Abstract
The significant mortality rate associated with metastatic melanoma, which exceeds the number of deaths attributed to the primary tumor, is primarily due to poor diagnosis and increased resistance to systemic therapy. Early detection and treatment of invasive melanoma are therefore crucial to increase survival rates. Low molecular weight proteins and peptides have garnered significant interest as biomarker candidates as they potentially represent a snap shot of pathological condition within the body and, by extension, the organism as a whole. We have developed a nanoporous silica-based platform to segregate the low molecular weight from the high molecular weight protein fraction to aid in the detection of peptides from serum samples using mass spectrometry. The combination of sample treatment with our platform, MALDI-TOF MS and following biostatistical analysis led to the discovery and identification of 27 peptides that are potential biomarkers associated with the development of pulmonary metastatic melanoma. We strongly believe our findings can assist to discover stage-specific peptide signatures and lead to more specific and personalized treatments for patients suffering from pulmonary metastatic melanoma.
Keywords: Peptide signatures, Nanoporous silica chip, pulmonary metastatic melanoma, MALDI-TOF MS, Principal component analysis
1. Introduction
In the United States alone, melanoma is the fifth most common type of cancer in men, the sixth most common in women and is responsible for no less than 5% of the total cancer cases [1]. The American Cancer Society estimates that the number of new cases and deaths in the U.S. from melanoma will be 70,230 and 8,790, respectively, in 2012 [1]. Recently, the incidence of metastatic melanoma has increased while the death rate has risen faster than most other cancers [2]. Patients with early-stage localized melanoma can be diagnosed reasonably accurately with the current staging classification system and have high survival rates. Patients suffering from later stage metastatic melanoma, however, are considerably more resistant to systemic therapy, leading to survival rates below 15% [3]. The identification of reliable markers to predict early metastasis in melanoma may enable immediate treatment for patients and improve their survival in the cases where early diagnosis is missed. Currently the American Joint Committee on Cancer (AJCC) only has one blood protein, lactate dehydrogenase (LDH), in their melanoma staging system. LDH cannot be used for early prediction, however, as only the late stages of metastatic melanoma expresses increased levels of this protein in serum[4]. Decreased expression levels of other proteins, including WNT5A and Foxp3, have also been demonstrated to correlate with tumor metastasis and decreased survival rates [5; 6]. As cancer is considered to be an invader of the human body, it triggers the immune system, eliciting biomarkers. Moreover, melanoma is considered to be a highly immunogenic tumor [7]. As the immune system is a complicated cascade of reactions, a single biomarker is insufficient to provide an accurate diagnosis. So far no biomarkers are available to sufficiently evaluate the risk of metastasis.
A limited number of oncogenes have already been identified for melanoma, most of them related to the RAS/MAPK signaling pathway, which is responsible for the cellular response to growth factor receptor signals[8]. These genetic studies require high numbers of fresh primary tumor tissues collected by surgery. Therefore it is desirable to develop other less invasive methods to augment our ability to characterize these tumors. Secreted proteins, which become part of the so-called secretome, reflect the state of a cell in a certain environment, and are therefore important indicators of a cell’s condition and by extension of the entire organism. The blood proteome can be used for the discovery of biomarkers that are released from both tumor cells and the immune system of the organism and can be further correlated to treatment efficacy [9; 10; 11; 12] .
Serum proteins and peptides could provide valuable information about the processes that occur in the body [13; 14; 15]. Some typical low molecular weight (LMW) protein compounds in serum are hormones, cytokines, chemokines and fragments of larger proteins [15; 16; 17]. The circulating LMW proteome has been associated with pathological conditions in patients. During recent years, several studies have reported that LMW proteins and peptides have been associated with cancer, such as breast, prostate, bladder and thyroid cancer [18; 19]. However, the isolation of LMW proteins/peptides is still a challenge due to the large dynamic range of serum proteins, and the presence of high abundant proteins, like albumin, immunoglobulin, and transferrin [14]. In previous studies, we have demonstrated the ability to optimize the nanotexture of silica films through the engineering of the physicochemical properties (pore size, pore structure, and surface affinity, etc.) on chips that can selectively parse and enrich peptides and low molecular weight proteins from serum samples [20; 21; 22; 23].
In this paper, we describe the use of nanoporous silica chips (NSC) for the high-throughput identification of biomarkers that are characteristic of pulmonary metastatic melanoma using the B16 mouse model. The lung is the most common and first site of visceral metastasis in melanoma, with over 85% of late stage melanoma patients having pulmonary metastasis [24]. The pulmonary metastatic melanoma mouse model is therefore a very important tool for human melanoma research. Serum samples were collected at four different time points before and after tail vein inoculation of B16 cells (7, 14 and 21 days). After on-chip sample fractionation, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF MS) was used for profiling of eluted serum peptides. All data were then subjected to principal component analysis (PCA) and peptides associated with tumor progression were selected to identify signatures unique to the different stages of cancer development. Finally, the peptides-of-interest were subjected to LC-MS/MS for identification. The identified peptides generally belonged to larger secreted proteins, though most of them were related to cancer. Expression profiles of the peptides showed different trends not only for different proteins, but also for several peptides belonging to the same protein.
2. Methods and Materials
2.1 Cell culture
B16 cells, a murine melanoma cell line, were obtained from the American Type Culture Collection (ATCC) and cultivated in RPMI-1640 cell culture medium supplemented with 10% heat-inactivated fetal bovine serum (FBS), 1% glutamine, and 0.1% penicillin/streptomycin at 37°C and 5% CO2.
