Abstract
Human immunodeficiency virus type 1 (HIV-1) is completely dependent upon the Env protein to enter cells. The virus typically replicates in activated CD4+ T cells due to viral entry requirements for the CCR5 coreceptor and for high surface levels of the CD4 receptor. This is the case for the transmitted virus and for most of the virus sampled in the blood. Over the course of infection, the env gene can evolve to encode a protein with altered receptor and coreceptor usage allowing the virus to enter alternative host cells. In about 50% of HIV-1 infections, the viral population undergoes coreceptor switching, usually late in disease, allowing the virus to use CXCR4 to enter a different subset of CD4+ T cells. Neurocognitive disorders occur in about 10% of infections, also usually late in disease, but caused (ultimately) by viral replication in the brain either in CD4+ T cells or macrophage and/or microglia. Expanded host range is significantly intertwined with pathogenesis. Identification and characterization of such HIV-1 variants may be useful for early detection which would allow intervention to reduce viral pathogenesis in these alternative cell types.
Keywords: human immunodeficiency virus, HIV, envelope, env, gp160, gp120, gp41, tropism, macrophage, monocyte, receptor, CD4, coreceptor, CCR5, R5, CXCR4, X4, transmission, evolution, entry
Introduction
Env Protein and Gene
The Human Immunodeficiency Virus type 1 (HIV-1) env gene encodes the only surface-expressed viral protein Env, a glycoprotein of 160 kD (gp160), which is solely required for binding and entry into host cells. After translation, gp160 is cleaved into gp120 and gp41, which remain non-covalently linked to form a single subunit of a trimeric “spike” on the virion surface. The C-terminal subunit, gp41, contains a cytoplasmic domain (ultimately inside the viral membrane), a membrane spanning domain and an extracellular domain, which mediates the conformational change needed for fusion. The N-terminal subunit, gp120, is completely outside the viral membrane. Although the protein has a complex fold it can also be viewed as being linearly organized into five conserved regions (C1-C5) interspersed with five variable regions (V1-V5). The host receptor CD4 interacts with residues in the conserved regions of gp120 on either side of V4, and the coreceptor CCR5 interacts both with a GPGR/Q motif at the apex of the V3 loop and at its base. The exposed surface of the spike is dominated by two features: the variable regions of gp120, insinuating that high variability plays and important role in extracellular interactions; and a large number of carbohydrates that help mask the surface of the protein.
The gp120 coding domain of the env gene evolves faster (changing 1–2% per year) than any other region of the genome [1]. Although sequence variability is generally troublesome for maintaining required structures and functions, env variability is concentrated within discrete regions, which protects the general architecture of the protein. Much of env variability is driven by immune escape; however, sequence changes in gp120 can also alter interactions with host receptors. Over the course of infection, viral populations can evolve to use the CD4 receptor differently and to use alternative coreceptors allowing entry into alternative host cells.
Tropism and Entry Phenotype
The Env protein is an entry machine, built to bind to CD4, undergo a series of conformational changes, fuse the cell and viral membranes, and deliver the viral core to the cytoplasm of the cell. Given that most transmissions occur across mucosal surfaces, persistent replication takes place in lymphoid tissues, and disease manifestations are apparent in many tissues/organs, it is clear that the virus has ample opportunity to encounter many different cell types. Much of the research on the virus side of viral pathogenesis is focused on the Env protein and its role in entry. In order to understand this, we need to be able to accurately identify and characterize variants able to infect alternative target cells (i.e. expanded host range), including when and where these variants first occur, when they become prevalent and how this affects pathogenesis. More mechanistically, we need to know how viruses with expanded host range differ from the rest of the viral population in terms of receptor use, coreceptor use, physical changes in Env and the consequences of these changes.
HIV-1 Comes In (At Least) Three Colors, Not Two
For 25 years the majority view of HIV-1 has been that it comes in two colors, or rather entry phenotypes. The initial dichotomy was based on the observation that only some HIV-1 isolates could grow in transformed CD4+ T cell lines. Furthermore, since HIV-1 Env protein mediates fusion without the need for reduced pH this allows infected cells, with the Env protein on the surface of the cell, to fuse with uninfected cells creating syncytia and giving the appearance of a pathogenic virus. The isolates that did not grow in the transformed cell lines could not cause syncytia, thus setting up the early classification of syncytium-inducing (SI) and nonsyncytium-inducing (NSI) viruses [2]. Some NSI viruses could also infect macrophages. The NSI and SI designations have changed with the discovery of the coreceptor, with SI being equal to CXCR4-using (X4) and with NSI being equal to CCR5-using (R5), and all of this can be traced to the fact that most transformed CD4+ T cell lines do not express CCR5 [3]. Thus, the legacy groupings are NSI/R5/macrophage-tropic and SI/X4 viruses.
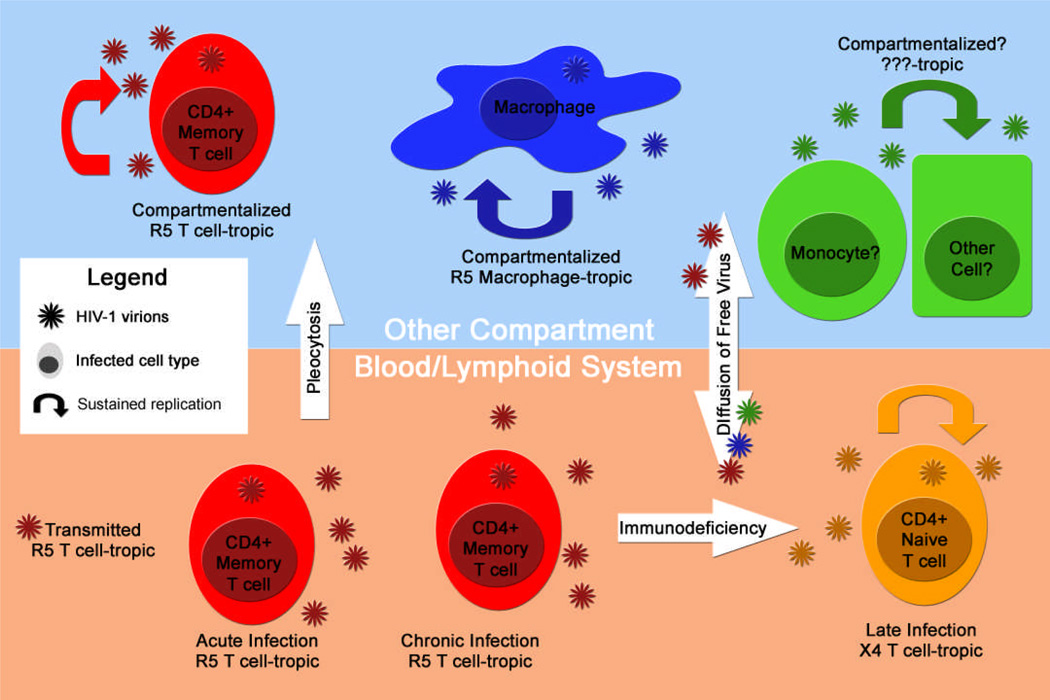
However, these two groups fail to capture the important concept that only a small (but important) fraction of R5 viruses are able to infect macrophages, necessitating the use of three groupings: R5 T cell-tropic, R5 macrophage-tropic, and X4. The distinction between T cell-tropic and macrophage-tropic R5 viruses is not a small point, because gaining the ability to infect macrophages represents an important but poorly understood part of HIV-1 pathogenesis. The salient features of these three groups are described in the following section then discussed in greater detail below in the context of evolution over the disease course of an infected person. Figure 1 provides a framework for the main topics covered in this review.
Figure 1. Compartmentalization and Replication in Alternative Cell Populations.
Compartmentalization has been frequently detected in the CNS of people with HIV-associated neurological disorders and there is evidence that compartmentalization can occur in other anatomical sites. Although a contribution of viral replication to pathogenesis in the CNS is well established, effects in other compartments will require further study.
R5 T cell-tropic Virus
HIV-1 spends most of its time replicating in activated CD4+ T cells [4–6]. Activated T cells are activated in the context of the immune response, but also are metabolically active making viral replication more efficient. These cells are infected by the form of the virus that is transmitted in the vast majority of cases, that persists in the host for years, that most often causes immunodeficiency, and that is sufficient to account for most of what we think of as the HIV-1 epidemic. The entry phenotype of this virus is clearly defined by the requirement for CCR5 to allow entry and also the need for high levels of CD4 on the surface of the cell as found on activated CD4+ T cells [7, 8].
