Abstract
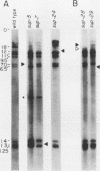
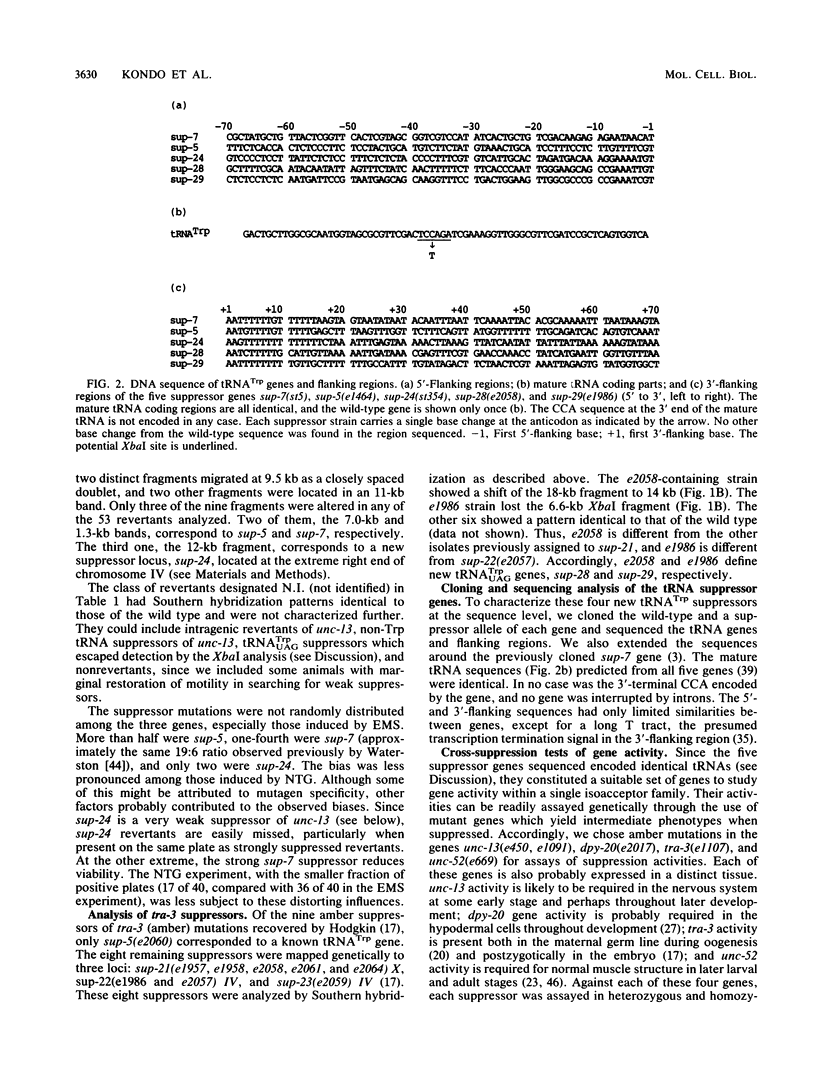
Caenorhabditis elegans has 12 tRNA(UGGTrp) genes as defined by Southern analysis. In order to evaluate the function of the individual members of this multigene family, we sought to recover amber (UAG)-suppressing mutations from reversion experiments with animals carrying amber mutations in a nervous system-affecting gene (unc-13) or a sex-determining gene (tra-3). Revertants were analyzed by Southern blot, exploiting the fact that the CCA to CTA change at the anticodon creates a new XbaI site. Five different members of the tRNATrp gene family were identified as suppressors: sup-7 X, sup-5 III, sup-24 IV, sup-28 X, and sup-29 IV. All five suppressor genes were sequenced and found to encode identical tRNA(UAGTrp) molecules with a single base change (CCA to CTA) at the anticodon compared with their wild-type counterparts. The flanking sequences had only limited homology. The relative expression of these five genes was determined by measuring the efficiencies of suppressers against amber mutations in genes affecting the nervous system, hypodermis, muscle, and sex determination. The results of these cross-suppression tests showed that the five members of the tRNA(Trp) gene family were differentially regulated in a tissue- or development stage-specific manner.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allison D. S., Hall B. D. Effects of alterations in the 3' flanking sequence on in vivo and in vitro expression of the yeast SUP4-o tRNATyr gene. EMBO J. 1985 Oct;4(10):2657–2664. doi: 10.1002/j.1460-2075.1985.tb03984.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barak L. S., Yocum R. R., Nothnagel E. A., Webb W. W. Fluorescence staining of the actin cytoskeleton in living cells with 7-nitrobenz-2-oxa-1,3-diazole-phallacidin. Proc Natl Acad Sci U S A. 1980 Feb;77(2):980–984. doi: 10.1073/pnas.77.2.980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolten S. L., Powell-Abel P., Fischhoff D. A., Waterston R. H. The sup-7(st5) X gene of Caenorhabditis elegans encodes a tRNATrpUAG amber suppressor. Proc Natl Acad Sci U S A. 1984 Nov;81(21):6784–6788. doi: 10.1073/pnas.81.21.6784. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974 May;77(1):71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen E. Y., Seeburg P. H. Supercoil sequencing: a fast and simple method for sequencing plasmid DNA. DNA. 1985 Apr;4(2):165–170. doi: 10.1089/dna.1985.4.165. [DOI] [PubMed] [Google Scholar]
- Ciliberto G., Castagnoli L., Melton D. A., Cortese R. Promoter of a eukaryotic tRNAPro gene is composed of three noncontiguous regions. Proc Natl Acad Sci U S A. 1982 Feb;79(4):1195–1199. doi: 10.1073/pnas.79.4.1195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coulondre C., Miller J. H. Genetic studies of the lac repressor. IV. Mutagenic specificity in the lacI gene of Escherichia coli. J Mol Biol. 1977 Dec 15;117(3):577–606. doi: 10.1016/0022-2836(77)90059-6. [DOI] [PubMed] [Google Scholar]
- DeFranco D., Sharp S., Söll D. Identification of regulatory sequences contained in the 5'-flanking region of Drosophila lysine tRNA2 genes. J Biol Chem. 1981 Dec 10;256(23):12424–12429. [PubMed] [Google Scholar]
- Dibb N. J., Brown D. M., Karn J., Moerman D. G., Bolten S. L., Waterston R. H. Sequence analysis of mutations that affect the synthesis, assembly and enzymatic activity of the unc-54 myosin heavy chain of Caenorhabditis elegans. J Mol Biol. 1985 Jun 25;183(4):543–551. doi: 10.1016/0022-2836(85)90170-6. [DOI] [PubMed] [Google Scholar]