2.2 B16 Melanoma Lung Metastasis Mouse Model
All animal work was done in accordance with a protocol approved by the Institutional Animal Care and Use Committee (IACUC) of The Methodist Hospital Research Institute in Houston, TX. Six to eight-week-old female nude mice were purchased from the Charles River Laboratories. They were housed in a specific pathogen-free facility under a 12-hour light-dark cycle, and fed with a pathogen-free diet and water ad libitum. To constitute a mouse model of melanoma lung metastatic, 1×105 of B16 melanoma cells were suspended in 100 μl PBS, and injected into the lateral tail vein. The tumor foci were well established in the lungs in 5-7 days.
2.3 Blood Sample Collection
Thirty female mice with pulmonary metastatic melanoma were used. Blood samples were collected by retro orbital bleeding at 4 different time points (before injection, 7, 14 and 21 days after injection). For each time point, 100 l blood was collected from each mouse. The blood samples were kept at room temperature for 1 hr and subsequently centrifuged at 3600 rpm for 15 min. The sera were carefully collected and stored at −80°C until further analysis.
2.4 Hematoxylin and Eosin (H&E) staining
Formalin fixed paraffin embedded sections of the lungs were cut at 4 microns, baked overnight in an oven at 60 °C, dewaxed in xylene and rehydrated by increasing the concentration of distilled water in reagent alcohol. The tissue was stained for 3 minutes in 3% Acetic acid/Harris Hematoxylin, washed with running tap water for 1 min and rinsed in 95% reagent alcohol prior to dipping 10 times in Alcoholic Eosin Y stain. Next, tissues were dehydrated in increasing concentration of reagent alcohol, cleared in xylene and permanently coversliped. Images of tissue sections after H&E staining were captured by Eclipse 80i microscope (Nikon Comp.).
2.5 Nanoporous silica thin films fabrication
The fabrication of NPS thin films start with the preparation of silicate sol by adding 14 mL of tetraethyl orthosilicate into the mixture of 17 mL of ethanol, 6.5 mL of distilled water, and 0.5 mL of 6M HCl and stirred 2 h at 80 °C to form a clear silicate sol. Separately, 1.2 g of Pluronic L121 three triblock copolymer were dissolved into 10 mL of ethanol by stirring at room temperature. Then the coating solution was prepared by mixing 10 mL silicate sol into the polymer solution followed by stirring for 2 h at room temperature. The coating solution was deposited on a Si(100) wafer by spin coating at the spin rate of 2500 rpm for 20 s. To increase the degree of polymerization of silica framework in the films and to further improve their thermal stability, the films were placed in an oven at 80 °C for 12 hours. The films were then calcinated at 450 °C for 5 hours to remove the organic surfactant. The furnace was heated at a rate of 1 °C per min and was maintained at the final temperature for 5 hours.
2.6 On chip fractionation
Serum samples were thawed on ice. For the pH 7 samples, a solution of 50 % acetonitrile (ACN; Sigma-Aldrich, St. Louis, MO, USA) and 0.1 % trifluoroacetic acid (TFA; Sigma-Aldrich, St. Louis, MO, USA) was prepared in deionized water and 2.2 l was added to 20 l sample. For the pH 5 samples, 1.7 l of a 5% TFA solution was added to 20 l serum samples. Samples were then mixed on a table vortex at room temperature for 30 min, followed by centrifugation at 5000 rpm for 5 min.
In this study, the nanoporous silica chips with Pluronic L121 as synthetic template were fabricated as described previously [20]. Chambered coverslips (3mm diameter×1mm height, Sigma-Aldrich, St. Louis, MO, USA) were washed in 100 % ethanol and placed on the chip surface. 5 l serum was pipetted into each well, ensuring that no precipitate in the base of a tube was placed on the chip. The samples were allowed to incubate at room temperature in a humidified chamber for 30 min. After incubation, the serum samples were removed and the wells were washed four times with 10 l deionized water. For elution a 50% ACN and 0.1% TFA solution was prepared in deionized water, and 5 l was applied into each well and left in it for 90 seconds to extract the peptides from nanopores. Then the elution solution was pipetted and transferred into a fresh microcentrifuge tube, which were stored at −80°C until further analysis.
2.7 Matrix-assisted laser desorption/ionization time-of-flight analysis
A fresh disposable MALDI target plate was used to avoid carry over from previous experiments. 0.5 l of each sample was spotted on the target plate and allowed to dry completely, while preparing the matrix solution (5 g/l of a-cyano-4-hydroxycinnamic acid (CHCA) in 50 % ACN and 0.1 % TFA). Following, 0.5 μl of matrix solution was spotted on the target plate and also allowed to dry. All samples were analyzed on an Applied Biosystems 4700 MALDI TOF Analyzer (Applied Biosystems, Inc., Framingham, MA, USA). The system was operated in positive ion mode with reflector (laser intensity 4300, 5000 shots/sample and mass range 800–5000 Da with target mass of 2000 Da).
2.8 LC MS/MS to identify
Reversed phase chromatography was performed on an Agilent 1200 series HPLC autosampler. Gradient solvents used for LC analysis are (A) 0.1% formic acid in water and (B) 0.1 % formic acid in acetonitrile. Samples were dried in a vacuum centrifuge prior to injection into the LC and resuspended in 1% formic acid, 5 mM NH4OAc. Analysis of the peptides was done on an Orbtrap-XL mass spectrometer (Thermo Scientific, Waltham MA). Acquisition parameters were: 1 FTMS scan at 60000 resolution followed by 3 MS/MS product ion scans (in the ion trap) of 2 microscans each. Total run time was 70 minutes. Proteins were identified by database searching of the fragment spectra against the SwissProt (EBI) protein database using Mascot (v 2.3, Matrix Science, London, UK) or Sequest (Thermo, San Jose, CA). Typical search settings were: mass tolerances, 15 ppm precursor, 0.8d fragments; variable modifications, methionine sulfoxide, pyro-glutamate formation; no enzyme and up to 9 missed cleavages.
