Abstract
Group II introns are self-splicing catalytic RNAs that are thought to be ancestral to the spliceosome. Here we report the 3.65 Å crystal structure of the group II intron from Oceanobacillus iheyensis in the pre-catalytic state. The structure reveals the conformation of the 5′ splice site in the catalytic core and represents the first structure of an intron prior to the first step of splicing.
Group II introns are mobile retroelements that have a self-splicing catalytic RNA component thought to be evolutionarily related to the spliceosome.1 Group II introns are postulated to have first originated in bacteria billions of years ago. These primitive molecules then invaded the first complex cells of eukaryotes, which subsequently evolved into plants, animals, and humans.2 In eukaryotes, the dual functionalities of splicing and mobility became separated into the spliceosomal introns and retroelements. These genetic elements comprise at least 50% of most mammalian genomes and have contributed greatly to genomic diversity.3 Because of this evolutionary impact, group II introns have been used as a tractable model system to gain insight into the structure and function of their eukaryotic descendants.
Group II intron RNAs catalyze splicing via two distinct transesterification events.4 In the first step, nucleophilic attack occurs at the 5′ splice site by the 2′-OH of a bulged adenosine within the intron, or by a water molecule from solution. This results in the formation of an intron-3′-exon intermediate. This is followed by the second step in which the 3′-OH of the newly cleaved 5′ exon attacks the 3′ splice site to form ligated exons and a liberated intron. During splicing, group II introns use exon binding sequences called EBS1 and EBS3 to properly position the 5′ and 3′ splice sites, respectively, in the catalytic core.5,6 The 3′ splice site has the additional structural requirement that the 3′ terminal residue of the intron must engage in the γ-γ′ interaction, which helps anchor the 3′ splice site in the core.7 The γ-γ′ interaction contains a single base pair between the terminal 3′ intron nucleotide (γ′) and a residue located between domains II and III of the intron (γ).
In this study, we set out to determine the structure of the group II intron in a pre-catalytic state directly preceding the first step of splicing. Previously, we had solved the crystal structure of a group II intron from O. iheyensis in the post-catalytic state.8,9 This structure revealed that splicing occurs through a two-metal-ion mechanism catalyzed by residues within the highly conserved intron domain V. In addition, the scissile phosphate of bound ligated exon substrate was found to be positioned directly over the active site. In this post-catalytic structure, the junction between the two exons probably represents a relaxed state that exists after the intron completes both steps of splicing, immediately before exon release. RNA splicing is a dynamic process involving multiple conformational changes to properly position substrates in the active site. In order to understand and visualize the different stages of splicing, it is necessary to trap the intron at each stage and determine its structure using X-ray crystallography. In particular, the pre-catalytic state has never been visualized in any intron splicing system.
Here we report the 3.65 Å crystal structure of the O. iheyensis intron trapped in the pre-splicing state, before the onset of catalysis. This represents the first structure of an intron before splicing and allows us to visualize the position and conformation of the 5′ splice site within the catalytic core of the group II intron. In order to trap the intron at this stage and prevent cleavage of the 5′ splice site, a splicing precursor construct of the O. iheyensis intron was deactivated through mutagenesis of the central guanosine of the 5′-CGC-3′ catalytic triad. This mutation changes the G•U wobble to an A-U pair, thereby maintaining pairing within the domain V helix, and minimizing potential disruption of the RNA structure. The mutant showed no detectable splicing activity when denaturing PAGE was used, however primer extension analysis revealed ~15% cleavage of the 5′ exon upon extended storage of the RNA at 4°C (see Supplementary Discussion). We transcribed and purified intron RNA in vitro, using a native purification procedure (see Supplementary Methods). In this procedure, the intron folds during in vitro transcription and is purified in a native state by using diafiltration.8 The intron RNA crystallized in space group P212121, and the structure was refined to Rwork=20.5% and Rfree=24.9% (Supplementary Table 1, PDB accession 4DS6).
