Abstract
Natural populations throughout the tree of life undergo range expansions in response to changes in the environment. Recent theoretical work suggests that range expansions can have a strong effect on evolution, even leading to the fixation of deleterious alleles that would normally be outcompeted in the absence of migration. However, little is known about how range expansions might influence alleles under frequency- or density-dependent selection. Moreover, there is very little experimental evidence to complement existing theory, since expanding populations are difficult to study in the natural environment. In this study, we have used a yeast experimental system to explore the effect of range expansions on the maintenance of cooperative behaviors, which commonly display frequency- and density-dependent selection and are widespread in nature. We found that range expansions favor the maintenance of cooperation in two ways: (i) through the enrichment of cooperators at the front of the expanding population and (ii) by allowing cooperators to “outrun” an invading wave of defectors. In this system, cooperation is enhanced through the coupling of population ecology and evolutionary dynamics in expanding populations, thus providing experimental evidence for a unique mechanism through which cooperative behaviors could be maintained in nature.
Keywords: microbes, Allee effect, eco-evolutionary feedback
Natural populations often increase the geographical area that they occupy through population growth and dispersal into new territory (1–4). Such events, termed range expansions, occur repeatedly during the life history of a species in response to changes in the environment (for instance, forest fires or seasonal changes in climate) as well as changes in phenotype that allow the species to colonize regions that were previously inaccessible to them (1, 5). Despite their widespread occurrence, the effect of range expansions on genetic diversity has only begun to be explored. Recent theoretical analyses suggest that genetic drift on the low-density front of an expanding population can lead to the fixation of neutral (or even deleterious) alleles in a way that mimics positive selection (6, 7). This effect, known as “allele surfing,” is a stochastic effect that has been demonstrated experimentally for neutral alleles in expanding bacterial colonies (8) and was recently suggested to underlie the global patterns of human phenotypic variation observed today (9–11).
Here, we chose to study the maintenance of cooperative alleles in the context of populations undergoing range expansions. By definition, a “cooperator” provides a benefit to other members of the population at a cost to itself (12). Examples of cooperation are ubiquitous in the wild, ranging from siderophore production in bacteria to the formation of communities in human populations (12–15). However, given the appearance of “defectors” that exploit the benefit provided by the cooperators without incurring the cost, explaining the origin and maintenance of cooperation in nature remains a key challenge in evolutionary biology (12, 14, 16, 17). Several studies (both experimental and theoretical) have suggested that spatial structuring may play a key role in the maintenance of cooperation in nature (18–21). Moreover, previous studies have shown that population dynamics can play a key role in maintaining cooperative alleles (22, 23). Thus, we hypothesized that the population dynamics of expanding populations might support the maintenance of cooperation in nature.
More broadly, cooperative alleles also comprise some of the most well-known examples of alleles for which selection is not constant, but instead depends upon instantaneous allele frequencies or population densities (24–30). Whereas range expansions are predicted to have a strong effect on the evolution of neutral, deleterious, and beneficial alleles, alleles under frequency- or density-dependent selection have not been considered previously. However, because range expansions often create spatial heterogeneity in both population density and allele frequency (1, 8), we hypothesized that, in contrast to stochastic allele surfing events, there might be a deterministic coupling between population and evolutionary dynamics that would be broadly generalizable to alleles under frequency- or density-dependent selection.
To probe how range expansions shape the maintenance of cooperation, we used a well-characterized budding yeast model system. In this system, a cooperator strain (expressing SUC2) catalyzes the enzymatic conversion of sucrose into a consumable energy source (glucose and fructose) (16, 24, 31, 32). Although the cooperators retain some preferential access to the sugars that they produce, roughly 99% of this resource is lost to neighboring cells (16, 24, 31). In our system, this behavior facilitates the growth of an obligate defector strain (Δsuc2) that cannot degrade sucrose but can consume the sugars produced by neighboring cells (16, 24, 31, 32). This system shows negative frequency-dependent selection, which leads to coexistence between the cooperator and defector genotypes in our system (16, 24, 31–34) (for a comprehensive study the eco-evolutionary feedback inherent in these cooperator–defector dynamics see ref. 35). The cooperator strain also displays an Allee effect (where the per capita growth rate of the population is maximal at intermediate population densities). This type of growth is predicted to have a strong effect on the dynamics of expanding populations (36–39).
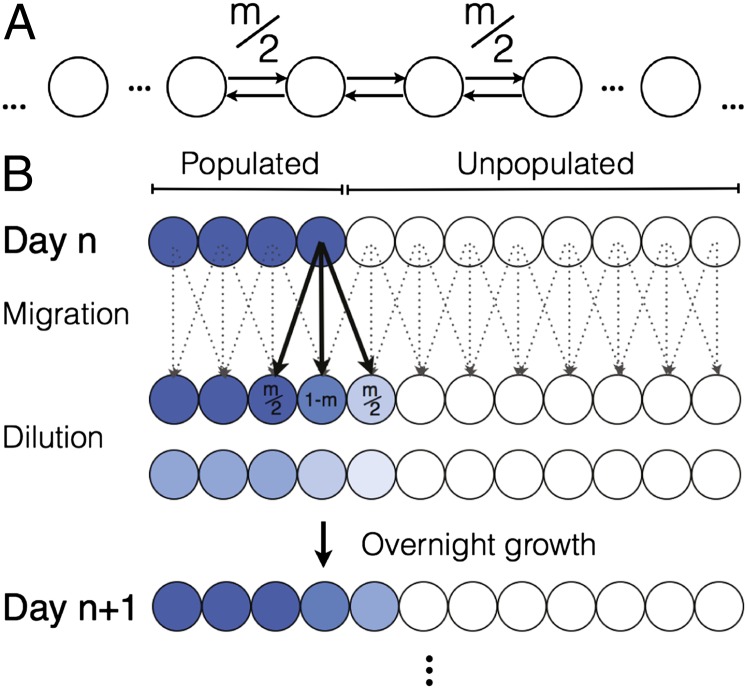
We used an experimental system (similar to that in ref. 40) based upon a linear stepping-stone model (also known as the Levins metapopulation model) (41–43). This theoretical framework, which is well established in the evolutionary and ecological literature, simulates short-range dispersal through nearest-neighbor migration between discrete, well-mixed subpopulations on a lattice (Fig. 1A) (41, 42). Following this scheme, we allowed populations to expand in a habitat of 12 wells (a single row of a 96-well plate), where each well contained a well-mixed population of cells growing in identical growth media (Fig. 1B). To start the experiment, a portion of the wells was populated with cells, and the remainder were unpopulated to allow for expansion. Each day, a fraction  of the contents of each well was transferred into the nearest neighbor wells. Next, the entire population was diluted by a fixed dilution factor (600 in Figs. 2–4), which represents a death process in the growth dynamics. Subsequently, the cells grew overnight. The frequency of each allele was measured via flow cytometry (Materials and Methods and Fig. S1). The process was repeated for 9 d. We note that previous experimental studies of range expansions have focused on expanding colonies on agar plates, which are continuous in both time and space and dominated by stochastic sampling events on the front (7, 8). However, our experimental setup offers a higher level of control over migration and growth parameters, better approximates the patchiness of the natural environment, and allows us to study deterministic effects that are unlikely to manifest themselves in expanding colonies.