R5 Macrophage-Tropic Virus
Some isolates of HIV-1 that can infect CD4+ T cells can also infect macrophages (i.e. are M-tropic). A commonly used viral clone, Ba-L, was the first M-tropic virus isolate, in this case from macrophages recovered from a bronchial/alveolar lavage [9]. Although M-tropic viruses are most often found in the central nervous system (CNS) [10–12], they have also been observed in the blood [10] and other compartments [10, 12]. In one case, the M-tropic virus in the blood was not derived from the CNS, suggesting alternative tissue sites where macrophage-tropic viruses can evolve beyond the CNS [13•].
X4 T Cell-Tropic Virus
As noted above, the evolution of variants that can use CXCR4 is a distinctive and consistent feature of the viral population in a significant fraction of those infected with the virus. There are continuing discussions as to whether these variants are selected against at the time of transmission, whether they can survive only in an immunodeficient host, whether they contribute to or are a marker for more rapid disease progression, and whether different subtypes have a different propensity to evolve these variants. Thus, while we know a great deal about the evolution of these variants and their properties, we cannot yet place them with confidence in the context of viral pathogenesis.
env Gene Evolution and Phenotypic Variation
The Transmitted Virus
Much attention is now being paid to the nature of the transmitted virus and, in particular, the properties of the Env protein associated with transmission. At one end of the debate is the idea that the transmitted virus is the random but lucky winner of a group of viruses deposited at a mucosal site. At the other end is the idea that only viruses with certain properties are transmitted; in this case, information about the transmitted virus would provide insight into the process of transmission and could be important to the development of prevention strategies such as vaccines and microbicides. Early studies of the virus present in people with acute infection demonstrated that there is a significant genetic bottleneck associated with sexual transmission. However, an important conceptual insight came with the understanding that, in a majority of cases, infection is initiated with a single variant [14•, 15].
There is still uncertainty about whether multiple variants are transmitted but a single variant is detected systemically, or if a single variant establishes the initial infection at the site of transmission and is the progenitor to all of the virus in the body. Sampling of the blood suggests a single variant is transmitted [16]; however, it is possible that there are abortive infections at the mucosa that do not contribute to the systemic viral population. If true, then there should be examples of abortive infections without systemic infection. In such a scenario, the level of viral replication must be low, since we do not observe people who are seropositive for HIV-1 but not infected (in contrast to HSV-2, for example, which induces seropositivity after a local infection in the mucosa). A subset of people are infected with multiple variants (i.e. two or more viruses from the same donor), which may be related to the route of infection or other cofactors that affect the frequency of transmission [17]. An important question is: does the extra genetic diversity associated with the transmission of multiple variants affect the rate of disease progression, a phenomenon that has been associated with dual infection from two sources [18]?
The severe transmission bottleneck begs the question of whether there is something consistently special about the single transmitted virus. If there is something special, then it is possible that this variant is selected at the donor site of transmission, during the transmission process, or after transmission. There can be local populations of virus at the donor site, i.e. compartmentalized virus in the seminal tract or the vaginal/cervix area (see [19] as an example), although the biological properties of these compartmentalized viruses have not been explored. At present, there is no evidence that these compartmentalized variants are favored in transmission.
Several studies have observed that the transmitted virus is underglycosylated [20, 21], and in our own work we have found that the virus in acutely infected men is more underglycosylated than the virus in acutely infected women (Ping et al., manuscript in preparation). This observation suggests that there is selective pressure in the transmission process itself, i.e. female-to-male transmission favors underglycosylated Env proteins. We observed a similar trend in intrapartum transmission where infants are also exposed to cervico-vaginal mucus [22]. Finally, it has been proposed that there is selective pressure after transmission either in the form of neutralization sensitive virus that replicates more quickly [21] or virus that binds to α4β7 integrin to promote attachment to cells in the mucosa and the gut-associated lymphoid tissue (GALT) [23].
To date, these hypotheses have been explored with a relatively small number of virus isolates and the role of these features in transmission will likely be examined in greater detail soon. It is worth repeating that even using a deep sequencing approach it has not been possible to find heterogeneity in the blood beyond the transmitted virus [16]. Thus, if there is selection in the new host among multiply transmitted variants, it must take place completely within tissue that is not releasing significant amounts of virus into the blood.
Early Evolution of the env Gene
The selective pressure on the env gene that is unique among viral gene products is that the Env protein is the only target of neutralizing antibodies. Thus, the host antibody response drives evolution with the initial autologous response to the transmitted virus and with sequential responses to the escape variants [24]. Important targets for the host antibodies are in the variable loops V1, V2, V4, and V5; to a certain extent this conclusion is inferred based on the extreme variability of these regions, since autologous responses to Env have been mapped in only a limited number of cases [25]. In addition, env is also under selection from cell-mediated immunity and this also results in escape mutations.
Because mutations are random, it is not that the regions encoding the variable loops are preferentially mutated, rather a greater fraction of the mutations in these regions give viable protein products that can be tested for escape. Evolution to avoid antibodies takes place on at least three levels. First, the Env protein is in a conformation that makes it neutralization resistant relative to other conformations the protein can assume. Amino acid changes between V1 and V2 (and perhaps elsewhere) control these conformation states and can expose the V3 loop and the coreceptor binding site to neutralization [26–28]. Second, escape occurs through point mutations that affect immune recognition [25]. In this aspect, the variable loops seem to serve the role of decoy, i.e. targets for host antibodies that easily evolve to allow escape. Third, the heavily glycosylated surface of Env can shield the protein surface from exposure to antibodies [29, 30]. One feature of these variable regions that may facilitate mutations is the concentration of specific trinucleotides that could enhance duplications or deletions through mispairing during viral DNA synthesis. In addition, these trinucleotides include the codons needed to encode N-linked glycosylation sites [31, 32]. Thus, HIV-1 Env mediates an intricate and delicate balance between the function of cell entry and evasion of antibody selection.
Evolution of X4 Variants
X4 virus was initially perceived as a more pathogenic variant that appeared late in disease. This idea came from the observation that viruses isolated from some late stage subjects could grow in transformed T cell lines and cause syncytia (SI viruses), a striking feature of the culture that gave the impression of pathogenesis. However, these viruses are not necessarily more pathogenic; viruses that use CCR5 (formerly NSI viruses) can grow in transformed T cell lines and cause syncytia if CCR5 is expressed.
The ability to replicate in transformed T cell lines did provide a useful assay for detecting X4 variants, which demonstrated the increased presence of X4 virus with decreasing CD4+ T cell count in cohorts of infected people [33]. These results were subsequently confirmed by entry assays that distinguished between R5 and X4 viruses [34]. There are two important questions about X4 viruses that remain unresolved. First, do X4 viruses cause more rapid disease progression or are they just a marker for increased immunodeficiency? The second question has to do with viruses that can use both CCR5 and CXCR4 to enter cells, known as dual-tropic viruses. Do these viruses continue to use CCR5 in vivo with dual tropism an important part of their biology, or is the residual CCR5 usage a mere vestige of ancestry and only CXCR4 usage is biologically relevant? Most X4 isolates are dual tropic, so this issue is relevant to our understanding of a majority of X4 variants. An unexplained observation is that X4 viruses appear to be more common in people who have been treated, but failed, therapy – although this is in part explained by low nadir CD4+ T cell counts prior to initiating therapy [35].
Evolution of the ability to infect using CXCR4 should expand the potential target cells available to X4 viruses. It seems unlikely that X4 variants would evolve to infect the same cells as R5 viruses; thus, the expectation is that CD4+ T cells expressing CXCR4, but not CCR5, are the targets of X4 viruses. This reasoning is consistent with the observations that memory CD4+ T cells express CCR5 and are the predominant cell infected [4–6], while naïve cells express very little CCR5 but both cell types express CXCR4 [36, 37]. R5 and X4 viruses can be separately cultured from populations of memory and naïve T cells, respectively [38]. However, the cell type where X4 variants first appear and if they compete with or outgrow R5 viruses is still difficult to assess [39, 40]. There is evidence that when both R5 and X4 viruses are present in a person, they are both replicating in short-lived cells, presumably activated CD4+ T cells [41].