2.9 Statistical analysis
The MALDI-TOF data were processed using CONVERTPEAKLIST software to generate text files, listing m/z-intensity pairs. These text files were then imported into SPECALIGN software for visualization and averaging of the spectra. The PCA-DA and t-test analysis were performed by MarkerView software v. 1.2.1 (AB SCIEX, Concord, Canada). Six repeated spectra for each sample were selected and imported into MarkerView. The data were normalized to total the area sum PCA-DA processing was carried out with a Pareto scaling. The t-value and corresponding p-value were also calculated between each group and all the other three groups using MarkerView.
3. Results
3.1 Melanoma Lung Metastasis Mouse Model
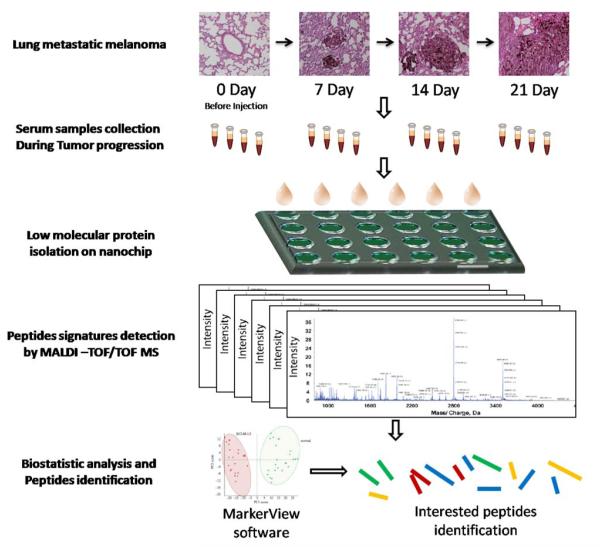
In our experiment, the B16 tumor cells were injected intravenously via the tail vein. The serum samples were collected before tumor cells injection, and 7, 14 and 21 days after injection. Two mice were sacrificed at each time point to monitor the development of melanoma metastasis in lung. The lung tissues were collected from each mouse for H&E staining. The surface lung metastatic foci were very tiny on day 7, but could be clearly observed by H&E staining (Figure 1). Tumor foci grew bigger on day 14 and by day 21, both size and the number of foci had increased comparing to the previous week. Tumor bearing mice were sacrificed after the last blood samples collection. Histological analysis revealed that all mice had developed lung metastasis after cell inoculation in our experiment. Thus, the B16 melanoma lung metastasis mouse model is a suitable animal model and can be used for further biomarker detection studies.
Figure 1.
Overview of this study design. The mouse serum samples were collected at different time points during the lung metastatic melanoma tumor progression. Serum peptides were enriched using nanoporous silica chips, peptide profiles were obtained by MALDI-TOF MS and the interesting peptides were selectedfor identification by LC-MS/MS after PCA analysis.
3.2 Serum peptides on chip isolation
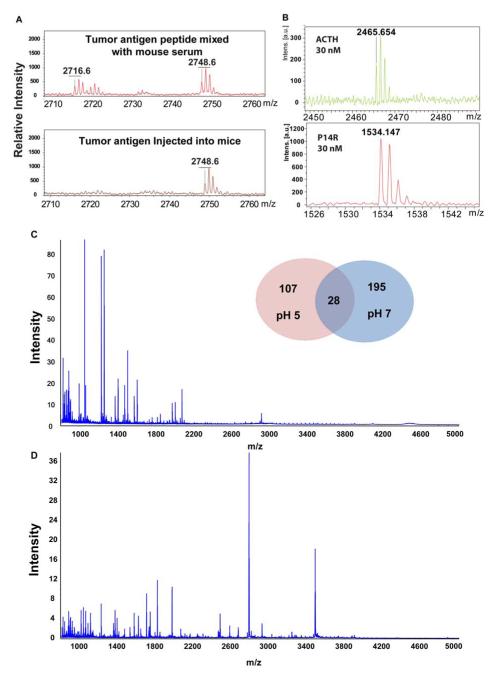
The procedure of our study is shown in Figure 1. After successful inoculation of mice with B16 melanoma cells, serum samples were collected at four different time points. The fractionation method based on our nanoporous silica chip was described previously [20; 23]. We also evaluated the capacity and sensitivity of our approach to capture tumor-specific biomarker with a vital control experiment. Since tyrosinase-related protein 2 (TRP-2) expressed by the murine B16 melanoma is as a tumor antigen [25], and a TRP-2 peptide has been identified as the major reactive epitope recognized by the anti-B16 cytotoxic T lymphocytes [26], we selected TRP-2 as a positive control peptide and spiked it into blank mouse serum directly to investigate its MALDI-TOF MS profile. Then we injected it into mouse circulating system following by serum collection. As shown in Figure 2A, TRP-2 peptide can be detected by MALDI-TOF MS after enrichment on our nanoporous silica chip. To evaluate the quality of our method, two commercial standard peptides (P14R at 1534 m/z and ACTH fragment 18-39 at 2465 m/z) were spiked into mouse serum at different concentration from 30 nM to 1μM. Figure 2B indicates that both standard peptides can be detected with desirable signal to noise ratio at very low concentration.
Figure 2.
MALDI-TOF mass spectrum of TRP-2 tumor antigen, standard peptides and averaged MS spectra for serum peptide fractionation at pH 7 and pH5. (A) TRP-2 tumor antigen peptide was respectively spiked (above) into mouse serum and injected (bottom) into mouse circulating system; (B) Two standard peptides, ACTH fragment 18-39 and P14R recovered by on-nanochip fractionation after spiked into mouse serum at low concentration; Averaged MS spectra for serum peptide fractionation at pH 7 (C) and 5 (D). Insert shows overview of detected peaks between both conditions.
