Abstract
Using deletion mutagenesis we localized the promoter for the light strand of mouse mitochondrial DNA to a 97-base-pair region, from -88 to +9 nucleotides of the transcriptional initiation site. Within this region the light-strand promoter could be dissected into at least three different functional domains. The specificity region, a maximum of 19 base pairs between -10 and +9 of the transcriptional initiation site, was essential and sufficient for accurate transcriptional initiation. A second region, extending to -29 nucleotides from the initiation site, facilitated the formation of a preinitiation complex between the template DNA and factor(s) present in the mitochondrial RNA polymerase fraction and was required for efficient transcription. A third, ill-defined upstream region, which extended up to -88 nucleotides from the initiation site, appeared to influence template transcriptional efficiencies in competition assays. Without the specificity domain, the upstream regions were incapable of supporting any transcription. The presence of multiple upstream domains was confirmed by disrupting nucleotide sequences in the upstream region by using linker insertion and linker replacement techniques.
Full text
PDF








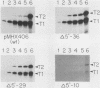
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Anderson S., Bankier A. T., Barrell B. G., de Bruijn M. H., Coulson A. R., Drouin J., Eperon I. C., Nierlich D. P., Roe B. A., Sanger F. Sequence and organization of the human mitochondrial genome. Nature. 1981 Apr 9;290(5806):457–465. doi: 10.1038/290457a0. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Van Etten R. A., Wright C. T., Walberg M. W., Clayton D. A. Sequence and gene organization of mouse mitochondrial DNA. Cell. 1981 Oct;26(2 Pt 2):167–180. doi: 10.1016/0092-8674(81)90300-7. [DOI] [PubMed] [Google Scholar]
- Bieker J. J., Martin P. L., Roeder R. G. Formation of a rate-limiting intermediate in 5S RNA gene transcription. Cell. 1985 Jan;40(1):119–127. doi: 10.1016/0092-8674(85)90315-0. [DOI] [PubMed] [Google Scholar]
- Biswas T. K., Edwards J. C., Rabinowitz M., Getz G. S. Characterization of a yeast mitochondrial promoter by deletion mutagenesis. Proc Natl Acad Sci U S A. 1985 Apr;82(7):1954–1958. doi: 10.1073/pnas.82.7.1954. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bogenhagen D. F., Wormington W. M., Brown D. D. Stable transcription complexes of Xenopus 5S RNA genes: a means to maintain the differentiated state. Cell. 1982 Feb;28(2):413–421. doi: 10.1016/0092-8674(82)90359-2. [DOI] [PubMed] [Google Scholar]
- Chang D. D., Clayton D. A. Identification of primary transcriptional start sites of mouse mitochondrial DNA: accurate in vitro initiation of both heavy- and light-strand transcripts. Mol Cell Biol. 1986 May;6(5):1446–1453. doi: 10.1128/mcb.6.5.1446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang D. D., Clayton D. A. Precise identification of individual promoters for transcription of each strand of human mitochondrial DNA. Cell. 1984 Mar;36(3):635–643. doi: 10.1016/0092-8674(84)90343-x. [DOI] [PubMed] [Google Scholar]
- Chomyn A., Mariottini P., Cleeter M. W., Ragan C. I., Matsuno-Yagi A., Hatefi Y., Doolittle R. F., Attardi G. Six unidentified reading frames of human mitochondrial DNA encode components of the respiratory-chain NADH dehydrogenase. Nature. 1985 Apr 18;314(6012):592–597. doi: 10.1038/314592a0. [DOI] [PubMed] [Google Scholar]
- Christianson T., Rabinowitz M. Identification of multiple transcriptional initiation sites on the yeast mitochondrial genome by in vitro capping with guanylyltransferase. J Biol Chem. 1983 Nov 25;258(22):14025–14033. [PubMed] [Google Scholar]