The overall fold of the pre-catalytic group II intron is markedly similar to that of the post-catalytic state (Fig. 1a), indicating that no large-scale conformational changes occur in domains I to V during the transition between the two steps of splicing. In the core of the ribozyme (Fig. 1b,c and Supplementary Figs. 1 and 2), strong electron density is now visible for the 5′ splice site docked into the catalytic pocket of domain V. The 5′ exon is paired with EBS1 indicating that this interaction persists throughout the splicing reaction into the post-catalytic state. The junction between the 5′ exon and the intron undergoes a marked reversal in the direction of the RNA chain. This produces a sharp kink in the backbone that presents the scissile phosphate of the splice site to the active site in domain V. Thus, the structure reveals the requirement for a structural distortion of the 5′ splice site just prior to catalysis. This also results in an unusual arrangement of two highly distorted backbones in close proximity to each other with the two-nucleotide bulge region of domain V being very near to the 5′ splice site (Supplementary Fig. 3). The position of the 5′ splice site within this crystal structure is entirely consistent with previous cross-linking data on the unspliced (pre-catalytic) forms of the aI5γ and L.l.ltrB group II introns.10–12 Based on the cumulative biochemical data, the 5′ splice site had been previously modeled in close proximity to the active site in domain V.6,10,12–15 The kinked 5′ splice site observed in our crystal structure was first proposed in 2005 (refs. 10, 15). Even though the scissile phosphate is positioned in the active site, no native density is visible for the catalytic metal ions. The most plausible explanation is that the mutation of the catalytic triad abolished binding of these metals. Analysis of the backbone of domain V in both the wild-type (post-catalytic) and mutant (pre-catalytic) introns reveals that the G359A mutation causes a slight shift in the active site that may abrogate binding of the catalytic metal ions (Supplementary Discussion and Supplementary Fig. 4). It should be emphasized that the exon-binding sequences within the intron are responsible for the kinking of the 5′ splice site and that these residues remain as the wild-type sequence. Therefore, binding of the catalytic metal ions is not expected to alter the conformation of the 5′ splice site.
Figure 1.
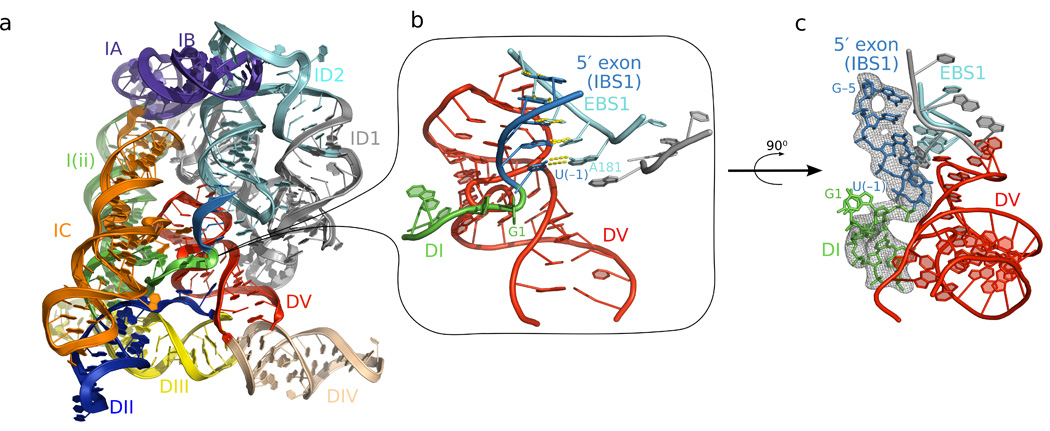
The pre-catalytic structure of the group II intron. a) The structure of the pre-catalytic state reveals the position of the 5′ splice site (junction between green and blue) in the center of the molecule. b) Close-up view of the 5′ splice site reveals a sharp kink in the backbone, which is positioned near the bulge and catalytic triad of domain V (red). c) Fo-Fc density for the 5′ splice site contoured at 3σ. The Fo-Fc density map was calculated using a model deleted for intron nucleotides 1–5 and the 5′ exon in order to avoid model bias.