- Elder R. T., Szabo P., Uhlenbeck O. C. In situ hybridization of three transfer RNAs to the polytene chromosomes of Drosophila melanogaster. J Mol Biol. 1980 Sep 5;142(1):1–17. doi: 10.1016/0022-2836(80)90202-8. [DOI] [PubMed] [Google Scholar]
- Fire A. Integrative transformation of Caenorhabditis elegans. EMBO J. 1986 Oct;5(10):2673–2680. doi: 10.1002/j.1460-2075.1986.tb04550.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galli G., Hofstetter H., Birnstiel M. L. Two conserved sequence blocks within eukaryotic tRNA genes are major promoter elements. Nature. 1981 Dec 17;294(5842):626–631. doi: 10.1038/294626a0. [DOI] [PubMed] [Google Scholar]
- Garel J. P., Mandel P., Chavancy G., Daillie J. Functional adaptation of tRNAs to fibroin biosynthesis in the silkgland of Bombyx mori L. FEBS Lett. 1970 May 1;7(4):327–329. doi: 10.1016/0014-5793(70)80196-x. [DOI] [PubMed] [Google Scholar]
- Greenwald I. S., Horvitz H. R. unc-93(e1500): A behavioral mutant of Caenorhabditis elegans that defines a gene with a wild-type null phenotype. Genetics. 1980 Sep;96(1):147–164. doi: 10.1093/genetics/96.1.147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodgkin J. Novel nematode amber suppressors. Genetics. 1985 Oct;111(2):287–310. doi: 10.1093/genetics/111.2.287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hofstetter H., Kressman A., Birnstiel M. L. A split promoter for a eucaryotic tRNA gene. Cell. 1981 May;24(2):573–585. doi: 10.1016/0092-8674(81)90348-2. [DOI] [PubMed] [Google Scholar]
- Horvitz H. R., Brenner S., Hodgkin J., Herman R. K. A uniform genetic nomenclature for the nematode Caenorhabditis elegans. Mol Gen Genet. 1979 Sep;175(2):129–133. doi: 10.1007/BF00425528. [DOI] [PubMed] [Google Scholar]
- Kimble J., Hodgkin J., Smith T., Smith J. Suppression of an amber mutation by microinjection of suppressor tRNA in C. elegans. Nature. 1982 Sep 30;299(5882):456–458. doi: 10.1038/299456a0. [DOI] [PubMed] [Google Scholar]
- Kurjan J., Hall B. D., Gillam S., Smith M. Mutations at the yeast SUP4 tRNATyr locus: DNA sequence changes in mutants lacking suppressor activity. Cell. 1980 Jul;20(3):701–709. doi: 10.1016/0092-8674(80)90316-5. [DOI] [PubMed] [Google Scholar]
- Lizardi P. M., Mahdavi V., Shields D., Candelas G. Discontinuous translation of silk fibroin in a reticulocyte cell-free system and in intact silk gland cells. Proc Natl Acad Sci U S A. 1979 Dec;76(12):6211–6215. doi: 10.1073/pnas.76.12.6211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mackenzie J. M., Jr, Garcea R. L., Zengel J. M., Epstein H. F. Muscle development in Caenorhabditis elegans: mutants exhibiting retarded sarcomere construction. Cell. 1978 Nov;15(3):751–762. doi: 10.1016/0092-8674(78)90261-1. [DOI] [PubMed] [Google Scholar]
- Müller F., Clarkson S. G. Nucleotide sequence of genes coding for tRNAPhe and tRNATyr from a repeating unit of X. laevis DNA. Cell. 1980 Feb;19(2):345–353. doi: 10.1016/0092-8674(80)90509-7. [DOI] [PubMed] [Google Scholar]
- Nishikura K., Kurjan J., Hall B. D., De Robertis E. M. Genetic analysis of the processing of a spliced tRNA. EMBO J. 1982;1(2):263–268. doi: 10.1002/j.1460-2075.1982.tb01157.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Priess J. R., Hirsh D. I. Caenorhabditis elegans morphogenesis: the role of the cytoskeleton in elongation of the embryo. Dev Biol. 1986 Sep;117(1):156–173. doi: 10.1016/0012-1606(86)90358-1. [DOI] [PubMed] [Google Scholar]