Even before the HIV-1 coreceptors had been identified, it was known that sequence changes in the V3 loop were major determinants defining the properties of late stage viruses [42]. The most consistent observation of sequence evolution is the appearance of basic amino acid substitutions (Lys and Arg), at positions 11, 24, and 25 of the 35-amino-acid-long V3 loop [43, 44]. Overall, diversity is higher in V3 when these basic amino acid substitutions are present, indicating that these substitutions are part of a more complex evolutionary pathway. These basic amino acid substitutions and increased diversity have made it possible to develop bioinformatic tools that predict coreceptor usage of viruses based on V3 loop sequence [45, 46].
Evolution to use CXCR4 likely includes more than just sequence changes in V3. Using a bioinformatics approach, it has been possible to identify sequence changes outside of V3 that are statistically linked to changes within V3, with the strongest association at position 440 in the Env protein, a position lying directly under the V3 loop in the gp120 crystal structure [47]. Molecular recombinants have been used to demonstrate the contribution of other regions of Env to entry phenotype [48]. However, the interpretation of these results is complicated for several reasons. First, the ability to enter cells using CXCR4 ranges on a continuous scale from nil to completely specific, meaning that in between are viruses that use both CCR5 and CXCR4 with every ratio of efficiency. Does a little CXCR4 tropism indicate the initial evolution of an X4 virus or do highly sensitive entry assays identify "sloppy" viruses that have some capacity to use CXCR4 but not in a biologically meaningful way? By extension, what is the significance of mutations that confer only a low level of CXCR4 tropism? Seeing mutations accumulate over time in Env genes from strongly X4 viruses provides the clearest evidence that these sequence changes are relevant to the coreceptor switch. However, we have too few examples of X4 viruses to create robust analyses to find the sets of mutations that are likely to define a number of evolutionary pathways.
An important, but difficult, question that speaks to the mechanism of X4 evolution is the propensity of viruses from different subtypes to undergo the coreceptor switch. A provocative observation was that X4 viruses were present in some subjects with AIDS but not others and this correlated with the subtype of the virus [49]. We have the most data for people infected with subtype B, where about 50% of those infected develop an X4 virus [33]. Subtype D, which is closely related to subtype B, appears to have an even greater propensity to evolve X4 variants [50]. Conversely, subtype C seems less inclined (but not unable) to evolve X4 variants [49]. In general, X4 variants appear late in disease. If we assume the host can suppress the replication of these variants until the time of immunodeficiency, and we assume different infected populations do not vary in a way that enhances or suppresses the ability of these viruses to evolve, then what remains is differences in the virus. Since all of these viruses can evolve X4 variants, perhaps it is the evolutionary distance the virus has to travel that defines the differences, i.e. subtype D viruses have the shortest distance followed by B then C.
Additional insights have been gained from studying R5 and X4 viruses in the macaque animal model. R5 Simian immunodeficiency virus (SIV) isolates only rarely undergo coreceptor switching [51]. However, there are several examples where SIV engineered to have an HIV-1 env gene, known as a SHIV, undergoes a coreceptor switch from R5 to X4 [52, 53]. The switch takes place in animals that are rapid progressors due to a poor initial immune response. The initial evolution of the X4 variant has been tracked to secondary lymphoid tissue [54], which is similar to an observation made in the humanized mouse model [26]. In animals infected with an X4 variant, there is rapid depletion of naïve CD4+ T cells followed by loss of memory cells, consistent with the idea that X4 viruses preferentially target a cell type different from R5 viruses [55, 56].
Evolution of M-tropic viruses
"Beauty is in the eye of the beholder" (a paraphrase of Plato). The definition of a macrophage-tropic virus is problematic. Monocyte-derived macrophages (MDM) from different donors can be very different in their ability to allow entry of a virus. Thus, a virus that appears macrophage-tropic on one preparation of MDMs may not appear to be macrophage-tropic on another preparation. Also, viruses have a continuum of entry phenotypes making it hard to draw a line that demarcates macrophage- and nonmacrophage-tropic viruses. The difficulty, as with defining X4 viruses, is deciding when the entry phenotype measured in the laboratory represents a biologically meaningful property for the virus in vivo. A major challenge for this field is accurately placing the phenotype of the ability to infect MDMs into the proper biological context.
Macrophage-tropic (M-tropic) viruses have most often been reported in the brain of HIV-infected individuals with neurocognitive disorders [11, 57, 58] and this is one place where the evolution of M-tropic virus is likely to play a direct role in viral pathogenesis. M-tropic viruses can also be detected in the cerebrospinal fluid (CSF) and, in one case, it was possible to detect this lineage before diagnosis of neurocognitive disorder [13•]. In the brain, M-tropic viruses may be replicating in perivascular macrophages or microglia, which are macrophage-like cells residing in the parenchyma of the brain. In the absence of pleocytosis, T cells are uncommon in the CNS; however, pleocytosis is associated with replication of R5 T cell-tropic virus within the CNS [13•]. An important tool in assessing localized viral replication is that viral populations in the CNS can be genetically diverse and genetically distinct from populations in the blood (i.e. compartmentalized) [59–62]. However, not all potential compartments (i.e. compartments with high concentrations of macrophage or macrophage-like cells) have been systematically explored (discussed below), and compartmentalized M-tropic virus has been detected in the semen of one person [10]. As we develop a better understanding of how to define macrophage-tropic viruses it will be worthwhile to explore other compartments with known HIV-associated pathogenesis (e.g. liver).
Currently, M-tropic variants are isolated from autopsy brain tissue or from the CSF, which can be used to sample virus longitudinally from living patients over the course of infection and disease. Later during infection, M-tropic virus may become detectable as a minor variant in the blood [63•, 64], though the anatomical origins of these variants have not been established. M-tropism has been associated predominantly with the ability to use low cell surface densities of CD4, but also with changes in CCR5 usage (currently under dispute with some groups observing ability to use lower levels of CCR5 [65, 66] and other observing no change in CCR5 usage [67, 68], enhanced fusogenicity [69], sensitivity to antibody neutralization [70–72] and an altered – specifically “open” – conformation (which may be a precursor for changes in both CD4 usage and neutralization sensitivity [65, 73•].
As noted above, the surrogate test for M-tropism in vivo is the ability of virus to infect MDM. An alternative method is to use cell lines engineered to have different levels of CD4 and measure the dependence of infectivity on CD4 density. The latest iteration of this approach is a novel cell line, Affinofile cells, developed by Benhur Lee [74], which have independently inducible CD4 and CCR5 expression. Affinofile cells are highly consistent in receptor expression and resulting infectivity; receptors can be induced to a wide range of densities, including normal physiological levels found on macrophages and T cells and beyond (allowing titration of receptor/coreceptor usage). Most importantly, these cells allow isolation of the entry requirement variables without potentially confounding post-entry factors.
Although there are likely multiple adaptations required to infect macrophages, M-tropic variants are largely defined by their ability to infect cells expressing low levels of CD4 [67], which reflects the relatively low CD4 density on the surface of macrophages as compared to activated memory T cells [37]. The search for sequence changes in gp120 to allow binding and entry into cells with lower densities of CD4 has focused on regions directly involved in receptor/coreceptor interactions: around the CD4 binding residues [57, 58, 70, 75] and around the V3 loop (and coreceptor binding residues) of gp120 [12, 76]. Others, however, have proposed that mutations in the V1-V2 loop [77, 78] or in gp41 [66] can induce significant changes in gp120 conformation, which may cause greater exposure of receptor/coreceptor binding sites. Although most M-tropic variants are R5 viruses, there is evidence for the existence of M-tropic X4 virus [79, 80], though their prevalence and role in pathogenesis is not yet clear.
Defining genetic determinants of macrophage tropism is essential to being able to screen for these viruses in mixtures and in large numbers of biological samples, as the totality of the effort required to identify these viruses by entry phenotype is cumbersome and quickly becomes limiting. Unfortunately, there is little overlap in the mutations identified by different groups to date (see Table 1). This discord could be explained by the time at which viruses were isolated (before appearance of neurocognitive symptoms or late in disease, including autopsy samples); a potentially diverse number of evolutionary pathways to reach M-tropism (open conformation to increase binding site exposure, greater affinity for receptor/coreceptor within the binding site, increased fusion efficiency to counteract the loss in infectivity wrought by low CD4 densities, etc.); or by different changes within a region of gp120 that result in a similar phenotype, but do not align precisely in sequence analysis. If essential or signature mutations can be identified for M-tropism, then HIV-infected individuals could be screened for M-tropic variants – by deep sequencing, if necessary, to deal with mixtures of M-tropic and T-tropic variants – to inform care and potentially reduce neurocognitive pathogenesis by early intervention.