In this study, to extend our detection range, we adjusted serum samples to both neutral and acidic conditions to affect the interaction between serum peptides and the surface of nanopores. To know how the change in pH of the samples influenced the fractionation, the aver age mass spectrums were generated per group and compared. Figure 2C and D show these average mass spectrums under pH5 and pH7 respectively. It can be concluded from Figure 2C and D that different peptides are captured under different pH conditions. In total, 330 peaks were detected, 223 peaks of which in neutral conditions and 135 in acidic conditions. Of these 330 peaks, 28 were detected in both conditions (Figure 2 insert). This shows that one condition is not necessarily superior to the other, but rather that both pH states are complementary and can potentially reveal different candidate biomarker peptides.
3.3 Principal component analysis
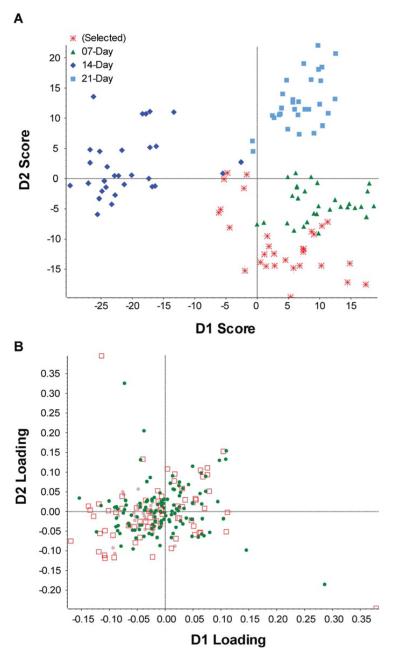
After mass spectrum acquisition for the peptide fractions, PCA-DA (principal component analysis and discriminant analysis) was performed using MarkerView. PCA is a statistical procedure commonly used for identifying clustering patterns in data by reducing dimensional variables, while still maintaining most of the original series [28]. To improve the separation of the four known groups during lung metastatic melanoma tumor progression, we performed PCA discriminant analysis and used D1 and D2 score rather than PC1 and PC2 (first and second principal component). The results of the PCA-DA were displayed as the variable projection scores and loadings plots. Score plots represent the variance of the original data, and show the score of each sample for the first principal component against the second. Figure 3 shows the score plot (A) and the loading plot (B) for these data sets. The first two principal components explained 71.40 % of total variance, suggesting that this score plot is representative for the original data. From the score plot in Figure 3A it can be concluded that after PCA-DA analysis, D1 and D2 scores are able to distinguish between the four different sample groups, representing different stages in tumor progressions. On the score plot, 3 clusters can be distinguished which are the 14-, the 21-, and the control together with the samples of day 7. When looking at the D2 scores, the samples of day 21 are the only ones with a positive score, while the group of day 14 is spread around zero and the control and samples of day 7 are negative. In general, of the 3 clusters, 2 of them each occupy 1 quadrant and the third cluster is spread over 2 quadrants.
Figure 3.
PCA analysis of four time-points groups under pH7.A) PCA-DA plot of four groups at different time-points during tumor growth (0 day, 7 day, 14 day and 21 day. B) Loading plot for the four time-point groups.
The loading plots described the variable behavior and difference among observed groups. In Figure 3B, the loading plots demonstrate the D1 and D2 loadings for each variable, indicating how each variable contributes to the first two principal components, and revealing the peptides specifically related to tumor metastasis progression.
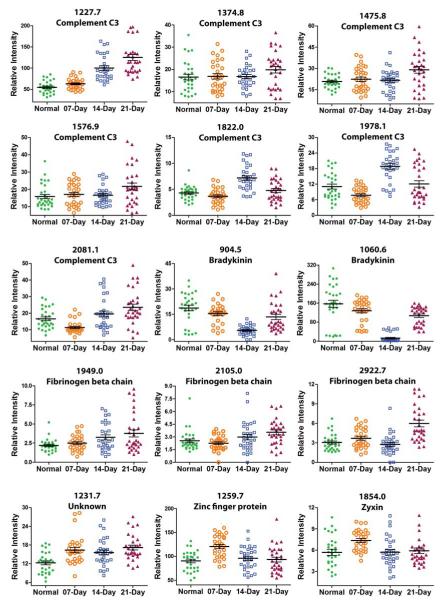
To examine the peptides associated with tumor metastasis revealed by the loading plot, a t-test analysis was performed using MarkerView. The t-test was carried out by comparing each group to the other three groups. The peaks resulting in lower p-values showed good agreement with PCA-DA results which also displayed higher absolute value of loading scores. Figures 4 and 5 show the expression profiles of the peptides of interest at neutral and acidic pH respectively. The vertical scattering plots show the smallest observation, lower quartile, median, upper quartile, and largest observation. Even though not all selected masses show significant changes in expression level related to metastasis development, most peptides have time-dependent fluctuating profiles.
Figure 4.
Box chart showing expression profiles of peptides associated with melanoma lung metastasis progression under pH7 conditions.
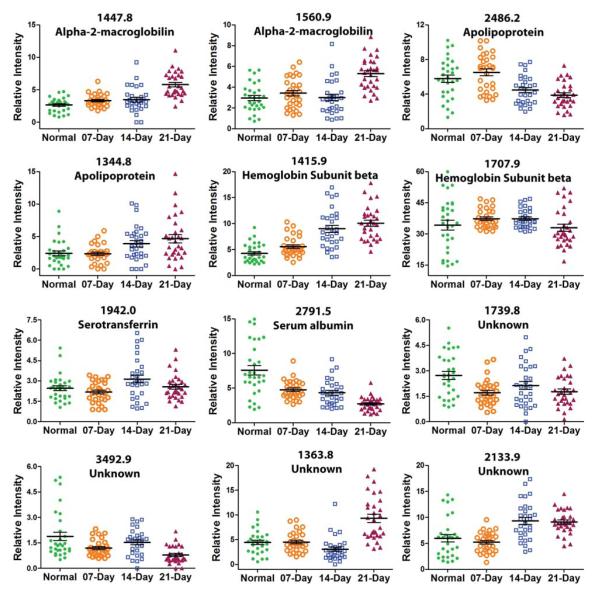
Figure 5.