of the contents of each well was transferred into the nearest neighbor wells. Next, the entire population was diluted by a fixed dilution factor (600 in Figs. 2–4), which represents a death process in the growth dynamics. Subsequently, the cells grew overnight. The frequency of each allele was measured via flow cytometry (Materials and Methods and Fig. S1). The process was repeated for 9 d. We note that previous experimental studies of range expansions have focused on expanding colonies on agar plates, which are continuous in both time and space and dominated by stochastic sampling events on the front (7, 8). However, our experimental setup offers a higher level of control over migration and growth parameters, better approximates the patchiness of the natural environment, and allows us to study deterministic effects that are unlikely to manifest themselves in expanding colonies.
Fig. 1.
Experimental realization of the stepping-stone model. (A) A schematic of a linear stepping-stone (or Levins metapopulation) model (41–43). The habitat consists of a linear array of subpopulations, each of which is either empty or contains a well-mixed population of individuals. Subpopulations are coupled to each other by nearest-neighbor migration with a fraction m leaving for the two neighboring wells at each time step. (B) Our daily protocol of growth, migration, and death follows the assumptions of the stepping-stone model. Each day, a fraction m/2 of the population from any individual well is migrated into each of the neighboring wells. Boundary conditions are reflective, meaning that wells on the edge receive a contribution m/2 from their only neighbor and 1 – m/2 from the corresponding well. Subsequently, the entire population is diluted by a dilution factor (ranging from 200 to 1,000) that is fixed for the duration of the experiment. The entire population is then grown for 22 h at 30 °C. The process is repeated for several days as the population expands into the unpopulated region. For all experiments, the environmental conditions are spatially homogeneous: all wells (including unpopulated wells) are seeded with identical growth medium [Yeast Extract Peptone Dextrose + Complete Supplement Mixture–His + 2% (wt/vol) sucrose + 0.4× histidine].
Fig. 2.
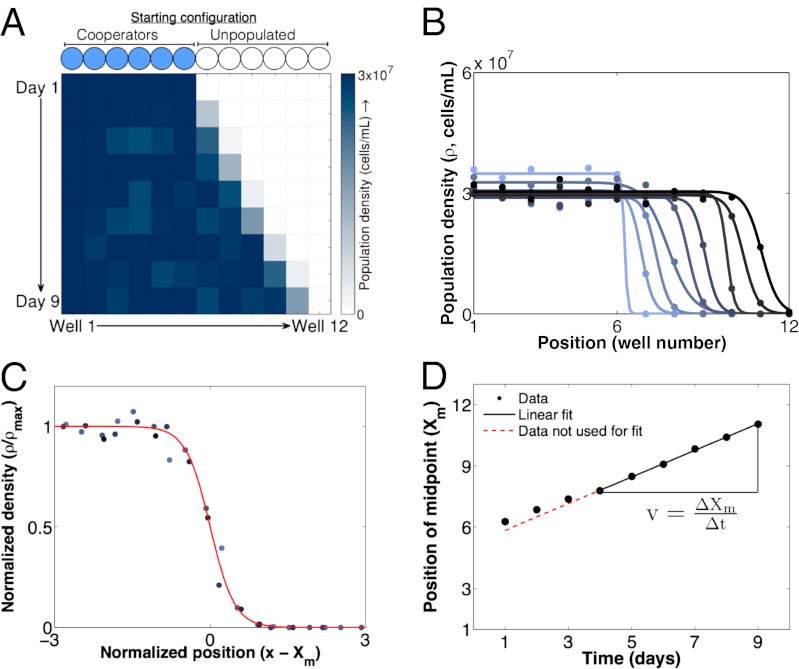
Cooperator populations move as a traveling wave. (A) The experimentally observed one-dimensional expansion of a pure cooperator population over 9 d (m = 0.5 and dilution factor = 600). (B) Density profiles over time for expanding populations of pure cooperators (darker colors indicate later time points). Circles are measurements of population density at a particular time and spatial position. Lines are fits of individual profiles to the hyperbolic tangent function derived in SI Text. (C) An overlay of the density profiles from the last 6 d of the experiment (darker circles indicate later time points). Each profile is normalized to the maximum density found in the bulk population ( max) and shifted by its midpoint position (Xm). The first 3 d are not included, because the expanding population had not yet reached a steady-state profile. The red line shows a theoretical fit to the hyperbolic tangent function derived from a standard reaction–diffusion model of expanding populations (discussed in SI Text). (D) We measure the velocity of the traveling wave by plotting the position of the density profile midpoint (Xm) vs. time and then finding the slope of the line. As in C, the first 3 d are not included.
max) and shifted by its midpoint position (Xm). The first 3 d are not included, because the expanding population had not yet reached a steady-state profile. The red line shows a theoretical fit to the hyperbolic tangent function derived from a standard reaction–diffusion model of expanding populations (discussed in SI Text). (D) We measure the velocity of the traveling wave by plotting the position of the density profile midpoint (Xm) vs. time and then finding the slope of the line. As in C, the first 3 d are not included.
Fig. 4.