Table 1.
Reported Amino Acid Changes Associated With Macrophage-Tropism
| Env position | |||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| V1 | V2 | C2 | V3 | C3 | V4 | ||||||||
| Year | First Author | Tropism | 153 | 178 | 188 | 283 | 308 | 317 | 362 | 363 | 364 | 373 | 386 |
| Subtype B Consensus | T cell | E | K | T | T | H | F | N | Q | S | M | N | |
| Observed Changes | Mac. | G | E | N | N | N | L | K | P | S | K | D | |
| 2011 | Musich [77] | T cell | E | ||||||||||
| Mac. | G | ||||||||||||
| 2005 | Walter [78] | T cell | K | T | |||||||||
| Mac | E | N | |||||||||||
| 2006 | Peters [11] | T cell | T | ||||||||||
| Mac | N | ||||||||||||
| 2006 | Dunfee [57] | T cell | T/I/V | ||||||||||
| Mac | N | ||||||||||||
| 2007 | Dunfee [58] | T cell | N | ||||||||||
| Mac | D | ||||||||||||
| 2007 | Thomas [12] | T cell | H/T | ||||||||||
| Mac | P | ||||||||||||
| 2007 | Sterjovski [69] | T cell | T | ||||||||||
| Mac | N | ||||||||||||
| 2008 | Duenas-Decamp [72] | T cell | R | N | |||||||||
| Mac | K | D | |||||||||||
| 2009 | Duenas-Decamp [70] | T cell | I/T | H | F | N | Q | P | |||||
| Mac | N | N | L | K | P | S | |||||||
| 2010 | Duenas-Decamp [75] | T cell | N | ||||||||||
| Mac | D | ||||||||||||
Infection of Monocytes
Given the clear evidence for infection of macrophages in vivo there is also interest in whether the precursors of macrophages, i.e. monocytes, can be infected by HIV-1. Monocytes are a diverse cell population that can be divided into three subsets - classical, intermediate and nonclassical. These subsets differ in size and shape [81, 82], expression of surface markers [83, 84], susceptibility to HIV infection [83, 85], cytokine production [81] and role in the immune system [81, 82]. After being produced from common myeloid progenitor cells in the bone marrow [86], they enter the blood as classical monocytes and can then differentiate into intermediate and then into nonclassical monocytes [87]. The three monocyte subsets are most easily separated by expression of CD14 and CD16, with classical monocytes expressing high levels of CD14 and lacking CD16 (CD14++, CD16−), intermediate monocytes expressing high levels of CD14 and low levels of CD16 (CD14++, CD16+) and nonclassical monocytes expressing low levels of CD14 and high levels of CD16 (CD14+, CD16++)[84, 87]. In healthy donors, approximately 85% of monocytes fall into the classical subset, 5% are intermediate and 10% are nonclassical [84].
Multiple in vitro studies have shown that freshly isolated monocytes are not productively infected by HIV-1, but can be productively infected after they are allowed to differentiate for at least one day and are increasingly susceptible to infection as they differentiate into MDMs [88, 89]. The resistance of monocytes to HIV-1 infection is likely due to blocks at multiple points in the virus life cycle. A block at entry could occur if densities of the receptor (CD4) or coreceptors (CCR5 or CXCR4) were very low on monocytes. This is probably not the case for CD4, which is expressed at appreciable levels on most monocytes [83], albeit at much lower levels than on T cells [37]. In contrast, CCR5 and/or CXCR4 are only expressed by a minority of monocytes and are expressed at low levels [83]. It has been observed that CCR5 density on monocytes and susceptibly to HIV-1 infection are correlated with the amount of time that monocytes spend in culture, thus fueling speculation that CCR5 levels are too low to facilitate virus entry into freshly isolated monocytes [88]. Multiple blocks to reverse transcription [90, 91] and transcription of the proviral genome [92] have been observed in monocytes.
An alternative model is that HIV-1 infection of monocytes is not blocked, but rather the mechanisms described above simply slow the virus lifecycle [93]. Considering that monocytes persist in the blood for only a few days [94] before dying or migrating into tissues and differentiating into DCs or macrophage, this alternative model suggests that infected monocytes are likely to differentiate before producing virus. While many questions remain about putative blocks to monocyte infection, it appears clear that HIV replication in monocytes is unlikely to contribute substantially to viral loads. However, given the ability of monocytes to enter tissues, infected monocytes could provide an important mechanism to disseminate virus to sites where independent replicating populations could be established.
One limitation of most of these in vitro studies is that they have not separately examined infection of the three monocyte subsets. Since approximately 85% of monocytes in the blood are of the classical subset [84], most studies to date have focused on infection of this subset. This is significant given that classical monocytes are the most undifferentiated subset, contain the lowest percentage of CCR5+ cells and are the least susceptible to infection in vitro [83]. Given the extensive evidence that differentiation increases susceptibility to infection of monocytes [88, 89], and the observation that monocytes expressing CD16 are more susceptible to infection [83], intermediate and nonclassical monocytes should be more susceptible to infection than classical monocytes.
In contrast to the in vitro studies examining whether HIV-1 can productively infect monocytes, multiple studies have isolated monocytes that contain HIV-1 DNA [63•, 95] and replication competent virus [96•], thus indicating that monocytes are infected in vivo. Analysis of env sequences isolated from these cells showed that monocytes can harbor viruses that appear distinct from those in the blood [95]. Further phenotypic analysis of these viruses revealed that they were all capable of using CCR5 as their coreceptor. Some were also able to use alternative coreceptors (CXCR4, CCR1, CCR3 and GPR15) and, while all of the monocyte derived viruses were capable of replicating in macrophage in vitro, only a subset of them could replicate in T cells [63•]. Given the short half-life of monocytes in the blood and the fact that the monocytes most susceptible to infection are likely to be near the end of this period, it is difficult to understand how a stable, diverse population of monocyte-tropic viruses could be maintained. One possibility is that monocytes are being infected by macrophage-tropic viruses replicating in tissue macrophage [63•]. Reports of monocytes being infected in individuals who are on otherwise suppressive therapy also present a challenge to understanding the virus life cycle in this enigmatic cell type.
Use of Alternative Co-receptors
People who carry two alleles of the delta32 deletion of the CCR5 gene are largely resistant to infection by HIV-1, consistent with the vast majority of infections being initiated with an R5 virus. In some isolates it is possible to show promiscuity in the ability to use different coreceptors in entry assays, but the biological significance of these observations is unknown, i.e. it is not clear if these alternative coreceptors are actually used in vivo. However, the potential to use an alternative coreceptor has clearly been demonstrated in the case of SIVsm infection of sooty mangabeys, which can also have an inactivating mutation in the CCR5 gene. Infection by SIVsm in these animals cannot use CCR5 but may use GPR15 and/or CXCR6 as an alternative coreceptor [97].
HIV infection of CD4–negative cells
It has long been thought that HIV-1 tropism is primarily limited by expression of CD4 and CXCR4 or CCR5; thus it is surprising that multiple studies have reported that HIV-1 can infect cells lacking these receptors [98–102]. The fact that most primary isolates require CD4 to infect [103] suggests that evolving the ability to be CD4-independent comes with an evolutionary cost. This cost could be in the form of increased sensitivity to neutralization [99, 103] or the ability to enter cells that do not support efficient viral replication, but could impact the range of viral pathogenesis. A possible example of CD4-indepence evolving in vivo is renal-tropic HIV which is thought to infect kidney epithelial cells. These cells lack CD4 [102, 104], CCR5 and CXCR4 [104]. Infection of these CD4-negative cells is suggested by studies showing that patients with HIV-associated renal diseases often have kidney epithelial cells containing HIV-1 mRNA [105] and DNA [102]. Surprisingly, a study of two patients with HIV-associated nephropathy found that that viral DNA isolated from kidney epithelial cells forms a lineage that is evolutionarily separate from that of viral DNA in the blood [102]. This result suggests that there is an independent population of viruses infecting kidney epithelial cells; however, this conclusion was obtained with a small number of kidney-derived gp120 sequences and additional studies are need to confirm this finding.