At the 5′ splice site, the first two nucleotides of the intron (G1 and U2) engage in a base-stacking interaction that probably stabilizes the position of the backbone in this single-stranded region. Unlike the post-catalytic structure, there is now electron density for the nucleobase of the γ residue (A287), which forms a continuous base stack with the domain V bulge nucleotide A376 and the G5 residue from the 5′ terminus (Supplementary Fig. 5). A287 is functionally important because it pairs to the 3′ terminal nucleotide of the intron and has a crucial role in positioning the 3′ splice site. The presence of the A287 nucleobase now enables us to model the probable position of the 3′ splice site within the catalytic core. Together with the pre- and post-catalytic structures, we have conceptualized a theoretical model of the complete group II intron-splicing pathway (Fig. 2). In this model, the 5′ splice site is first drawn near the catalytic metal ions and the kink is induced (Fig. 2a). This allows the nucleophile to attack and cleave the 5′ exon-intron junction. This is immediately followed by entry of the 3′ splice site to the active site. Nucleotides at the 3′ splice site are expected to pair simultaneously to both the γ nucleotide and EBS3 (Fig. 2b), resulting in a bifurcated set of interactions that are likely to kink the 3′ splice site. This arrangement is also consistent with previous biochemical data regarding the involvement of these tertiary interactions during 3′ splice site selection by group II introns.6,7,16 The resulting conformation enables the 3′ splice site to approach the active site at an angle of attack that is ~90 degrees from that of the 5′ splice site. As a result, the 3′-OH of the 5′ exon is located in close proximity to both the scissile phosphate of the 3′ splice site and one of the catalytic metal ions. This is consistent with a reaction mechanism in which this hydroxyl group is activated by a metal ion and engages in nucleophilic attack at the 3′ splice site with subsequent ligation of the exons.17 After ligation, the junction between the 5′ and 3′ exons exists in a relaxed state prior to product release (Fig. 2c).
Figure 2.

Theoretical model for the complete group II intron splicing pathway. a) The 5′ splice site is kinked immediately before the onset of catalysis. The kink is positioned in close proximity to the catalytic metal ions (M1 and M2). A ribose 2′-OH group or water molecule (not depicted) is activated for nucleophilic attack and cleaves the splice site. Residue 288 is not shown because there is no electron density for the base of this nucleotide in the pre-catalytic structure. b) The 3′ splice site is positioned in the active site through its interaction with EBS3 and the γ nucleotide. These tertiary interactions cause the 3′ splice site to also adopt a sharp kink which presents the scissile phosphate to the active site metal ions. The 3′-OH (shown in stick format) of the 5′ exon is in a position to coordinate to M1 as well as to the 3′ splice site (coordination indicated by black lines). G1 and U2 are not shown due to the fact that these nucleotides must depart before the 3′ splice site enters the active site. c) The 3′ splice site is cleaved, the exons ligated, and the product adopts a “relaxed” conformation prior to release by the intron. Abbreviation: J2/3 - junction sequence between domains II and III.
This theoretical model suggests that group II intron active sites are versatile and can accommodate both steps of splicing with small conformational alterations. Given their mechanistic similarities, it is likely that both group II introns and the spliceosome induce sharp bends at the splice sites for proper positioning of the scissile phosphates. The generality of this behavior is underscored by the actions of the evolutionarily unrelated group I introns, which induce a sharp kink at the 3′ splice site in constructs designed to capture the second step of splicing.18 Therefore, the sharp structural distortion of splice sites may represent a common mechanism of all RNA splicing reactions.
Supplementary Material
Acknowledgements
We thank the staff of the NE-CAT beamline ID-24 at the Advanced Photon Source of Argonne National Laboratory. We thank Kevin Keating for valuable discussions. This work was supported by startup funds from the University of California, San Diego (N.T.) and the Howard Hughes Medical Institute (A.M.P.). R.T.C. was supported by the Cell, Molecular, and Genetics (CMG) Training Program funded by NIH predoctoral training grant T32GM007240.
Footnotes
Accession Codes. Protein Data Bank: coordinates and structure factors have been deposited under accession code 4DS6.