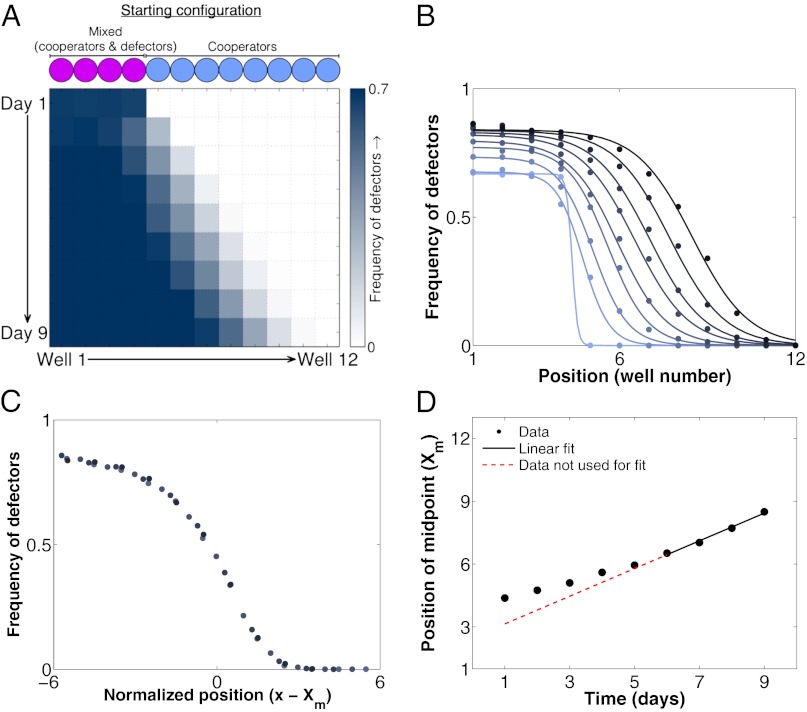
Defectors invade a pure cooperator population as a genetic wave. Note that, unlike in the previous figures, the frequency of defectors, not cooperators, is plotted here. (A) The experimentally observed one-dimensional invasion of defectors into a spatially extended population of cooperators (m = 0.5 and dilution factor = 600). Because defectors (∆suc2) cannot grow in the medium used here without cooperators present, the invasion begins from a mixed cooperator–defector population near its equilibrium genetic composition. (B) Frequency profiles over time for an invading population of defectors (darker colors indicate later time points). Circles are measurements of the frequency of defectors at a particular time and spatial position. Lines are fits of individual profiles to the hyperbolic tangent function derived in SI Text. It is important to note that this fit is based upon a theoretical prediction but is simply used as an approximation to the observed sigmoidal shape to infer the wave midpoint (Xm). (C) An overlay of the frequency profiles from the last 4 d of the experiment (darker circles indicate later time points). Each profile is shifted by its midpoint position (Xm). We note that no red theoretical fit line is shown here because, to our knowledge, there is no theoretical prediction for the shape of the genetic wave profile. (D) We can measure the velocity of the invading genetic wave by using the same procedure that we used for the population density wave (Fig. 2D). The first 6 d of the experiment are excluded from the fit, because the population had not yet reached a steady-state frequency profile and density.
Results
We first explored the spatiotemporal dynamics of pure cooperators undergoing a range expansion. In agreement with theoretical predictions (43), we found that an expanding population of cooperators moves as a traveling wave. Over time, the population adopted a characteristic profile in space consisting of a high-density bulk region and a low-density front (Fig. 2 A and B). The shape of the spatial profile was time-invariant and well fit by the solution to a simple model of range expansions with an Allee effect (SI Text; ref. 43 gives a more detailed treatment). The functional form of the solution has three free parameters: (i) ρmax, the density in the bulk population, (ii) w, the width of the traveling wave, and (iii) Xm, the midpoint of the wave (Fig. 2C). We also found that the wave profile traveled over time with a constant velocity (vcoop = 0.65 ± 0.01 wells/d), which we measured by fitting the time trajectory of the wave midpoint (Xm) to a line (Fig. 2D). The high quality of these fits (R2 = 0.99 for the fit shown here) suggests that the velocity is indeed constant over time. Given the migration scheme, the maximum wave velocity (vmax) is one well/d, because one additional well is populated each day. However, as illustrated above, the velocity can certainly be less than vmax, because it is an emergent property of the expanding population.
Next, we considered the range expansion of a mixed population of cooperators and defectors. Similar to the expanding cooperator population, the mixed population also moved as a traveling wave with a constant profile and velocity (vmixed = 0.49 ± 0.02 wells/d) (Fig. 3A and Fig. S2). Interestingly, we observed that mixed waves of cooperators and defectors traveled more slowly than waves of pure cooperators (vmixed < vcoop), even though the bulk density of the mixed population was higher than that of the cooperators alone (33) (Figs. 2A and 3A and Fig. S3A). This result suggests that, in general, intraspecies interactions can have strong and complex effects on the growth and spread of populations.
Fig. 3.
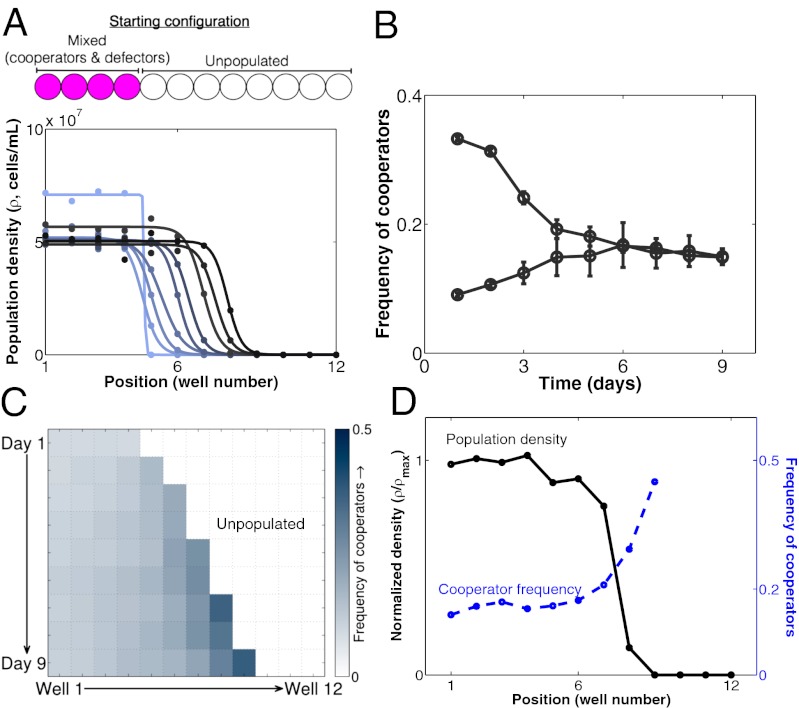
Cooperators are enriched on the front of mixed cooperator–defector waves. (A) Population density profiles for the one-dimensional expansion of a mixed cooperator–defector population (m = 0.5 and dilution factor = 600) at its equilibrium cooperator frequency. Darker colors indicate later time points. Circles are density measurements from a particular day of the experiment. Lines are fits of individual profiles to the hyperbolic tangent function derived in SI Text. (B) In a well-mixed population with no dispersal, the cooperator and defector alleles coexist at an intermediate frequency (in this case, roughly 15%). (C) The frequency of cooperators over time in an expanding mixed cooperator–defector population. The frequency of cooperators is strongly enriched on the front of the expanding population wave (∼45%), whereas the bulk population remains at the equilibrium frequency predicted in B. (D) An overlay of the density profile (in black) with the cooperator frequency profile (in blue) from d 9 of the expansion of the mixed cooperator–defector population. The density profile is normalized to the maximum density found in the bulk population ( max). Cooperators are enriched at the low-density front of the expanding population.