Since many viruses have evolved the ability to enter their hosts using a single receptor, it is curious that HIV-1 requires two receptors (CD4 and CCR5 or CXCR4), thus fueling speculation that this dual receptors system confers an evolutionary advantage. One possibility is that having CD4 as a receptor allows the Env protein to be in one conformation prior to binding and a different conformation (and immunogenic state) after binding when the fusion mechanisms of Env become active. It is clear that the regions of Env that interact with CD4 are distinct from those that interact with CCR5 and also distinct from the actual fusion machinery. Thus it is possible to imagine a form of Env that is CD4-independent, i.e. in the alternative or bound conformation that allows interaction with CCR5 and subsequent fusion. It has been possible to select for such variants in culture; in vitro studies have shown that HIV-1 can evolve CD4-independence [98, 100, 101]. The exact nature of these viruses is still being worked out since there is an important distinction between viruses that can use very low levels of CD4 versus those that are truly CD4 independent. The mutations that confer CD4 independence include changes in C2, C3 and V3 that are thought to alter coreceptor binding [98] and changes in gp41 with unknown effects on coreceptor binding [66]. This concept of alternative conformations has also carried over into the idea that late in disease there may be relaxed antibody selection on Env allowing it to evolve an alternative conformation for the purpose of using CD4 more efficiently; the corollary of this model is that the alternative conformation may also potentiate the evolution to use CXCR4 or the evolution to use low levels of CD4 and in this way expand the host range, and the pathogenic potential, of HIV-1.
Conclusions
HIV-1 env mutates rapidly under evolutionary pressure from multiple sources such that the encoded Env protein displays a delicate balance between continual immune evasion and retention of the vital function of host cell entry. There are at least three different colors or host range variants of HIV-1 as determined by the Env protein. The first is the R5 T cell-tropic virus, which is typically the single transmitted virus and also the virus most commonly found throughout chronic infection. The second is the R5 macrophage-tropic virus, which is most often reported in the CNS of people with HIV-associated neurocognitive disorders – although macrophage-tropic viruses with a non-CNS (and unknown) origin have also been detected. The third is the X4 virus, which frequently emerges in late disease and is associated with advanced immunodeficiency. While the mechanisms and evolutionary “motivations” are not fully clear, evolving altered receptor and coreceptor usage allows the virus to infect new cell types with potentially serious pathogenic consequences.
Acknowledgements
We would like to thank the NIH for funding that supports our ongoing research in this field. We also apologize to our colleagues whose work we did not cite due to editorial limitations.
Footnotes
The final publication is available at link.springer.com: http://link.springer.com/article/10.1007/s11904-011-0107-3/fulltext.html
Contributor Information
Kathryn Twigg Arrildt, Email: arrildt@email.unc.edu.
Sarah Beth Joseph, Email: sbjoseph@email.unc.edu.
Ronald Swanstrom, Email: risunc@med.unc.edu.
References
- 1.Leitner T, Kumar S, Albert J. Tempo and mode of nucleotide substitutions in gag and env gene fragments in human immunodeficiency virus type 1 populations with a known transmission history. J Virol. 1997;71:4761–4770. doi: 10.1128/jvi.71.6.4761-4770.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Asjo B, Morfeldt-Manson L, Albert J, et al. Replicative capacity of human immunodeficiency virus from patients with varying severity of HIV infection. Lancet. 1986;2:660–662. [PubMed] [Google Scholar]
- 3.Berger EA, Murphy PM, Farber JM. Chemokine receptors as HIV-1 coreceptors: roles in viral entry, tropism, and disease. Annu Rev Immunol. 1999;17:657–700. doi: 10.1146/annurev.immunol.17.1.657. [DOI] [PubMed] [Google Scholar]
- 4.Douek DC, Brenchley JM, Betts MR, et al. HIV preferentially infects HIVspecific CD4+ T cells. Nature. 2002;417:95–98. doi: 10.1038/417095a. [DOI] [PubMed] [Google Scholar]
- 5.Brenchley JM, Hill BJ, Ambrozak DR, et al. T-cell subsets that harbor human immunodeficiency virus (HIV) in vivo: implications for HIV pathogenesis. J Virol. 2004;78:1160–1168. doi: 10.1128/JVI.78.3.1160-1168.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Sleasman JW, Aleixo LF, Morton A, et al. CD4+ memory T cells are the predominant population of HIV-1-infected lymphocytes in neonates and children. AIDS. 1996;10:1477–1484. doi: 10.1097/00002030-199611000-00004. [DOI] [PubMed] [Google Scholar]
- 7.Alexander M, Lynch R, Mulenga J, et al. Donor and recipient envs from heterosexual human immunodeficiency virus subtype C transmission pairs require high receptor levels for entry. J Virol. 2010;84:4100–4104. doi: 10.1128/JVI.02068-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Isaacman-Beck J, Hermann EA, Yi Y, et al. Heterosexual transmission of human immunodeficiency virus type 1 subtype C: Macrophage tropism, alternative coreceptor use, and the molecular anatomy of CCR5 utilization. J Virol. 2009;83:8208–8220. doi: 10.1128/JVI.00296-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gartner S, Markovits P, Markovitz DM, et al. The role of mononuclear phagocytes in HTLV-III LAV infection. Science. 1986;233:215–219. doi: 10.1126/science.3014648. [DOI] [PubMed] [Google Scholar]
- 10.Brown RJP, Peters PJ, Caron C, et al. Intercompartmental Recombination of HIV-1 Contributes to env Intrahost Diversity and Modulates Viral Tropism and Sensitivity to Entry Inhibitors. J Virol. 2011;85:6024–6037. doi: 10.1128/JVI.00131-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Peters PJ, Sullivan WM, Duenas-Decamp MJ, et al. Non-macrophage-tropic human immunodeficiency virus type 1 R5 envelopes predominate in blood, lymph nodes, and semen: implications for transmission and pathogenesis. J Virol. 2006;80:6324–6332. doi: 10.1128/JVI.02328-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Thomas ER, Dunfee RL, Stanton J, et al. Macrophage entry mediated by HIV Envs from brain and lymphoid tissues is determined by the capacity to use low CD4 levels and overall efficiency of fusion. Virology. 2007;360:105–119. doi: 10.1016/j.virol.2006.09.036. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Schnell G, Joseph S, Spudich SS, et al. Two classes of viral encephalitis contribute to the development of HIV-1-associated dementia. PLoS Pathog. In press.. This paper demonstrates the production of macrophage-tropic virus from a long-lived cell in a subset of subjects with HIV-associated dementia.
- 14. Keele BF, Giorgi EE, Salazar-Gonzalez JF, et al. Identification and characterization of transmitted and early founder virus envelopes in primary HIV-1 infection. Proc Natl Acad Sci U S A. 2008;105:7552–7557. doi: 10.1073/pnas.0802203105.. This paper demonstrates that in most transmissions of HIV-1, the systemic infection is derived from a single viral genome.