- Ciliberto G., Raugei G., Costanzo F., Dente L., Cortese R. Common and interchangeable elements in the promoters of genes transcribed by RNA polymerase iii. Cell. 1983 Mar;32(3):725–733. doi: 10.1016/0092-8674(83)90058-2. [DOI] [PubMed] [Google Scholar]
- Clayton D. A. Transcription of the mammalian mitochondrial genome. Annu Rev Biochem. 1984;53:573–594. doi: 10.1146/annurev.bi.53.070184.003041. [DOI] [PubMed] [Google Scholar]
- Dierks P., van Ooyen A., Cochran M. D., Dobkin C., Reiser J., Weissmann C. Three regions upstream from the cap site are required for efficient and accurate transcription of the rabbit beta-globin gene in mouse 3T6 cells. Cell. 1983 Mar;32(3):695–706. doi: 10.1016/0092-8674(83)90055-7. [DOI] [PubMed] [Google Scholar]
- Engelke D. R., Ng S. Y., Shastry B. S., Roeder R. G. Specific interaction of a purified transcription factor with an internal control region of 5S RNA genes. Cell. 1980 Mar;19(3):717–728. doi: 10.1016/s0092-8674(80)80048-1. [DOI] [PubMed] [Google Scholar]
- Fisher R. P., Clayton D. A. A transcription factor required for promoter recognition by human mitochondrial RNA polymerase. Accurate initiation at the heavy- and light-strand promoters dissected and reconstituted in vitro. J Biol Chem. 1985 Sep 15;260(20):11330–11338. [PubMed] [Google Scholar]
- Levens D., Lustig A., Rabinowitz M. Purification of mitochondrial RNA polymerase from Saccharomyces cerevisiae. J Biol Chem. 1981 Feb 10;256(3):1474–1481. [PubMed] [Google Scholar]
- Levens D., Morimoto R., Rabinowitz M. Mitochondrial transcription complex from Saccharomyces cerevisiae. J Biol Chem. 1981 Feb 10;256(3):1466–1472. [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- McKnight S. L., Gavis E. R., Kingsbury R., Axel R. Analysis of transcriptional regulatory signals of the HSV thymidine kinase gene: identification of an upstream control region. Cell. 1981 Aug;25(2):385–398. doi: 10.1016/0092-8674(81)90057-x. [DOI] [PubMed] [Google Scholar]
- Montoya J., Christianson T., Levens D., Rabinowitz M., Attardi G. Identification of initiation sites for heavy-strand and light-strand transcription in human mitochondrial DNA. Proc Natl Acad Sci U S A. 1982 Dec;79(23):7195–7199. doi: 10.1073/pnas.79.23.7195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Sollner-Webb B., Wilkinson J. A., Roan J., Reeder R. H. Nested control regions promote Xenopus ribosomal RNA synthesis by RNA polymerase I. Cell. 1983 Nov;35(1):199–206. doi: 10.1016/0092-8674(83)90222-2. [DOI] [PubMed] [Google Scholar]
- Sommerville J. RNA polymerase I promoters and transcription factors. Nature. 1984 Jul 19;310(5974):189–190. doi: 10.1038/310189a0. [DOI] [PubMed] [Google Scholar]
- Suissa M. Spectrophotometric quantitation of silver grains eluted from autoradiograms. Anal Biochem. 1983 Sep;133(2):511–514. doi: 10.1016/0003-2697(83)90117-3. [DOI] [PubMed] [Google Scholar]
- Walberg M. W., Clayton D. A. In vitro transcription of human mitochondrial DNA. Identification of specific light strand transcripts from the displacement loop region. J Biol Chem. 1983 Jan 25;258(2):1268–1275. [PubMed] [Google Scholar]
- Winkley C. S., Keller M. J., Jaehning J. A. A multicomponent mitochondrial RNA polymerase from Saccharomyces cerevisiae. J Biol Chem. 1985 Nov 15;260(26):14214–14223. [PubMed] [Google Scholar]
- Yoza B. K., Bogenhagen D. F. Identification and in vitro capping of a primary transcript of human mitochondrial DNA. J Biol Chem. 1984 Mar 25;259(6):3909–3915. [PubMed] [Google Scholar]