max). Cooperators are enriched at the low-density front of the expanding population.
In addition, we observed significant spatial heterogeneity in allele frequencies within the mixed population wave. Under well-mixed conditions, the cooperator and defector alleles are mutually invasible in our system (24), resulting in stable coexistence between the two alleles at a low frequency of cooperators (Fig. 3B and Fig. S3B). In agreement with the well-mixed prediction, the high-density bulk population maintained a stable equilibrium frequency of cooperators of about 15%. However, we observed that the frequency of cooperators was significantly larger on the low-density front of the expanding population wave, reaching a frequency that was three times higher than that found in the bulk (roughly 45%) (Fig. 3 C and D). It is also interesting to note that, despite the decline in total population density at the front of the expanding population, the density of cooperators actually reaches a peak near the front before declining to zero at the tip of the wave (Fig. S4A).
The cooperator cells are enriched at the front of the traveling wave due to their ability to outcompete defectors at low cell densities. At high densities, defector cells can outcompete the cooperators, because they can take advantage of the sugar produced by the cooperators without having to pay the metabolic “cost” of production. However, at low densities, defectors in the population cannot rely on a high density of cooperator cells to provide the sugar needed for growth (24, 35, 44). Cooperators are able to outcompete defector cells under these conditions because they (selfishly) retain a small fraction of the sugars that they produce (24). Thus, density-dependent growth dynamics, which allow for stable coexistence between the two alleles (24, 35, 44), also lead to strong deterministic enrichment of the cooperative allele on the low-density front of expanding population waves.
Next, we considered the process by which defectors invade a spatially extended population of cooperators. As noted previously, in the absence of migration, defectors can invade a population of cooperators over time (Fig. 3B). We also observed the invasion of defectors into a spatially extended region of cooperators (Fig. 4 A and B). This invasion takes place via a genetic wave with a time-invariant frequency profile (Fig. 4C) and a constant velocity (Fig. 4D) that we termed the “invasion velocity” (vinvasion = 0.55 ± 0.03 wells/d). Thus, we used our experimental system to characterize the genetic wave of invasion for a competing allele. The notion of a genetic wave was first introduced by Fisher (45) and Kolmogorov et al. (46), but thus far experimental characterizations of genetic waves in spatially extended populations have been scarce (however, see examples in refs. 11 and 47).
Notably, we found that the invasion velocity was less than the velocity of migrating cooperators (vinvasion < vcoop). We were intrigued by this comparison, because it suggested that, if a population of cooperators continued to migrate as it was invaded, the two populations might never completely mix. To explore this idea further, we repeated our measurements of vcoop, vinvasion, and vmixed over a range of dilution factors from 200 to 1,000 (Fig. 5A). The dilution factor, which modulates the daily death rate, allowed us to vary the dynamics of the system by changing the effective “severity” of the environment. Using this experimental control knob, we hoped to probe a broader range of dynamics in the system.
Fig. 5.
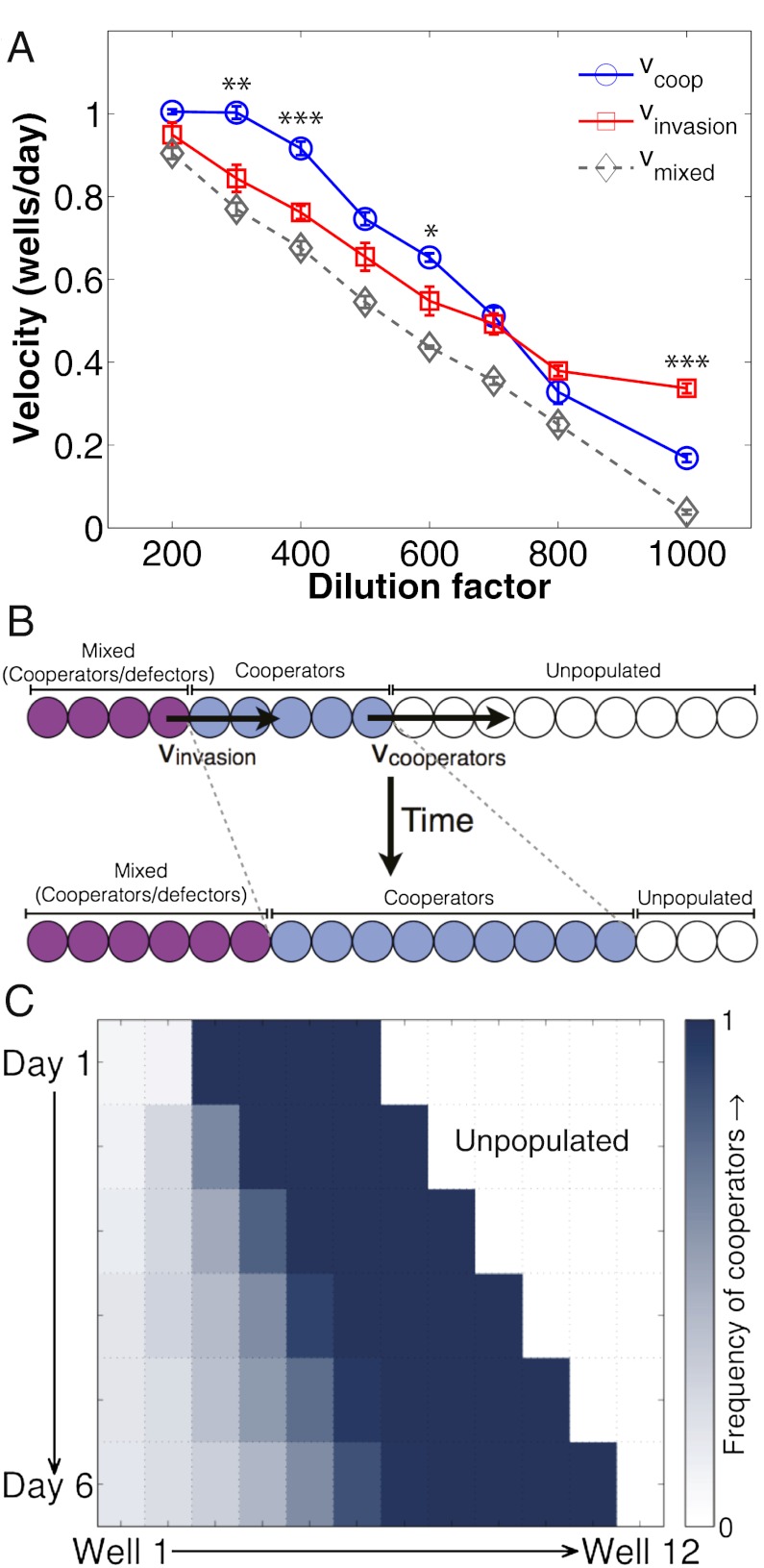
Cooperators can “outrun” an invading defector wave. (A) Measurement of the velocities of pure cooperators (vcoop), invading defectors (vinvasion), and the mixed cooperator-defector wave (vmixed) over a range of dilution factors indicates two regimes: (i) at high dilution factors, defectors invade more quickly than the cooperators can escape (vinvasion > vcoop) and (ii) at low dilution factors, cooperators can outrun the invasion (vcoop > vinvasion). Error bars for the velocities are SEs in the slope of the Xm vs. time plot. Asterisks indicate the magnitude of the P value for the difference between vcoop and vinvasion at a particular dilution factor (*P < 0.05, **P < 0.01, ***P < 0.001). (B) A schematic depicting the case in which cooperators can outrun defectors, in which the region occupied by the cooperators increases over time, even as the defectors invade. (C) Experimental observation of cooperators outrunning an invading wave of defectors (m = 0.5 and dilution factor = 400). At this dilution factor, ∆v = vcoop – vinvasion ∼ 0.2 wells/d. Over 5 d, the “headstart” region containing pure cooperators increases from four to nearly five wells, consistent with the observed ∆v.