- 15.Abrahams MR, Anderson JA, Giorgi EE, et al. Quantitating the multiplicity of infection with human immunodeficiency virus type 1 subtype C reveals a non-poisson distribution of transmitted variants. J Virol. 2009;83:3556–3567. doi: 10.1128/JVI.02132-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fischer W, Ganusov VV, Giorgi EE, et al. Transmission of single HIV-1 genomes and dynamics of early immune escape revealed by ultra-deep sequencing. PLoS One. 2010;5:e12303. doi: 10.1371/journal.pone.0012303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Li H, Bar KJ, Wang S, et al. High Multiplicity Infection by HIV-1 in Men Who Have Sex with Men. PLoS Pathog. 2010;6:e1000890. doi: 10.1371/journal.ppat.1000890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gottlieb GS, Nickle DC, Jensen MA, et al. Dual HIV-1 infection associated with rapid disease progression. Lancet. 2004;363:619–622. doi: 10.1016/S0140-6736(04)15596-7. [DOI] [PubMed] [Google Scholar]
- 19.Anderson JA, Ping LH, Dibben O, et al. HIV-1 Populations in Semen Arise through Multiple Mechanisms. PLoS Pathog. 2010;6:e1001053. doi: 10.1371/journal.ppat.1001053. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chohan B, Lang D, Sagar M, et al. Selection for human immunodeficiency virus type 1 envelope glycosylation variants with shorter V1-V2 loop sequences occurs during transmission of certain genetic subtypes and may impact viral RNA levels. J Virol. 2005;79:6528–6531. doi: 10.1128/JVI.79.10.6528-6531.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Derdeyn CA, Decker JM, Bibollet-Ruche F, et al. Envelope-constrained neutralization-sensitive HIV-1 after heterosexual transmission. Science. 2004;303:2019–2022. doi: 10.1126/science.1093137. [DOI] [PubMed] [Google Scholar]
- 22.Russell ES, Kwiek JJ, Keys J, et al. The Genetic Bottleneck in Vertical Transmission of Subtype C HIV-1 Is Not Driven by Selection of Especially Neutralization Resistant Virus from the Maternal Viral Population. J Virol. 2011:8253–8262. doi: 10.1128/JVI.00197-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nawaz F, Cicala C, Van Ryk D, et al. The genotype of early-transmitting HIV gp120s promotes alphabeta-reactivity, revealing alphabetaCD4+ T cells as key targets in mucosal transmission. PLoS Pathog. 2011;7:e1001301. doi: 10.1371/journal.ppat.1001301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Richman DD, Wrin T, Little SJ, Petropoulos CJ. Rapid evolution of the neutralizing antibody response to HIV type 1 infection. Proc Natl Acad Sci U S A. 2003;100:4144–4149. doi: 10.1073/pnas.0630530100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Moore PL, Gray ES, Morris L. Specificity of the autologous neutralizing antibody response. Curr Opin HIV AIDS. 2009;4:358–363. doi: 10.1097/COH.0b013e32832ea7e8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Ince WL, Zhang L, Jiang Q, et al. Evolution of the HIV-1 env gene in the Rag2−/− gammaC−/− humanized mouse model. J Virol. 2010;84:2740–2752. doi: 10.1128/JVI.02180-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Pugach P, Kuhmann SE, Taylor J, et al. The prolonged culture of human immunodeficiency virus type 1 in primary lymphocytes increases its sensitivity to neutralization by soluble CD4. Virology. 2004;321:8–22. doi: 10.1016/j.virol.2003.12.012. [DOI] [PubMed] [Google Scholar]
- 28.Shibata J, Yoshimura K, Honda A, et al. Impact of V2 mutations on escape from a potent neutralizing anti-V3 monoclonal antibody during in vitro selection of a primary human immunodeficiency virus type 1 isolate. J Virol. 2007;81:3757–3768. doi: 10.1128/JVI.01544-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Binley JM, Ban YE, Crooks ET, et al. Role of complex carbohydrates in human immunodeficiency virus type 1 infection and resistance to antibody neutralization. J Virol. 2010;84:5637–5655. doi: 10.1128/JVI.00105-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Wei X, Decker JM, Wang S, et al. Antibody neutralization and escape by HIV-1. Nature. 2003;422:307–312. doi: 10.1038/nature01470. [DOI] [PubMed] [Google Scholar]
- 31.Bosch ML, Andeweg AC, Schipper R, Kenter M. Insertion of N-linked glycosylation sites in the variable regions of the human immunodeficiency virus type 1 surface glycoprotein through AAT triplet reiteration. J Virol. 1994;68:7566–7569. doi: 10.1128/jvi.68.11.7566-7569.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kitrinos KM, Hoffman NG, Nelson JA, Swanstrom R. Turnover of env variable region 1 and 2 genotypes in subjects with late-stage human immunodeficiency virus type 1 infection. J Virol. 2003;77:6811–6822. doi: 10.1128/JVI.77.12.6811-6822.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Koot M, Keet IP, Vos AH, et al. Prognostic value of HIV-1 syncytium-inducing phenotype for rate of CD4+ cell depletion and progression to AIDS. Ann Intern Med. 1993;118:681–688. doi: 10.7326/0003-4819-118-9-199305010-00004. [DOI] [PubMed] [Google Scholar]
- 34.Connor RI, Sheridan KE, Ceradini D, et al. Change in coreceptor use correlates with disease progression in HIV-1--infected individuals. J Exp Med. 1997;185:621–628. doi: 10.1084/jem.185.4.621. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hunt PW, Harrigan PR, Huang W, et al. Prevalence of CXCR4 tropism among antiretroviral-treated HIV-1-infected patients with detectable viremia. J Infect Dis. 2006;194:926–930. doi: 10.1086/507312. [DOI] [PubMed] [Google Scholar]
- 36.Bleul CC, Wu L, Hoxie JA, et al. The HIV coreceptors CXCR4 and CCR5 are differentially expressed and regulated on human T lymphocytes. Proc Natl Acad Sci U S A. 1997;94:1925–1930. doi: 10.1073/pnas.94.5.1925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lee B, Sharron M, Montaner LJ, et al. Quantification of CD4, CCR5, and CXCR4 levels on lymphocyte subsets, dendritic cells, and differentially conditioned monocyte-derived macrophages. Proc Natl Acad Sci U S A. 1999;96:5215–5220. doi: 10.1073/pnas.96.9.5215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Blaak H, van't Wout AB, Brouwer M, et al. In vivo HIV-1 infection of CD45RA(+)CD4(+) T cells is established primarily by syncytium-inducing variants and correlates with the rate of CD4(+) T cell decline. Proc Natl Acad Sci U S A. 2000;97:1269–1274. doi: 10.1073/pnas.97.3.1269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Heeregrave EJ, Geels MJ, Brenchley JM, et al. Lack of in vivo compartmentalization among HIV-1 infected naive and memory CD4+ T cell subsets. Virology. 2009;393:24–32. doi: 10.1016/j.virol.2009.07.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.van Rij RP, Blaak H, Visser JA, et al. Differential coreceptor expression allows for independent evolution of non-syncytium-inducing and syncytium-inducing HIV-1. J Clin Invest. 2000;106:1569. doi: 10.1172/jci7953c1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ince WL, Harrington PR, Schnell GL, et al. Major coexisting human immunodeficiency virus type 1 env gene subpopulations in the peripheral blood are produced by cells with similar turnover rates and show little evidence of genetic compartmentalization. J Virol. 2009;83:4068–4080. doi: 10.1128/JVI.02486-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Hwang SS, Boyle TJ, Lyerly HK, Cullen BR. Identification of the envelope V3 loop as the primary determinant of cell tropism in HIV-1. Science. 1991;253:71–74. doi: 10.1126/science.1905842. [DOI] [PubMed] [Google Scholar]
- 43.de Jong JJ, Goudsmit J, Keulen W, et al. Human immunodeficiency virus type1 clones chimeric for the envelope V3 domain differ in syncytium formation and replication capacity. J Virol. 1992;66:757–765. doi: 10.1128/jvi.66.2.757-765.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Milich L, Margolin BH, Swanstrom R. Patterns of amino acid variability in NSI-like and SI-like V3 sequences and a linked change in the CD4-binding domain of the HIV-1 Env protein. Virology. 1997;239:108–118. doi: 10.1006/viro.1997.8821. [DOI] [PubMed] [Google Scholar]
