504.081306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kijlstra A, et al. The role of rodents and shrews in the transmission of Toxoplasma gondii to pigs. Vet Parasitol. 2008;156(3-4):183–190. doi: 10.1016/j.vetpar.2008.05.030. [DOI] [PubMed] [Google Scholar]
- 7.Murphy RG, et al. The urban house mouse (Mus domesticus) as a reservoir of infection for the human parasite Toxoplasma gondii: An unrecognised public health issue? Int J Environ Health Res. 2008;18(3):177–185. doi: 10.1080/09603120701540856. [DOI] [PubMed] [Google Scholar]
- 8.Boyle JP, Saeij JP, Boothroyd JC. Toxoplasma gondii: Inconsistent dissemination patterns following oral infection in mice. Exp Parasitol. 2007;116(3):302–305. doi: 10.1016/j.exppara.2007.01.010. [DOI] [PubMed] [Google Scholar]
- 9.Frenkel JK, Dubey JP, Miller NL. Toxoplasma gondii in cats: Fecal stages identified as coccidian oocysts. Science. 1970;167(3919):893–896. doi: 10.1126/science.167.3919.893. [DOI] [PubMed] [Google Scholar]
- 10.Frenkel JK, Dubey JP. Hammondia hammondi: A new coccidium of cats producing cysts in muscle of other mammals. Science. 1975;189(4198):222–224. doi: 10.1126/science.806116. [DOI] [PubMed] [Google Scholar]
- 11.Su C, et al. Recent expansion of Toxoplasma through enhanced oral transmission. Science. 2003;299(5605):414–416. doi: 10.1126/science.1078035. [DOI] [PubMed] [Google Scholar]
- 12.Heydorn AO, Mehlhorn H. Further remarks on Hammondia hammondi and the taxonomic importance of obligate heteroxeny. Parasitol Res. 2001;87(7):573–577. doi: 10.1007/s004360100381. [DOI] [PubMed] [Google Scholar]
- 13.Frenkel JK, Dubey JP. Hammondia hammondi gen. nov., sp.nov., from domestic cats, a new coccidian related to Toxoplasma and Sarcocystis. Z Parasitenkd. 1975;46(1):3–12. doi: 10.1007/BF00383662. [DOI] [PubMed] [Google Scholar]
- 14.Saeij JP, et al. Polymorphic secreted kinases are key virulence factors in toxoplasmosis. Science. 2006;314(5806):1780–1783. doi: 10.1126/science.1133690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Reese ML, Zeiner GM, Saeij JP, Boothroyd JC, Boyle JP. Polymorphic family of injected pseudokinases is paramount in Toxoplasma virulence. Proc Natl Acad Sci USA. 2011;108(23):9625–9630. doi: 10.1073/pnas.1015980108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Behnke MS, et al. Virulence differences in Toxoplasma mediated by amplification of a family of polymorphic pseudokinases. Proc Natl Acad Sci USA. 2011;108(23):9631–9636. doi: 10.1073/pnas.1015338108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Taylor S, et al. A secreted serine-threonine kinase determines virulence in the eukaryotic pathogen Toxoplasma gondii. Science. 2006;314(5806):1776–1780. doi: 10.1126/science.1133643. [DOI] [PubMed] [Google Scholar]
- 18.Pawlowski N, et al. The activation mechanism of Irga6, an interferon-inducible GTPase contributing to mouse resistance against Toxoplasma gondii. BMC Biol. 2011;9:7. doi: 10.1186/1741-7007-9-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Steinfeldt T, et al. Phosphorylation of mouse immunity-related GTPase (IRG) resistance proteins is an evasion strategy for virulent Toxoplasma gondii. PLoS Biol. 2010;8(12):e1000576. doi: 10.1371/journal.pbio.1000576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Khaminets A, et al. Coordinated loading of IRG resistance GTPases on to the Toxoplasma gondii parasitophorous vacuole. Cell Microbiol. 2010;12(7):939–961. doi: 10.1111/j.1462-5822.2010.01443.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Zhao YO, Khaminets A, Hunn JP, Howard JC. Disruption of the Toxoplasma gondii parasitophorous vacuole by IFNgamma-inducible immunity-related GTPases (IRG proteins) triggers necrotic cell death. PLoS Pathog. 2009;5(2):e1000288. doi: 10.1371/journal.ppat.1000288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Martens S, et al. Disruption of Toxoplasma gondii parasitophorous vacuoles by the mouse p47-resistance GTPases. PLoS Pathog. 2005;1(3):e24. doi: 10.1371/journal.ppat.0010024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Niedelman W, et al. The rhoptry proteins ROP18 and ROP5 mediate Toxoplasma gondii evasion of the murine, but not the human, interferon-gamma response. PLoS Pathog. 