As we increased the dilution factor, we found that each of the velocities we measured (vcoop, vinvasion, and vmixed) decreased monotonically (Fig. 5A and Fig. S4B for vcoop). For population waves (e.g., vcoop and vmixed), this result can be explained by the fact that newly populated wells on the front of the wave cannot grow to measurable densities at higher dilution factors, thereby impeding the spread of the population wave. For the genetic wave (vinvasion), this monotonic decrease is consistent with the dependence predicted by the Fisher–Kolmogorov model (refs. 36 and 37 and SI Text). Interestingly, we also found that the mixed cooperator–defector wave traveled more slowly than both the pure cooperator wave and the defector invasion wave over the entire range of dilution factors that we probed (Fig. 5A). This result draws comparison with theoretical predictions suggesting that opportunistic pathogens can slow the migration of biological species (48), and it indicates that interactions between cells can have a significant influence on population expansions.
Most surprisingly, although vcoop and vinvasion showed similar qualitative trends, a comparison of the two velocities immediately revealed two distinct regimes (Fig. 5A). At high dilution factors (above 700), we observed that vinvasion was greater than vcoop, whereas vinvasion was less than vcoop at low dilution factors (below 700). These relationships between vcoop and vinvasion are surprising, given that under well-mixed conditions cooperators have a growth advantage at low densities (high dilution factors), whereas defectors are favored at high densities (low dilution factors). However, it is important to note that, although these two expansion processes are both emergent properties of the well-mixed dynamics in individual wells, they occur under significantly different environmental conditions. In particular, cooperators spread by colonizing previously unoccupied regions; thus, at the front of the wave, population densities (and therefore, the concentration of glucose available for consumption) are low. However, defectors spread into wells that are already occupied by cooperators, in which population densities are high and growth conditions are favorable (glucose concentration is high). Thus, our analysis shows that the velocities of range expansions are strongly coupled to the environment and can produce relationships that are not immediately apparent from dynamics observed in a well-mixed context. Interestingly, a simple phenomenological model of yeast growth in sucrose that we have used previously (24, 35, 44) predicts our experimentally observed crossing of velocities as a function of dilution factor (Fig. S5).
From an ecological perspective, the relationship between vcoop and vinvasion has several interesting implications. At high dilution factors (vinvasion > vcoop), cooperators and defectors will eventually become completely mixed in space, after which the two alleles will spread together into new territories (Fig. S6). In contrast, at low dilution factors (vcoop > vinvasion), cooperators can “outrun” an invading wave of defectors (Fig. 5B). In this case, a sufficiently large region of pure cooperators expands over time, even in the presence of an invasive allele (Fig. 5C, Fig. S7, and SI Text). Broadly, the ability of cooperators to outrun an invading defector wave provides a second mechanism through which range expansions could help to maintain cooperation in nature.
Discussion
We have presented an experimental study of the effect of range expansions on the maintenance of cooperation. Using yeast populations in the laboratory to study range expansions gave us direct control over all migration parameters and experimental conditions. As a result, we were able to perform a high-resolution analysis of an expanding population of cooperating alleles in a manner that could not easily be replicated in the natural environment.
Using this approach, we found that range expansions favor the maintenance of cooperation in two ways. First, cooperation is strongly enriched on the front of expanding populations compared with the bulk, even without spatial heterogeneity in the environmental conditions. Given that range expansions are common in natural populations, they could aid in the maintenance of cooperation in nature. Moreover, the mechanism of enrichment can be applied more generally to any allele favored at low densities. Second, we demonstrated that cooperators can outrun an invading wave of defectors under certain conditions. Because most natural populations are spatially extended, outrunning provides a plausible mechanism through which cooperation could be maintained in spatially extended populations.