- 45.Jensen MA, Li FS, van 't Wout AB, et al. Improved coreceptor usage prediction and genotypic monitoring of R5-to-X4 transition by motif analysis of human immunodeficiency virus type 1 env V3 loop sequences. J Virol. 2003;77:13376–13388. doi: 10.1128/JVI.77.24.13376-13388.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Resch W, Hoffman N, Swanstrom R. Improved success of phenotype prediction of the human immunodeficiency virus type 1 from envelope variable loop 3 sequence using neural networks. Virology. 2001;288:51–62. doi: 10.1006/viro.2001.1087. [DOI] [PubMed] [Google Scholar]
- 47.Hoffman NG, Seillier-Moiseiwitsch F, Ahn J, et al. Variability in the human immunodeficiency virus type 1 gp120 Env protein linked to phenotypeassociated changes in the V3 loop. J Virol. 2002;76:3852–3864. doi: 10.1128/JVI.76.8.3852-3864.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Huang W, Toma J, Fransen S, et al. Coreceptor tropism can be influenced by amino acid substitutions in the gp41 transmembrane subunit of human immunodeficiency virus type 1 envelope protein. J Virol. 2008;82:5584–5593. doi: 10.1128/JVI.02676-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Tscherning C, Alaeus A, Fredriksson R, et al. Differences in chemokine coreceptor usage between genetic subtypes of HIV-1. Virology. 1998;241:181–188. doi: 10.1006/viro.1997.8980. [DOI] [PubMed] [Google Scholar]
- 50.Huang W, Eshleman SH, Toma J, et al. Coreceptor tropism in human immunodeficiency virus type 1 subtype D: high prevalence of CXCR4 tropism and heterogeneous composition of viral populations. J Virol. 2007;81:7885–7893. doi: 10.1128/JVI.00218-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Sina ST, Ren W, Cheng-Mayer C. Coreceptor use in nonhuman primate models of HIV infection. J Transl Med. 2011;9(Suppl 1):S7. doi: 10.1186/1479-5876-9-S1-S7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Ho SH, Tasca S, Shek L, et al. Coreceptor switch in R5-tropic simian/human immunodeficiency virus-infected macaques. J Virol. 2007;81:8621–8633. doi: 10.1128/JVI.00759-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Nishimura Y, Shingai M, Willey R, et al. Generation of the pathogenic R5- tropic simian/human immunodeficiency virus SHIVAD8 by serial passaging in rhesus macaques. J Virol. 2010;84:4769–4781. doi: 10.1128/JVI.02279-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Ren W, Tasca S, Zhuang K, et al. Different tempo and anatomic location of dual-tropic and X4 virus emergence in a model of R5 simian-human immunodeficiency virus infection. J Virol. 2010;84:340–351. doi: 10.1128/JVI.01865-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Ho SH, Shek L, Gettie A, et al. V3 loop-determined coreceptor preference dictates the dynamics of CD4+-T-cell loss in simian-human immunodeficiency virus-infected macaques. J Virol. 2005;79:12296–12303. doi: 10.1128/JVI.79.19.12296-12303.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Nishimura Y, Igarashi T, Donau OK, et al. Highly pathogenic SHIVs and SIVs target different CD4+ T cell subsets in rhesus monkeys, explaining their divergent clinical courses. Proc Natl Acad Sci U S A. 2004;101:12324–12329. doi: 10.1073/pnas.0404620101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Dunfee RL, Thomas ER, Gorry PR, et al. The HIV Env variant N283 enhances macrophage tropism and is associated with brain infection and dementia. Proc Natl Acad Sci U S A. 2006;103:15160–15165. doi: 10.1073/pnas.0605513103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Dunfee RL, Thomas ER, Wang J, et al. Loss of the N-linked glycosylation site at position 386 in the HIV envelope V4 region enhances macrophage tropism and is associated with dementia. Virology. 2007;367:222–234. doi: 10.1016/j.virol.2007.05.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Harrington PR, Haas DW, Ritola K, Swanstrom R. Compartmentalized human immunodeficiency virus type 1 present in cerebrospinal fluid is produced by short-lived cells. J Virol. 2005;79:7959–7966. doi: 10.1128/JVI.79.13.7959-7966.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Harrington PR, Schnell G, Letendre SL, et al. Cross-sectional characterization of HIV-1 env compartmentalization in cerebrospinal fluid over the full disease course. AIDS. 2009;23:907–915. doi: 10.1097/QAD.0b013e3283299129. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Pillai SK, Pond SLK, Liu Y, et al. Genetic attributes of cerebrospinal fluidderived HIV-1 env. Brain. 2006;129:1872–1883. doi: 10.1093/brain/awl136. [DOI] [PubMed] [Google Scholar]
- 62.Ritola K, Robertson K, Fiscus SA, et al. Increased human immunodeficiency virus type 1 (HIV-1) env compartmentalization in the presence of HIV-1-associated dementia. J Virol. 2005;79:10830–10834. doi: 10.1128/JVI.79.16.10830-10834.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Xu Y, Zhu H, Wilcox CK, et al. Blood monocytes harbor HIV type 1 strains with diversified phenotypes including macrophage-specific CCR5 virus. J Infect Dis. 2008;197:309–318. doi: 10.1086/524847.. This is a provocative report identifying virus in monocytes that have lost the ability to replicate in T cells.
- 64.Li S, Juarez J, Alali M, et al. Persistent CCR5 utilization and enhanced macrophage tropism by primary blood human immunodeficiency virus type1 isolates from advanced stages of disease and comparison to tissuederived isolates. J Virol. 1999;73:9741–9755. doi: 10.1128/jvi.73.12.9741-9755.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Sterjovski J, Roche M, Churchill MJ, et al. An altered and more efficient mechanism of CCR5 engagement contributes to macrophage tropism of CCR5-using HIV-1 envelopes. Virology. 2010;404:269–278. doi: 10.1016/j.virol.2010.05.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Taylor BM, Foulke JS, Flinko R, et al. An alteration of human immunodeficiency virus gp41 leads to reduced CCR5 dependence and CD4 independence. J Virol. 2008;82:5460–5471. doi: 10.1128/JVI.01049-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67.Peters PJ, Duenas-Decamp MJ, Sullivan WM, et al. Variation in HIV-I R5 macrophage-tropism correlates with sensitivity to reagents that block envelope: CD4 interactions but not with sensitivity to other entry inhibitors. Retrovirology. 2008;5:5. doi: 10.1186/1742-4690-5-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Roche M, Jakobsen MR, Sterjovski J, et al. HIV-1 Escape from the CCR5 Antagonist Maraviroc Associated with an Altered and Less-Efficient Mechanism of gp120-CCR5 Engagement That Attenuates Macrophage Tropism. J Virol. 2011;85:4330–4342. doi: 10.1128/JVI.00106-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Sterjovski J, Churchill MJ, Ellett A, et al. Asn 362 in gp120 contributes to enhanced fusogenicity by CCR5-restricted HIV-1 envelope glycoprotein variants from patients with AIDS. Retrovirology. 2007;4:89. doi: 10.1186/1742-4690-4-89. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Duenas-Decamp MJ, Peters P, Burton D, Clapham PR. Natural resistance of human immunodeficiency virus type 1 to the CD4bs antibody b12 conferred by a glycan and an arginine residue close to the CD4 binding loop. J Virol. 2008;82:5807–5814. doi: 10.1128/JVI.02585-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 71.Dunfee RL, Thomas ER, Gabuzda D. Enhanced macrophage tropism of HIV in brain and lymphoid tissues is associated with sensitivity to the broadly neutralizing CD4 binding site antibody b12. Retrovirology. 2009;6:69. doi: 10.1186/1742-4690-6-69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72.Duenas-Decamp MJ, Peters PJ, Burton D, Clapham PR. Determinants flanking the CD4 binding loop modulate macrophage tropism of human immunodeficiency virus type 1 R5 envelopes. J Virol. 2009;83:2575–2583. doi: 10.1128/JVI.02133-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Zhuang K, Finzi A, Tasca S, et al. Adoption of an "Open" Envelope Conformation Facilitating CD4 Binding and Structural Remodeling Precedes Coreceptor Switch in R5 SHIV-Infected Macaques. Plos One. 2011;6:e21350. doi: 10.1371/journal.pone.0021350.. This paper reports a new cell line where CD4 and CCR5 are under control of a regulatable promoters, allowing quantitative assessment of the need for these receptors in entry.