2012;8(6):e1002784. doi: 10.1371/journal.ppat.1002784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fleckenstein MC, et al. A Toxoplasma gondii pseudokinase inhibits host IRG resistance proteins. PLoS Biol. 2012;10(7):e1001358. doi: 10.1371/journal.pbio.1001358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Reid AJ, et al. Comparative genomics of the apicomplexan parasites Toxoplasma gondii and Neospora caninum: Coccidia differing in host range and transmission strategy. PLoS Pathog. 2012;8(3):e1002567. doi: 10.1371/journal.ppat.1002567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Collantes-Fernandez E, et al. Infected dendritic cells facilitate systemic dissemination and transplacental passage of the obligate intracellular parasite Neospora caninum in mice. PLoS ONE. 2012;7(3):e32123. doi: 10.1371/journal.pone.0032123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Khan A, Taylor S, Ajioka JW, Rosenthal BM, Sibley LD. Selection at a single locus leads to widespread expansion of Toxoplasma gondii lineages that are virulent in mice. PLoS Genet. 2009;5(3):e1000404. doi: 10.1371/journal.pgen.1000404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Ellis JT, et al. The genus Hammondia is paraphyletic. Parasitology. 1999;118(Pt 4):357–362. doi: 10.1017/s0031182098003801. [DOI] [PubMed] [Google Scholar]
- 29.Reese ML, Boothroyd JC. A helical membrane-binding domain targets the Toxoplasma ROP2 family to the parasitophorous vacuole. Traffic. 2009;10(10):1458–1470. doi: 10.1111/j.1600-0854.2009.00958.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Sibley LD, Boothroyd JC. Virulent strains of Toxoplasma gondii comprise a single clonal lineage. Nature. 1992;359(6390):82–85. doi: 10.1038/359082a0. [DOI] [PubMed] [Google Scholar]
- 31.Boyle JP, Saeij JP, Harada SY, Ajioka JW, Boothroyd JC. Expression quantitative trait locus mapping of toxoplasma genes reveals multiple mechanisms for strain-specific differences in gene expression. Eukaryot Cell. 2008;7(8):1403–1414. doi: 10.1128/EC.00073-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Perna NT, et al. Genome sequence of enterohaemorrhagic Escherichia coli O157:H7. Nature. 2001;409(6819):529–533. doi: 10.1038/35054089. [DOI] [PubMed] [Google Scholar]
- 33.Toledo-Arana A, et al. The Listeria transcriptional landscape from saprophytism to virulence. Nature. 2009;459(7249):950–956. doi: 10.1038/nature08080. [DOI] [PubMed] [Google Scholar]
- 34.Herrmann DC, et al. Atypical Toxoplasma gondii genotypes identified in oocysts shed by cats in Germany. Int J Parasitol. 2010;40(3):285–292. doi: 10.1016/j.ijpara.2009.08.001. [DOI] [PubMed] [Google Scholar]
- 35.Schares G, et al. Characterization of a repetitive DNA fragment in Hammondia hammondi and its utility for the specific differentiation of H. hammondi from Toxoplasma gondii by PCR. Mol Cell Probes. 2008;22(4):244–251. doi: 10.1016/j.mcp.2008.04.003. [DOI] [PubMed] [Google Scholar]
- 36.Boisvert S, Laviolette F, Corbeil J. Ray: Simultaneous assembly of reads from a mix of high-throughput sequencing technologies. J Comput Biol. 2010;17(11):1519–1533. doi: 10.1089/cmb.2009.0238. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Larkin MA, et al. Clustal W and Clustal X version 2.0. Bioinformatics. 2007;23(21):2947–2948. doi: 10.1093/bioinformatics/btm404. [DOI] [PubMed] [Google Scholar]
- 38.Donald RG, Roos DS. Insertional mutagenesis and marker rescue in a protozoan parasite: Cloning of the uracil phosphoribosyltransferase locus from Toxoplasma gondii. Proc Natl Acad Sci USA. 1995;92(12):5749–5753. doi: 10.1073/pnas.92.12.5749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rosowski EE, et al. Strain-specific activation of the NF-kappaB pathway by GRA15, a novel Toxoplasma gondii dense granule protein. J Exp Med. 2011;208(1):195–212. doi: 10.1084/jem.20100717. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Hartley JL, Temple GF, Brasch MA. DNA cloning using in vitro site-specific recombination. Genome Res. 2000;10(11):1788–1795. doi: 10.1101/gr.143000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Donald RG, Carter D, Ullman B, Roos DS. Insertional tagging, cloning, and expression of the Toxoplasma gondii hypoxanthine-xanthine-guanine phosphoribosyltransferase gene. Use as a selectable marker for stable transformation. J Biol Chem. 1996;271(24):14010–14019. doi: 10.1074/jbc.271.24.14010. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.