- Reed K. C., Mann D. A. Rapid transfer of DNA from agarose gels to nylon membranes. Nucleic Acids Res. 1985 Oct 25;13(20):7207–7221. doi: 10.1093/nar/13.20.7207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Riddle D. L., Brenner S. Indirect suppression in Caenorhabditis elegans. Genetics. 1978 Jun;89(2):299–314. doi: 10.1093/genetics/89.2.299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein R. J., Esposito R. E., Esposito M. S. The effect of ochre suppression on meiosis and ascospore formation in Saccharomyces. Genetics. 1977 Jan;85(1):35–54. doi: 10.1093/genetics/85.1.35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp S. J., Schaack J., Cooley L., Burke D. J., Söll D. Structure and transcription of eukaryotic tRNA genes. CRC Crit Rev Biochem. 1985;19(2):107–144. doi: 10.3109/10409238509082541. [DOI] [PubMed] [Google Scholar]
- Shaw K. J., Olson M. V. Effects of altered 5'-flanking sequences on the in vivo expression of a Saccharomyces cerevisiae tRNATyr gene. Mol Cell Biol. 1984 Apr;4(4):657–665. doi: 10.1128/mcb.4.4.657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silverman S., Schmidt O., Söll D., Hovemann B. The nucleotide sequence of a cloned Drosophila arginine tRNA gene and its in vitro transcription in Xenopus germinal vesicle extracts. J Biol Chem. 1979 Oct 25;254(20):10290–10294. [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Sprague K. U., Hagenbüchle O., Zuniga M. C. The nucleotide sequence of two silk gland alanine tRNAs: implications for fibroin synthesis and for initiator tRNA structure. Cell. 1977 Jul;11(3):561–570. doi: 10.1016/0092-8674(77)90074-5. [DOI] [PubMed] [Google Scholar]
- Sprague K. U., Larson D., Morton D. 5' flanking sequence signals are required for activity of silkworm alanine tRNA genes in homologous in vitro transcription systems. Cell. 1980 Nov;22(1 Pt 1):171–178. doi: 10.1016/0092-8674(80)90165-8. [DOI] [PubMed] [Google Scholar]
- Sulston J. E., Brenner S. The DNA of Caenorhabditis elegans. Genetics. 1974 May;77(1):95–104. doi: 10.1093/genetics/77.1.95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vogelstein B., Gillespie D. Preparative and analytical purification of DNA from agarose. Proc Natl Acad Sci U S A. 1979 Feb;76(2):615–619. doi: 10.1073/pnas.76.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward S. Chemotaxis by the nematode Caenorhabditis elegans: identification of attractants and analysis of the response by use of mutants. Proc Natl Acad Sci U S A. 1973 Mar;70(3):817–821. doi: 10.1073/pnas.70.3.817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterston R. H. A second informational suppressor, SUP-7 X, in Caenorhabditis elegans. Genetics. 1981 Feb;97(2):307–325. doi: 10.1093/genetics/97.2.307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waterston R. H., Brenner S. A suppressor mutation in the nematode acting on specific alleles of many genes. Nature. 1978 Oct 26;275(5682):715–719. doi: 10.1038/275715a0. [DOI] [PubMed] [Google Scholar]
- Waterston R. H., Thomson J. N., Brenner S. Mutants with altered muscle structure of Caenorhabditis elegans. Dev Biol. 1980 Jun 15;77(2):271–302. doi: 10.1016/0012-1606(80)90475-3. [DOI] [PubMed] [Google Scholar]
- Wills N., Gesteland R. F., Karn J., Barnett L., Bolten S., Waterston R. H. The genes sup-7 X and sup-5 III of C. elegans suppress amber nonsense mutations via altered transfer RNA. Cell. 1983 Jun;33(2):575–583. doi: 10.1016/0092-8674(83)90438-5. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Young L. S., Takahashi N., Sprague K. U. Upstream sequences confer distinctive transcriptional properties on genes encoding silkgland-specific tRNAAla. Proc Natl Acad Sci U S A. 1986 Jan;83(2):374–378. doi: 10.1073/pnas.83.2.374. [DOI] [PMC free article] [PubMed] [Google Scholar]