Box chart showing expression profiles of peptides related with lung metastatic melanoma tumor progression under pH5 conditions.
3.4 Identification of interested peptides
To further study the peptide signatures under neutral and acidic conditions, we identified the peptides-of-interest using LC-MS/MS. Of the 27 selected peptides, 21 were positively identified by LC-MS/MS (Table 1 and 2). Note that the m/z values listed in Table 1 and 2 are monoisotopic. Seven sequences, which were identified in neutral condition, originate from complement C3, a component of the complement system (Figure 4). All these 7 fragments displayed higher level during pulmonary metastatic growth and changed most through day 14 to 21. Bradykinin and desArg-bradykinin also identified at neutral conditions exhibited a decreasing expression level up to day 14, followed by an increase up to 21 days (Figure 4). Three fragments derived from fibrinogen beta chain enriched under pH 7 were increased in tumor groups, especially in 14- and 21-day groups (Figure 4). Under acidic conditions, two fragments from alpha-2 macroglobulin, one fragment from apolipoprotein A-I (1344.8 m/z) and one fragment from serotransferrin show increasing expression levels throughout the tumor development (Figure 5). Three identified peptides show a decreasing expression profile in the acidic condition. These peptides originate from apolipoproteins A-1 (m/z 2468.2), hemoglobin subunit beta and serum albumin (Figure 5). Four unknown fragments were also found under acidic conditions, peptides with 1739.8 and 3492.9 m/z show lower expression level at 21-days, peptide with 1363.8 m/z shows a significant increase at 21 days and peptide with 2133.9 m/z shows an increase going from 7 days to 14 days, followed by a status quo.
Table 1.
Identification of interested peptides incubated on chip at pH7
| Protein Description | Peptide mH+ | Peptide Sequence | Peptide pI |
|---|---|---|---|
| Complement C3 | 1227.6844 | RLLWENGNLL | 6.00 |
| 1374.7528 | FRLLWENGNLL | 6.00 | |
| 1475.8005 | TFRLLWENGNLL | 5.66 | |
| 1576.8409 | TTFRLLWENGNLL | 5.66 | |
| 1821.9494 | SSATTFRLLWENGNLL | 5.72 | |
| 1978.0505 | SSATTFRLLWENGNLLR | 9.31 | |
| 2081.0622 | EETKQNEAFSLTAKGKGR | 8.59 | |
| Zinc finger protein 827 | 1259.6702 | SSQKNKGGNNLL | 10.0 |
| Fibrinogen beta chain | 1949.0127 | KEEPPSLRPAPPPISGGGY | 6.14 |
| 2105.1065 | RKEEPPSLRPAPPPISGGGY | 8.59 | |
| 2922.5372 | GHRPVDRRKEEPPSLRPAPPPISGGGY | 9.97 | |
| Zyxin | 1854.0451 | TPAPKFAPVAPKFTPVVS | 10.0 |
| Bradykinin | 904.4603 | RPPGFSPF | 9.75 |
| 1060.5614 | RPPGFSPFR | 12.00 |
Table 2.
Identification of interested peptides incubation on chip at pH5 condition.
| Protein Description | Peptide mH+ | Peptide Sequence | Peptide pI |
|---|---|---|---|
| Alpha-2-macroglobulin | 1447.84 | NYRPGLPFSGQVL | 8.75 |
| Alpha-2-macroglobulin | 1560.91 | NYRPGLPFSGQVLL | 8.75 |
| Apolipoprotein A I | 2486.22 | WHVWQQDEPQSQWDKVKDF | 4.66 |
| Apolipoprotein A-I | 1344.78 | YRQKVAPLGAEL | 8.59 |
| Hemoglobin subunit beta | 1707.88 | VVYPWTQRYFDSF | 5.81 |
| Serotransferrin | 1942.04 | FSSPLGKDLLFKDSAFGL | 5.96 |
| Serum albumin | 2791.50 | EAHKSEIAHRYNDLGEQHFKGLVL | 6.29 |
4. Discussion
The aim of this study was to detect and identify possible peptides associated with pulmonary metastatic melanoma progression from mouse serum. Furthermore, B16 is an ideal model to test whether our platform is sensitive enough to distinguish the tumor progression stages within a short time interval. In previous works, researchers frequently compared the healthy group and disease group [27; 28; 29]. Few studies focus on the discovery of markers to predict tumor development, which is valuable for personalized diagnosis and treatment evaluation. The level of serum proteins reflects the physiological status of tumor progression [30]. Traditional proteomic methods usually focus on the high molecular weight (HMW) circulating proteins, while the low molecular weight proteins or peptides in serum are also of great interest in disease development and evolution [17]. Our study describes the development of a simple method for peptide enrichment from serum using nanopore-based silica chips combined with MALDI-TOF MS. In the vital control experiment, our results indicated that our method is able to capture the potential peptides biomarkers from circulating biological fluids, e.g. serum-based tumor antigen. Our device combines a size-exclusion effect with a surface charge effect due to the silica. The nanopores on our device act as molecular sieve to exclude the HMW proteins in the sample, while the LMW proteins and peptides bind to the surface of the pores. As the pH of the solution is known to influence the surface charge of proteins and peptides, the samples were incubated at both neutral and slightly acidic conditions. Furthermore, our study confirmed that our nanopore-based silica chip coupled with MALDI TOF MS is a suitable technique in a high-throughput serum peptidome study. Combined with biostatistic analysis by MarkerView, the tumor progression-associated peptide signatures were successfully identified. Most of the peptides identified belonged to secreted proteins, many of which have been described in literature to be cancer related. Their integrated variation in proteomic pattern provides a more powerful platform in diagnosis and prognosis by means of reliability and accuracy, compared to the profiling of single biomarkers.