- 74.Johnston SH, Lobritz MA, Nguyen S, et al. A quantitative affinity-profiling system that reveals distinct CD4/CCR5 usage patterns among human immunodeficiency virus type 1 and simian immunodeficiency virus strains. J Virol. 2009;83:11016–11026. doi: 10.1128/JVI.01242-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Duenas-Decamp MJ, Clapham PR. HIV-1 gp120 Determinants Proximal to the CD4 Binding Site Shift Protective Glycans That Are Targeted by Monoclonal Antibody 2G12. J Virol. 2010;84:9608–9612. doi: 10.1128/JVI.00185-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.Richards KH, Aasa-Chapman MMI, McKnight A, Clapham PR. Modulation of HIV-1 macrophage-tropism among R5 envelopes occurs before detection of neutralizing antibodies. Retrovirology. 2010;7:48. doi: 10.1186/1742-4690-7-48. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 77.Musich T, Peters PJ, Duenas-Decamp MJ, et al. A Conserved Determinant in the V1 Loop of HIV-1 Modulates the V3 Loop To Prime Low CD4 Use and Macrophage Infection. J Virol. 2011;85:2397–2405. doi: 10.1128/JVI.02187-10. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 78.Walter BL, Wehrly K, Swanstrom R, et al. Role of low CD4 levels in the influence of human immunodeficiency virus type 1 envelope V2 and V2 regions on entry and spread in macrophages. J Virol. 2005;79:4828–4837. doi: 10.1128/JVI.79.8.4828-4837.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Cashin K, Roche M, Sterjovski J, et al. Alternative coreceptor requirements for efficient CCR5- and CXCR4-mediated HIV-1 entry into macrophages. J Virol. doi: 10.1128/JVI.05510-11. In press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 80.Ghaffari G, Tuttle DL, Briggs D, et al. Complex determinants in human immunodeficiency virus type 1 envelope gp120 mediate CXCR4-dependent infection of macrophages. J Virol. 2005;79:13250–13261. doi: 10.1128/JVI.79.21.13250-13261.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 81.Auffray C, Sieweke MH, Geissmann F. Blood Monocytes: Development, Heterogeneity, and Relationship with Dendritic Cells. Annu Rev Immunol. 2009;27:669–692. doi: 10.1146/annurev.immunol.021908.132557. [DOI] [PubMed] [Google Scholar]
- 82.Grage-Griebenow E, Flad HD, Ernst M. Heterogeneity of human peripheral blood monocyte subsets. J Leuk Biol. 2001;69:11–20. [PubMed] [Google Scholar]
- 83.Ellery PJ, Tippett E, Chiu YL, et al. The CD16(+) monocyte subset is more permissive to infection and preferentially harbors HIV-1 in vivo. J Immunol. 2007;178:6581–6589. doi: 10.4049/jimmunol.178.10.6581. [DOI] [PubMed] [Google Scholar]
- 84.Wong KL, Tai JJ, Wong WC, et al. Gene expression profiling reveals the defining features of the classical, intermediate, and nonclassical human monocyte subsets. Blood. 2011;118:e16–e31. doi: 10.1182/blood-2010-12-326355. [DOI] [PubMed] [Google Scholar]
- 85.Crowe S, Zhu TF, Muller WA. The contribution of monocyte infection and trafficking to viral persistence, and maintenance of the viral reservoir in HIV infection. J Leuk Biol. 2003;74:635–641. doi: 10.1189/jlb.0503204. [DOI] [PubMed] [Google Scholar]
- 86.Fogg DK, Sibon C, Miled C, et al. A clonogenic bone marrow progenitor specific for macrophages and dendritic cells. Science. 2006;311:83–87. doi: 10.1126/science.1117729. [DOI] [PubMed] [Google Scholar]
- 87.Ziegler-Heitbrock L, Ancuta P, Crowe S, et al. Nomenclature of monocytes and dendritic cells in blood. Blood. 2010;116:E74–E80. doi: 10.1182/blood-2010-02-258558. [DOI] [PubMed] [Google Scholar]
- 88.Naif HM, Li S, Alali M, et al. CCR5 expression correlates with susceptibility of maturing monocytes to human immunodeficiency virus type 1 infection. J Virol. 1998;72:830–836. doi: 10.1128/jvi.72.1.830-836.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 89.Sonza S, Maerz A, Uren S, et al. Susceptibility of Human Monocytes to HIV Type 1 Infection in Vitro Is Not Dependent on Their Level of CD4 Expression. AIDS Res Hum Retroviruses. 1995;11:769–776. doi: 10.1089/aid.1995.11.769. [DOI] [PubMed] [Google Scholar]
- 90.Dong C, Kwas C, Wu L. Transcriptional restriction of human immunodeficiency virus type 1 gene expression in undifferentiated primary monocytes. J Virol. 2009;83:3518–3527. doi: 10.1128/JVI.02665-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 91.Sung T-L, Rice AP. miR-198 Inhibits HIV-1 Gene Expression and Replication in Monocytes and Its Mechanism of Action Appears To Involve Repression of Cyclin T1. Plos Pathog. 2009;5:e1000263. doi: 10.1371/journal.ppat.1000263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 92.Lewin SR, Lambert P, Deacon NJ, et al. Constitutive expression of p50 homodimer in freshly isolated human monocytes decreases in vitro and in vivo differentiation: A possible mechanism influencing human immunodeficiency virus replication in monocytes and mature macrophages. J Virol. 1997;71:2114–2119. doi: 10.1128/jvi.71.3.2114-2119.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 93.Arfi V, Riviere L, Jarrosson-Wuilleme L, et al. Characterization of the early steps of infection of primary blood monocytes by human immunodeficiency virus type 1. J Virol. 2008;82:6557–6565. doi: 10.1128/JVI.02321-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 94.Meuret G, Bammert J, Hoffmann G. Kinetics of Human Monocytopoiesis. Blood. 1974;44:801–816. [PubMed] [Google Scholar]
- 95.Zhu TF, Muthui D, Holte S, et al. Evidence for human immunodeficiency virus type 1 replication in vivo in CD14(+) monocytes and its potential role as a source of virus in patients on highly active antiretroviral therapy. J Virol. 2002;76:707–716. doi: 10.1128/JVI.76.2.707-716.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 96. Sonza S, Mutimer HP, Oelrichs R, et al. Monocytes harbour replicationcompetent, non-latent HIV-1 in patients on highly active antiretroviral therapy. AIDS. 2001;15:17–22. doi: 10.1097/00002030-200101050-00005..
- 97.Riddick NE, Hermann EA, Loftin LM, et al. A novel CCR5 mutation common in sooty mangabeys reveals SIVsmm infection of CCR5-null natural hosts and efficient alternative coreceptor use in vivo. PLoS Pathog. 2010;6:e1001064. doi: 10.1371/journal.ppat.1001064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 98.Dumonceaux J, Nisole S, Chanel C, et al. Spontaneous mutations in the env gene of the human immunodeficiency virus type 1 NDK isolate are associated with a CD4-independent entry phenotype. J Virol. 1998;72:512–519. doi: 10.1128/jvi.72.1.512-519.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 99.Haim H, Strack B, Kassa A, et al. Contribution of Intrinsic Reactivity of the HIV-1 Envelope Glycoproteins to CD4-Independent Infection and Global Inhibitor Sensitivity. Plos Pathog. 2011;7:e1002101. doi: 10.1371/journal.ppat.1002101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 100.Hoffman TL, LaBranche CC, Zhang WT, et al. Stable exposure of the coreceptor-binding site in a CD4-independent HIV-1 envelope protein. Proc Natl Acad Sci U S A. 1999;96:6359–6364. doi: 10.1073/pnas.96.11.6359. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 101.Kolchinsky P, Mirzabekov T, Farzan M, et al. Adaptation of a CCR5-using, primary human immunodeficiency virus type 1 isolate for CD4-independent replication. J Virol. 1999;73:8120–8126. doi: 10.1128/jvi.73.10.8120-8126.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 102.Marras D, Bruggeman LA, Gao F, et al. Replication and compartmentalization of HIV-1 in kidney epithelium of patients with HIV-associated nephropathy. Nature Medicine. 2002;8:522–526. doi: 10.1038/nm0502-522. [DOI] [PubMed] [Google Scholar]
- 103.Edwards TG, Hoffman TL, Baribaud F, et al. Relationships between CD4 independence, neutralization sensitivity, and exposure of a CD4-induced epitope in a human immunodeficiency virus type 1 envelope protein. J Virol. 2001;75:5230–5239. doi: 10.1128/JVI.75.11.5230-5239.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 104.Eitner F, Cui Y, Hudkins KL, et al. Chemokine receptor CCR5 and CXCR4 expression in HIV-associated kidney disease. Journal of the American Society of Nephrology. 2000;11:856–867. doi: 10.1681/ASN.V115856. [DOI] [PubMed] [Google Scholar]
- 105.Bruggeman LA, Ross MD, Tanji N, et al. Renal epithelium is a previously unrecognized site of HIV-1 infection. Journal of the American Society of Nephrology. 2000;11:2079–2087. doi: 10.1681/ASN.V11112079. [DOI] [PubMed] [Google Scholar]