To our knowledge, the two mechanisms that we have demonstrated here are different from those described previously. These unique mechanisms do not invoke kin selection or any form of reciprocity, both of which are deterministic effects that have been shown to lead to stable coexistence between cooperators and defectors under well-mixed conditions (12, 25). Stochastic effects due to population bottlenecks [i.e., allele surfing (1) and Simpson’s paradox (22)] are also unlikely to play a large role in our experiments, given the relatively large population size, rates of dispersal between wells, and an Allee effect, which limits population growth below a critical density. Moreover, the mechanisms that we described do not rely on spatial heterogeneity in environmental conditions generated by deteriorating conditions (26, 49) or habitat destruction (49, 50). Thus, our experiments suggest two distinct deterministic mechanisms in which cooperation could be maintained in a spatially homogeneous environment.
We also believe that our results are broadly generalizable, because many examples of obligate cheating behaviors like those in our system exist in nature. Indeed, several recent studies of microbes isolated from the environment (see ref. 51 for an example) indicate that complete gene loss—which leads to obligate cheating—is a common mechanism through which defectors can arise in natural populations. However, we do note that facultative cheating has been observed in experimental populations (52, 53), which we have not studied here.
Given that cooperation is enriched at the expanding wave front and migrating cooperators can outrun an invading defector wave, we might expect cooperators to be able to “split”—that is, spatially separate themselves—from a mixed population of cooperators and defectors (Fig. S7 shows a schematic). This splitting effect would allow a cooperator population effectively to purify itself of defectors in a way that could not be achieved in a nonmigrating population. However, recent theoretical predictions suggest that these two features are necessary but not sufficient for splitting to occur (54). Indeed, in line with this prediction, we found that the cooperator and defector populations moved at the same velocity within the mixed population wave (e.g., Fig. S4A), even though the velocities of each population individually were significantly different. This finding suggests that splitting does not occur in our system, even at dilution factors at which both cooperation enrichment and outrunning are observed.
In part, the absence of splitting can be attributed to the overall lowered velocity of the mixed cooperator–defector wave compared with the pure cooperators (vmixed < vcoop over all dilution factors tested). This suggests that cooperators are “slowed down” by their interactions with defectors, allowing the two alleles to move together in the mixed wave in a way that maintains allelic diversity in expanding populations. A second corollary of this slowing-down effect is that, given that both vmixed and vcoop decrease monotonically with the severity of the environment, vmixed will reach zero while vcoop is still positive. As such, our results suggest that a population of pure cooperators may be able to expand into harsh environments that would be inaccessible to the mixed population. Thus, our experiments provide support for an additional mechanism, which had previously been described theoretically (26), through which cooperation could be aided via ecological constraints.
Overall, we suggest that the coupling of ecological and evolutionary effects drives the spatiotemporal dynamics of cooperation in our system. However, given the generality of our analyses, these ideas can easily be extended to the general case of alleles under frequency- or density-dependent selection, which is a widespread feature of many natural ecosystems. Thus, our findings support the growing, but still underappreciated, notion that eco-evolutionary feedback may dictate the growth and survival of natural populations (26, 35, 44).
Materials and Methods
Strains.
All strains are identical to those used in Gore et al. (24). Strains were derived from the haploid BY4741 strain of Saccharomyces cerevisiae [mating type a, European Saccharomyces Cerevisiae Archive for Functional Analysis (EUROSCARF)]. The cooperator strain has a wild-type SUC2 gene, a mutated HIS3 (his3Δ1), and has yellow fluorescent protein expressed constitutively from the ADH1 promoter (inserted using plasmid pRS401 with a MET17 marker). The defector strain lacks the SUC2 gene (EUROSCARF Y02321, SUC2::kanMX4), has the wild-type HIS3 gene, and has the fluorescent protein tdTomato expressed constitutively from the PGK1 promoter (inserted using plasmid pRS301 containing a HIS3 marker).
Experimental Protocols.
All experiments were performed in 200-μL batch culture in BD Biosciences Falcon 96-well Microtest plates. All cultures were grown at 30 °C in synthetic media (yeast nitrogen base and Complete Supplement Mixture–His) supplemented with 2% (wt/vol) sucrose and 0.4× (8 μg/mL) histidine. It is important to note that, because the cooperator strain is a histidine auxotroph, the histidine concentration can be used to tune the “cost” of cooperation by preferentially limiting the growth of the cooperators (24).
Cultures were shaken continuously at 800 rpm during growth. To avoid evaporation and cross-contamination between wells, plates were covered for the duration of the experiment with Parafilm Laboratory Film. Each day, the frequency of cooperators and defectors was measured via flow cytometry (FACS LSR II HTS; BD Biosciences). The population density was estimated with flow cytometry and benchmarked with optical density measurements at 620 nm with a Thermo Scientific Multiskan FC microplate spectrophotometer.
Cultures underwent a migration step and a dilution step each day following 22 h of overnight growth. During the migration step, a portion 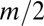 of the cells from each well was transferred into each of the two neighboring wells on a new 96-well plate. The remaining
of the cells from each well was transferred into each of the two neighboring wells on a new 96-well plate. The remaining 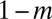 cells were transferred to the corresponding well on the new plate. We used m = 0.5 in all experiments. Boundary conditions were reflective, meaning that wells on the edge received a contribution
cells were transferred to the corresponding well on the new plate. We used m = 0.5 in all experiments. Boundary conditions were reflective, meaning that wells on the edge received a contribution 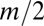 from their single neighbor, with the remainder coming from the corresponding well.
from their single neighbor, with the remainder coming from the corresponding well.
Serial dilutions were performed each day with a fixed dilution factor ranging from 200 to 1,000. The data presented in Figs. 2–4 were diluted by a factor of 600 each day, in Fig. 5C by a factor of 400, and in Fig. 5A and Figs. S4B and S3 with dilution factors of 200, 300, 400, 500, 600, 800, and 1,000.
It is important to note that all data shown in the main text was obtained from a single 9-d run of the experiment. In three independent experiments, we found that the dilution factor at which particular velocities were observed was highly variable (±20%). However, our core conclusions were robust to this variation in each experiment (Fig. S8 gives examples).