Author Contributions:
R.T.C., A.R.R., K.R.R. and N.T. performed experiments. R.T.C., A.R.R. and N.T. designed the experiments. R.T.C., A.R.R., K.R.R., A.M.P. and N.T. analyzed the data. R.T.C. and N.T. wrote the manuscript with input from all authors
References
- 1.Sharp PA. Five easy pieces. Science. 1991;254:663. doi: 10.1126/science.1948046. [DOI] [PubMed] [Google Scholar]
- 2.Martin W, Koonin EV. Introns and the origin of nucleus-cytosol compartmentalization. Nature. 2006;440:41–45. doi: 10.1038/nature04531. [DOI] [PubMed] [Google Scholar]
- 3.Koonin EV. Intron-dominated genomes of early ancestors of eukaryotes. J Hered. 2009;100:618–623. doi: 10.1093/jhered/esp056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Robart AR, Zimmerly S. Group II intron retroelements: function and diversity. Cytogenet Genome Res. 2005;110:589–597. doi: 10.1159/000084992. [DOI] [PubMed] [Google Scholar]
- 5.Jacquier A, Michel F. Multiple exon-binding sites in class II self-splicing introns. Cell. 1987;50:17–29. doi: 10.1016/0092-8674(87)90658-1. [DOI] [PubMed] [Google Scholar]
- 6.Costa M, Michel F, Westhof E. A three-dimensional perspective on exon binding by a group II self-splicing intron. EMBO J. 2000;19:5007–5018. doi: 10.1093/emboj/19.18.5007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Jacquier A, Michel F. Base-pairing interactions involving the 5′ and 3′-terminal nucleotides of group II self-splicing introns. J Mol Biol. 1990;213:437–447. doi: 10.1016/S0022-2836(05)80206-2. [DOI] [PubMed] [Google Scholar]
- 8.Toor N, Keating KS, Taylor SD, Pyle AM. Crystal structure of a self-spliced group II intron. Science. 2008;320:77–82. doi: 10.1126/science.1153803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Toor N, Rajashankar K, Keating KS, Pyle AM. Structural basis for exon recognition by a group II intron. Nat Struct Mol Biol. 2008;15:1221–1222. doi: 10.1038/nsmb.1509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.de Lencastre A, Hamill S, Pyle AM. A single active-site region for a group II intron. Nat Struct Mol Biol. 2005;12:626–627. doi: 10.1038/nsmb957. [DOI] [PubMed] [Google Scholar]
- 11.Noah JW, Lambowitz AM. Effects of maturase binding and Mg2+ concentration on group II intron RNA folding investigated by UV cross-linking. Biochemistry. 2003;42:12466–12480. doi: 10.1021/bi035339n. [DOI] [PubMed] [Google Scholar]
- 12.Dai L, et al. A three-dimensional model of a group II intron RNA and its interaction with the intron-encoded reverse transcriptase. Mol Cell. 2008;30:472–485. doi: 10.1016/j.molcel.2008.04.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Boudvillain M, de Lencastre A, Pyle AM. A tertiary interaction that links active-site domains to the 5′ splice site of a group II intron. Nature. 2000;406:315–318. doi: 10.1038/35018589. [DOI] [PubMed] [Google Scholar]
- 14.Wang J. Inclusion of weak high-resolution X-ray data for improvement of a group II intron structure. Acta Crystallogr D Biol Crystallogr. 2010;66:988–1000. doi: 10.1107/S0907444910029938. [DOI] [PubMed] [Google Scholar]
- 15.de Lencastre A, Pyle AM. Three essential and conserved regions of the group II intron are proximal to the 5′-splice site. RNA. 2008;14:11–24. doi: 10.1261/rna.774008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Robart AR, Montgomery NK, Smith KL, Zimmerly S. Principles of 3′ splice site selection and alternative splicing for an unusual group II intron from Bacillus anthracis. RNA. 2004;10:854–862. doi: 10.1261/rna.5246804. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Roitzsch M, Fedorova O, Pyle AM. The 2′-OH group at the group II intron terminus acts as a proton shuttle. Nat Chem Biol. 2010;6:218–224. doi: 10.1038/nchembio.312. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Adams PL, Stahley MR, Kosek AB, Wang J, Strobel SA. Crystal structure of a self-splicing group I intron with both exons. Nature. 2004;430:45–50. doi: 10.1038/nature02642. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.