Most of the peptides identified by our method are actually breakdown products of large and abundant serum proteins, including complement C3, fibrinogen beta chain, bradykinin, apolipoprotein A-I and alpha-2-macroglobulin. This however does not mean they cannot act as potential biomarkers for pulmonary metastatic melanoma. Villanueva et al. published a study recently showing that breakdown products actually should be taken into consideration to be signatures for diseases. They performed a large screening of serum collected from healthy versus patients diagnosed with different types of cancer. They reduced the number of key peptides by filtering the data based on several features to get a group that can be used to discriminate between cancer and control samples [19]. Keeping this in mind, we are convinced that also the peptides we identified might contribute to the development of a signature for pulmonary metastatic melanoma. The peptide fragments might have been generated by common enzymes such as thrombin, plasmin and coagulation factors, and subsequently processed by exoproteases, e.g. aminopeptidase and carboxypeptidase [31].
The protein that had most peptides identified in our study was complement C3. The complement system is essential in inflammation and in the innate immunity to infection and has been linked to many types of cancer and other diseases [32]. The complement C3 is considered central to the complement cascade, which may aid tumor growth through immunosuppression, further promoting angiogenesis, invasion and migration [33]. In our study, the nanopore-based protein chip enriched seven C3 fragments truncations, and all the seven C3 fragments show higher level in tumor groups. Other research also demonstrated that DM-4, a highly metastatic human melanoma cell line in nude mice, actually is known to co-express and co-secrete human C3 and a C3-cleaving cysteine proteinase [34]. This suggests that C3 fragments may act as a factor, which improves growth and metastasis of melanoma cells in nude mice. We do however think further testing is necessary to claim it as a signature peptide for pulmonary metastatic melanoma. Another independent predictor of distant metastasis, fibrinogen, was also identified. Fibrinogen is a large protein (340 KDa) consisting of two sets of three polypeptide chains termed Aα, Bβ and γ [35; 36]. This protein is highly involved in coagulation and has been associated both with melanoma and with lung cancer in the past. Plasmin can digest fibrinogen into two D-fragments and numbers of small peptides including the N-terminus of the beta chain [37]. We observed three peptides degraded from fibrinogen beta chain. All these three peptides increased during tumor progression. High plasma fibrinogen could be found in renal cell carcinoma [38]. Furthermore, fibrinogen was demonstrated as a critical component to promote tumor cell metastasis [39]. Whereas the expression profiles of the different complement C3 peptide profiles do not all show a similar trend, this is much more the case for the fibrinogen peptides. The expression of the fibrinogen peptides shows an increasing trend, while the complement C3 expression profiles are rather fluctuating. This can be explained by the differential expression of exoproteases with higher affinity for the complement C3 proteins. It should be noted that the peptides identified for fibrinogen beta chain under neutral conditions have high proline content, an amino acid often impeding proteases, explaining why those peptides have higher masses compared to the complement C3 peptides. A key issue for tumor growth is the supply of blood and oxygen, supporting our findings of these haemostasis and coagulation related proteins in our study. A potent endothelium dependent vasodilator peptide, bradykinin, was observed. Bradykinin was known to be a cancer growth factor, and has been tested as anticancer agents [40]. We found that bradykinin and desArg-bradykinin levels showed lower level in metastatic tumor groups, especially in the group of day 14. This can be explained by the fact that bradykinin is required for tumor growth, being the first 2 weeks, after which tumor growth slowed down and thus allowing bradykinin levels to increase again.
This study illustrates that the use of our nanopore based silica chip combined with mass spectrometry is an attractive method for the enrichment and identification of the low molecular weight fraction of blood serum, and also confirms the stages of tumor progression can be reflected more accurately by the cooperative profiling of multiple peptides. With the need of therapeutic evaluation and personalized diagnosis and treatment, a peptide signature is highly desirable. Our study revealed peptide signatures related pulmonary metastatic melanoma development (Figure 4 and 5). It provides the great potential to use these peptide signatures for clinical diagnosis. By further optimizing the pore size, structure and chemical characteristics, we can possibly use nanopore-based chips to selectively enrich for certain types of peptides, e.g. phospho- or glycopeptides. In addition, it is an ideal tool for clinical application due to the low cost, low sample amount required and simple pretreatment.