Supplementary Material
Acknowledgments
We thank members of the J.G. laboratory and Nathan Cermak for helpful discussions, as well as the personnel at the Koch Institute Flow Cytometry Core at Massachusetts Institute of Technology (MIT) for experimental help (supported by Cancer Center Support Grant P30CCA14051 from the National Cancer Institute). M.S.D. was supported by the Department of Defense through the National Defense Science and Engineering Graduate Fellowship Program. K.S.K. was supported by a Pappalardo Fellowship at MIT. C.D. was supported by MIT’s Undergraduate Research Opportunities Program. The laboratory acknowledges support from National Institutes of Health (NIH) R00 Pathways to Independence Award GM085279-02, National Science Foundation Career Award PHY-1055154, Pew Fellowship 2010-000224-007, Sloan Foundation Fellowship BR2011-066, and NIH New Innovator Award DP2.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1217517110/-/DCSupplemental.
References
- 1.Excoffier L, Foll M, Petit RJ. Genetic consequences of range expansions. Annu Rev Ecol Evol Syst. 2009;40:481–501. [Google Scholar]
- 2.Gray ME, Sappington TW, Miller NJ, Moeser J, Bohn MO. Adaptation and invasiveness of western corn rootworm: Intensifying research on a worsening pest. Annu Rev Entomol. 2009;54:303–321. doi: 10.1146/annurev.ento.54.110807.090434. [DOI] [PubMed] [Google Scholar]
- 3.Van Bocxlaer I, et al. Gradual adaptation toward a range-expansion phenotype initiated the global radiation of toads. Science. 2010;327(5966):679–682. doi: 10.1126/science.1181707. [DOI] [PubMed] [Google Scholar]
- 4.Moreau C, et al. Deep human genealogies reveal a selective advantage to be on an expanding wave front. Science. 2011;334(6059):1148–1150. doi: 10.1126/science.1212880. [DOI] [PubMed] [Google Scholar]
- 5.Walther G-R, et al. Ecological responses to recent climate change. Nature. 2002;416(6879):389–395. doi: 10.1038/416389a. [DOI] [PubMed] [Google Scholar]
- 6.Excoffier L, Ray N. Surfing during population expansions promotes genetic revolutions and structuration. Trends Ecol Evol. 2008;23(7):347–351. doi: 10.1016/j.tree.2008.04.004. [DOI] [PubMed] [Google Scholar]
- 7.Korolev KS, Avlund M, Hallatschek O, Nelson DR. Genetic demixing and evolution in linear stepping stone models. Rev Mod Phys. 2010;82(2):1691–1718. doi: 10.1103/RevModPhys.82.1691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hallatschek O, Hersen P, Ramanathan S, Nelson DR. Genetic drift at expanding frontiers promotes gene segregation. Proc Natl Acad Sci USA. 2007;104(50):19926–19930. doi: 10.1073/pnas.0710150104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Manica A, Amos W, Balloux F, Hanihara T. The effect of ancient population bottlenecks on human phenotypic variation. Nature. 2007;448(7151):346–348. doi: 10.1038/nature05951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Atkinson QD. Phonemic diversity supports a serial founder effect model of language expansion from Africa. Science. 2011;332(6027):346–349. doi: 10.1126/science.1199295. [DOI] [PubMed] [Google Scholar]
- 11.Novembre J, Di Rienzo A. Spatial patterns of variation due to natural selection in humans. Nat Rev Genet. 2009;10(11):745–755. doi: 10.1038/nrg2632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Nowak MA. Five rules for the evolution of cooperation. Science. 2006;314(5805):1560–1563. doi: 10.1126/science.1133755. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Wingreen NS, Levin SA. Cooperation among microorganisms. PLoS Biol. 2006;4(9):e299. doi: 10.1371/journal.pbio.0040299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.West SA, Griffin AS, Gardner A. Evolutionary explanations for cooperation. Curr Biol. 2007;17(16):R661–R672. doi: 10.1016/j.cub.2007.06.004. [DOI] [PubMed] [Google Scholar]
- 15.Xavier JB. Social interaction in synthetic and natural microbial communities. Mol Syst Biol. 2011;7(483):1–11. doi: 10.1038/msb.2011.16. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Damore JA, Gore J. Understanding microbial cooperation. J Theor Biol. 2012;299:31–41. doi: 10.1016/j.jtbi.2011.03.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Celiker H, Gore J. 2013. Cellular cooperation: Insights from microbes. Trends Cbell Biol 23(1):9–15.
- 18.Griffin AS, West SA, Buckling A. Cooperation and competition in pathogenic bacteria. Nature. 2004;430(7003):1024–1027. doi: 10.1038/nature02744. [DOI] [PubMed] [Google Scholar]
- 19.Craig Maclean R, Brandon C. Stable public goods cooperation and dynamic social interactions in yeast. J Evol Biol. 2008;21(6):1836–1843. doi: 10.1111/j.1420-9101.2008.01579.x. [DOI] [PubMed] [Google Scholar]
- 20.Nadell CD, Foster KR, Xavier JB. Emergence of spatial structure in cell groups and the evolution of cooperation. PLOS Comput Biol. 2010;6(3):e1000716. doi: 10.1371/journal.pcbi.1000716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Xavier JB, Kim W, Foster KR. A molecular mechanism that stabilizes cooperative secretions in Pseudomonas aeruginosa. Mol Microbiol. 2011;79(1):166–179. doi: 10.1111/j.1365-2958.2010.07436.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chuang JS, Rivoire O, Leibler S. Simpson’s paradox in a synthetic microbial system. Science. 2009;323(5911):272–275. doi: 10.1126/science.1166739. [DOI] [PubMed] [Google Scholar]
- 23.Cremer J, Melbinger A, Frey E. Evolutionary and population dynamics: a coupled approach. Phys Rev E Stat Nonlin Soft Matter Phys. 2011;84(5 Pt 1):051921. doi: 10.1103/PhysRevE.84.051921. [DOI] [PubMed] [Google Scholar]
- 24.Gore J, Youk H, van Oudenaarden A. Snowdrift game dynamics and facultative cheating in yeast. Nature. 2009;459(7244):253–256. doi: 10.1038/nature07921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Turner PE, Chao L. Prisoner’s dilemma in an RNA virus. Nature. 1999;398(6726):441–443. doi: 10.1038/18913. [DOI] [PubMed] [Google Scholar]
- 26.Zhang F, Hui C. Eco-evolutionary feedback and the invasion of cooperation in prisoner’s dilemma games. PLoS ONE. 2011;6(11):e27523. doi: 10.1371/journal.pone.0027523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Ross-Gillespie A, Gardner A, Buckling A, West SA, Griffin AS. Density dependence and cooperation: Theory and a test with bacteria. Evolution. 2009;63(9):2315–2325. doi: 10.1111/j.1558-5646.2009.00723.x. [DOI] [PubMed] [Google Scholar]
- 28.Lampert A, Tlusty T. Density-dependent cooperation as a mechanism for persistence and coexistence. Evolution. 2011;65(10):2750–2759. doi: 10.1111/j.1558-5646.2011.01364.x. [DOI] [PubMed] [Google Scholar]
- 29.Pruitt JN, Riechert SE. Frequency-dependent success of cheaters during foraging bouts might limit their spread within colonies of a socially polymorphic spider. Evolution. 2009;63(11):2966–2973. doi: 10.1111/j.1558-5646.2009.00771.x. [DOI] [PubMed] [Google Scholar]
- 30.Ross-Gillespie A, Gardner A, West SA, Griffin AS. Frequency dependence and cooperation: Theory and a test with bacteria. Am Nat. 2007;170(3):331–342. doi: 10.1086/519860. [DOI] [PubMed] [Google Scholar]
- 31.Velenich A, Gore J. Synthetic approaches to understanding biological constraints. Curr Opin Chem Biol. 2012;16(3–4):323–328. doi: 10.1016/j.cbpa.2012.05.199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Greig D, Travisano M. The prisoner’s dilemma and polymorphism in yeast SUC genes. Proc Biol Sci. 2004;271(Suppl 3):S25–S26. doi: 10.1098/rsbl.2003.0083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.MaClean RC, Fuentes-Hernandez A, Greig D, Hurst LD, Gudelj I. A mixture of “cheats” and “co-operators” can enable maximal group benefit. PLoS Biol. 2010;8(9):e1000486. doi: 10.1371/journal.pbio.1000486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Koschwanez JH, Foster KR, Murray AW. Sucrose utilization in budding yeast as a model for the origin of undifferentiated multicellularity. PLoS Biol. 2011;9(8):e1001122. doi: 10.1371/journal.pbio.1001122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Sanchez A, Gore J. 2013. Feedback between population and evolutionary dynamics determines the fate of social microbial populations. PLoS Biology, arXiv:13012791.
- 36.Courchamp F, Clutton-Brock T, Grenfell B. Inverse density dependence and the Allee effect. Trends Ecol Evol. 1999;14(10):405–410. doi: 10.1016/s0169-5347(99)01683-3. [DOI] [PubMed] [Google Scholar]
- 37.Dai L, Vorselen D, Korolev KS, Gore J. Generic indicators for loss of resilience before a tipping point leading to population collapse. Science. 2012;336(6085):1175–1177. doi: 10.1126/science.1219805. [DOI] [PubMed] [Google Scholar]
- 38.Roques L, Garnier J, Hamel F, Klein EK. Allee effect promotes diversity in traveling waves of colonization. Proc Natl Acad Sci USA. 2012;109(23):8828–8833. doi: 10.1073/pnas.1201695109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Tobin PC, Berec L, Liebhold AM. Exploiting Allee effects for managing biological invasions. Ecol Lett. 2011;14(6):615–624. doi: 10.1111/j.1461-0248.2011.01614.x. [DOI] [PubMed] [Google Scholar]
- 40.Kerr B, Neuhauser C, Bohannan BJM, Dean AM. Local migration promotes competitive restraint in a host-pathogen ‘tragedy of the commons.’. Nature. 2006;442(7098):75–78. doi: 10.1038/nature04864. [DOI] [PubMed] [Google Scholar]
- 41.Kimura M, Weiss GH. The stepping stone model of population structure and the decrease of genetic correlation with distance. Genetics. 1964;49(4):561–576. doi: 10.1093/genetics/49.4.561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hanski I. Metapopulation Ecology. New York: Oxford Univ Press; 1999. [Google Scholar]
- 43.Murray JD. Mathematical Biology II. 3rd Ed. Berlin: Springer; 2003. [Google Scholar]
- 44.Celiker H, Gore J. Competition between species can stabilize public-goods cooperation within a species. Mol Syst Biol. 2012 doi: 10.1038/msb.2012.54. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Fisher RA. The wave of advance of advantageous genes. Ann Eugen. 1937;7:355–369. [Google Scholar]
- 46.Kolmogorov A, Petrovskii I, Piscounov N. Selected Works of A. N. Kolmogorov I. Dordrecht, The Netherlands: Kluwer; 1991. pp. 248–270. [Google Scholar]
- 47.Matsushita M, et al. 1999. Formation of colony patterns by a bacterial cell population. Physica A: Statistical Mechanics Its Applications 274:190–199.
- 48.Hilker FM, Lewis MA, Seno H, Langlais M, Malchow H. Pathogens can slow down or reverse invasion fronts of their hosts. Biol Invasions. 2005;7:817–832. [Google Scholar]
- 49.Zhang F, Hui C, Han X, Li Z. Evolution of cooperation in patchy habitat under patch decay and isolation. Ecol Res. 2005;20:461–469. [Google Scholar]
- 50.Brockhurst MA, Buckling A, Gardner A. Cooperation peaks at intermediate disturbance. Curr Biol. 2007;17(9):761–765. doi: 10.1016/j.cub.2007.02.057. [DOI] [PubMed] [Google Scholar]
- 51.Cordero OX, Ventouras L-A, DeLong EF, Polz MF. 2012. Public good dynamics drive evolution of iron acquisition strategies in natural bacterioplankton populations. Proc Natl Acad Sci USA 109(49):20059–20064.
- 52.Strassmann JE, Queller DC. Evolution of cooperation and control of cheating in a social microbe. Proc Natl Acad Sci USA. 2011;108(Suppl 2):10855–10862. doi: 10.1073/pnas.1102451108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Santorelli LA, et al. Facultative cheater mutants reveal the genetic complexity of cooperation in social amoebae. Nature. 2008;451(7182):1107–1110. doi: 10.1038/nature06558. [DOI] [PubMed] [Google Scholar]
- 54.Korolev KS. 2012. The fate of cooperation during range expansions. PLOS Computational Biology, arXiv:12100498.
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.