Acknowledgement
The work at the Methodist Hospital Research Institute (TMHRI) was primarily supported by research funding provided by TMHRI. The authors also acknowledge the support from the following grants: Alliance of Nanohealth pre-center award - DoD W81XWH-11-2-0168, DOD W81XWH-09-1-0212, NIH U54CA151668. Authors also like to thank Dr. David A. Engler at Proteomics core of TMHRI and Dr. David Hawke in Proteomic facilities at the University of Texas M.D. Anderson Cancer Center for their help in operating mass spectroscopy and suggestion in data analysis.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- [1].Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012;62:10–29. doi: 10.3322/caac.20138. [DOI] [PubMed] [Google Scholar]
- [2].Longo C, Gambara G, Espina V, Luchini A, Bishop B, Patanarut AS, Petricoin EF, 3rd, Beretti F, Ferrari B, Garaci E, De Pol A, Pellacani G, Liotta LA. A novel biomarker harvesting nanotechnology identifies Bak as a candidate melanoma biomarker in serum. Exp Dermatol. 2011;20:29–34. doi: 10.1111/j.1600-0625.2010.01187.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [3].Markovic SN, Erickson LA, Rao RD, Weenig RH, Pockaj BA, Bardia A, Vachon CM, Schild SE, McWilliams RR, Hand JL, Laman SD, Kottschade LA, Maples WJ, Pittelkow MR, Pulido JS, Cameron JD, Creagan ET. Malignant melanoma in the 21st century, part 2: staging, prognosis, and treatment. Mayo Clin Proc. 2007;82:490–513. doi: 10.4065/82.4.490. [DOI] [PubMed] [Google Scholar]
- [4].Balch CM, Gershenwald JE, Soong SJ, Thompson JF, Atkins MB, Byrd DR, Buzaid AC, Cochran AJ, Coit DG, Ding S, Eggermont AM, Flaherty KT, Gimotty PA, Kirkwood JM, McMasters KM, Mihm MC, Jr., Morton DL, Ross MI, Sober AJ, Sondak VK. Final version of 2009 AJCC melanoma staging and classification. J Clin Oncol. 2009;27:6199–6206. doi: 10.1200/JCO.2009.23.4799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [5].Weeraratna AT, Jiang Y, Hostetter G, Rosenblatt K, Duray P, Bittner M, Trent JM. Wnt5a signaling directly affects cell motility and invasion of metastatic melanoma. Cancer Cell. 2002;1:279–288. doi: 10.1016/s1535-6108(02)00045-4. [DOI] [PubMed] [Google Scholar]
- [6].Knol AC, Nguyen JM, Quereux G, Brocard A, Khammari A, Dreno B. Prognostic value of tumor-infiltrating Foxp3+ T-cell subpopulations in metastatic melanoma. Exp Dermatol. 2011;20:430–434. doi: 10.1111/j.1600-0625.2011.01260.x. [DOI] [PubMed] [Google Scholar]
- [7].Hegde UP, Chakraborty N, Kerr P, Grant-Kels JM. Melanoma in the elderly patient: relevance of the aging immune system. Clin Dermatol. 2009;27:537–544. doi: 10.1016/j.clindermatol.2008.09.012. [DOI] [PubMed] [Google Scholar]
- [8].Polsky D, Cordon-Cardo C. Oncogenes in melanoma. Oncogene. 2003;22:3087–3091. doi: 10.1038/sj.onc.1206449. [DOI] [PubMed] [Google Scholar]
- [9].Sabel MS, Liu Y, Lubman DM. Proteomics in melanoma biomarker discovery: great potential, many obstacles. Int J Proteomics. 2011;2011:181890. doi: 10.1155/2011/181890. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [10].Tarro G, Perna A, Esposito C. Early diagnosis of lung cancer by detection of tumor liberated protein. J Cell Physiol. 2005;203:1–5. doi: 10.1002/jcp.20195. [DOI] [PubMed] [Google Scholar]
- [11].Indovina P, Marcelli E, Pentimalli F, Tanganelli P, Tarro G, Giordano A. Mass spectrometry-based proteomics: The road to lung cancer biomarker discovery. Mass Spectrom Rev. 2012 doi: 10.1002/mas.21355. [DOI] [PubMed] [Google Scholar]
- [12].Beachy SH, Repasky EA. Using extracellular biomarkers for monitoring efficacy of therapeutics in cancer patients: an update. Cancer Immunol Immun. 2008;57:759–775. doi: 10.1007/s00262-007-0445-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [13].Tanaka Y, Akiyama H, Kuroda T, Jung G, Tanahashi K, Sugaya H, Utsumi J, Kawasaki H, Hirano H. A novel approach and protocol for discovering extremely low-abundance proteins in serum. Proteomics. 2006;6:4845–4855. doi: 10.1002/pmic.200500774. [DOI] [PubMed] [Google Scholar]
- [14].Anderson NL, Anderson NG. The human plasma proteome: history, character, and diagnostic prospects. Mol Cell Proteomics. 2002;1:845–867. doi: 10.1074/mcp.r200007-mcp200. [DOI] [PubMed] [Google Scholar]
- [15].Tirumalai RS, Chan KC, Prieto DA, Issaq HJ, Conrads TP, Veenstra TD. Characterization of the low molecular weight human serum proteome. Mol Cell Proteomics. 2003;2:1096–1103. doi: 10.1074/mcp.M300031-MCP200. [DOI] [PubMed] [Google Scholar]
- [16].Chertov O, Simpson JT, Biragyn A, Conrads TP, Veenstra TD, Fisher RJ. Enrichment of low-molecular-weight proteins from biofluids for biomarker discovery. Expert Rev Proteomics. 2005;2:139–145. doi: 10.1586/14789450.2.1.139. [DOI] [PubMed] [Google Scholar]
- [17].Kawashima Y, Fukutomi T, Tomonaga T, Takahashi H, Nomura F, Maeda T, Kodera Y. High-yield peptide-extraction method for the discovery of subnanomolar biomarkers from small serum samples. J Proteome Res. 2010;9:1694–1705. doi: 10.1021/pr9008018. [DOI] [PubMed] [Google Scholar]
- [18].Villanueva J, Martorella AJ, Lawlor K, Philip J, Fleisher M, Robbins RJ, Tempst P. Serum Peptidome Patterns That Distinguish Metastatic Thyroid Carcinoma from Cancer-free Controls Are Unbiased by Gender and Age. Molecular & Cellular Proteomics. 2006;5:1840–1852. doi: 10.1074/mcp.M600229-MCP200. [DOI] [PubMed] [Google Scholar]
- [19].Villanueva J, Shaffer DR, Philip J, Chaparro CA, Erdjument-Bromage H, Olshen AB, Fleisher M, Lilja H, Brogi E, Boyd J, Sanchez-Carbayo M, Holland EC, Cordon-Cardo C, Scher HI, Tempst P. Differential exoprotease activities confer tumor-specific serum peptidome patterns. J Clin Invest. 2006;116:271–284. doi: 10.1172/JCI26022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [20].Hu Y, Bouamrani A, Tasciotti E, Li L, Liu X, Ferrari M. Tailoring of the nanotexture of mesoporous silica films and their functionalized derivatives for selectively harvesting low molecular weight protein. ACS Nano. 2010;4:439–451. doi: 10.1021/nn901322d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [21].Hu Y, Peng Y, Lin K, Shen H, Brousseau LC, 3rd, Sakamoto J, Sun T, Ferrari M. Surface engineering on mesoporous silica chips for enriching low molecular weight phosphorylated proteins. Nanoscale. 2011;3:421–428. doi: 10.1039/c0nr00720j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [22].Bouamrani A, Hu Y, Tasciotti E, Li L, Chiappini C, Liu X, Ferrari M. Mesoporous silica chips for selective enrichment and stabilization of low molecular weight proteome. Proteomics. 2010;10:496–505. doi: 10.1002/pmic.200900346. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [23].Fan J, Deng X, Gallagher JW, Huang H, Huang Y, Wen J, Ferrari M, Shen H, Hu Y. Monitoring the progression of metastatic breast cancer on nanoporous silica chips. Philos Transact A Math Phys Eng Sci. 2012;370:2433–2447. doi: 10.1098/rsta.2011.0444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [24].Damsky WE, Rosenbaum LE, Bosenberg M. Decoding Melanoma Metastasis. Cancers. 2010;3:126–163. doi: 10.3390/cancers3010126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [25].Wang RF, Appella E, Kawakami Y, Kang XQ, Rosenberg SA. Identification of TRP-2 as a human tumor antigen recognized by cytotoxic T lymphocytes. Journal of Experimental Medicine. 1996;184:2207–2216. doi: 10.1084/jem.184.6.2207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [26].Bloom MB, PerryLalley D, Robbins PF, Li Y, ElGamil M, Rosenberg SA, Yang JC. Identification of tyrosinase-related protein 2 as a tumor rejection antigen for the B16 melanoma. Journal of Experimental Medicine. 1997;185:453–459. doi: 10.1084/jem.185.3.453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [27].Lam K.W.-k., Lo S.C.-l. Discovery of diagnostic serum biomarkers of gastric cancer using proteomics. PROTEOMICS – Clinical Applications. 2008;2:219–228. doi: 10.1002/prca.200780015. [DOI] [PubMed] [Google Scholar]
- [28].Guo R, Pan C, Shen J, Liu C. New serum biomarkers for detection of esophageal carcinoma using Matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. J Cancer Res Clin Oncol. 2011;137:513–519. doi: 10.1007/s00432-010-0899-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [29].Schwamborn K, Krieg RC, Grosse J, Reulen N, Weiskirchen R, Knuechel R, Jakse G, Henkel C. Serum proteomic profiling in patients with bladder cancer. Eur Urol. 2009;56:989–996. doi: 10.1016/j.eururo.2009.02.031. [DOI] [PubMed] [Google Scholar]
- [30].Mian S, Ugurel S, Parkinson E, Schlenzka I, Dryden I, Lancashire L, Ball G, Creaser C, Rees R, Schadendorf D. Serum Proteomic Fingerprinting Discriminates Between Clinical Stages and Predicts Disease Progression in Melanoma Patients. Journal of Clinical Oncology. 2005;23:5088–5093. doi: 10.1200/JCO.2005.03.164. [DOI] [PubMed] [Google Scholar]
- [31].Diamandis EP. Peptidomics for cancer diagnosis: present and future. J Proteome Res. 2006;5:2079–2082. doi: 10.1021/pr060225u. [DOI] [PubMed] [Google Scholar]
- [32].Markiewski MM, Lambris JD. Unwelcome complement. Cancer Res. 2009;69:6367–6370. doi: 10.1158/0008-5472.CAN-09-1918. [DOI] [PMC free article] [PubMed] [Google Scholar]
- [33].Rutkowski MJ, Sughrue ME, Kane AJ, Mills SA, Parsa AT. Cancer and the complement cascade. Mol Cancer Res. 2010;8:1453–1465. doi: 10.1158/1541-7786.MCR-10-0225. [DOI] [PubMed] [Google Scholar]
- [34].Jean D, Bar-Eli M, Huang S, Xie K, Rodrigues-Lima F, Hermann J, Frade R. A cysteine proteinase, which cleaves human C3, the third component of complement, is involved in tumorigenicity and metastasis of human melanoma. Cancer Res. 1996;56:254–258. [PubMed] [Google Scholar]
- [35].Mosesson MW. Fibrinogen and fibrin structure and functions. J Thromb Haemost. 2005;3:1894–1904. doi: 10.1111/j.1538-7836.2005.01365.x. [DOI] [PubMed] [Google Scholar]
- [36].Doolittle RF. Fibrinogen and fibrin. Sci Am. 1981;245:126–135. doi: 10.1038/scientificamerican1281-126. [DOI] [PubMed] [Google Scholar]
- [37].Bootle-Wilbraham CA, Tazzyman S, Marshall JM, Lewis CE. Fibrinogen E-fragment inhibits the migration and tubule formation of human dermal microvascular endothelial cells in vitro. Cancer Res. 2000;60:4719–4724. [PubMed] [Google Scholar]
- [38].Du J, Zheng JH, Chen XS, Yang Q, Zhang YH, Zhou L, Yao X. High preoperative plasma fibrinogen is an independent predictor of distant metastasis and poor prognosis in renal cell carcinoma. Int J Clin Oncol. 2012 doi: 10.1007/s10147-012-0412-x. [DOI] [PubMed] [Google Scholar]
- [39].Palumbo JS, Kombrinck KW, Drew AF, Grimes TS, Kiser JH, Degen JL, Bugge TH. Fibrinogen is an important determinant of the metastatic potential of circulating tumor cells. Blood. 2000;96:3302–3309. [PubMed] [Google Scholar]
- [40].Stewart JM. Bradykinin antagonists as anti-cancer agents. Curr Pharm Des. 2003;9:2036–2042. doi: 10.2174/1381612033454171. [DOI] [PubMed] [Google Scholar]