A PubMed search for HIV-related publications from 2011 yields a whopping 15 091 total, or over 40 studies a day. No one can read or remember all this; unsettlingly, no one knows the full HIV literature. The explosion of high throughput data makes the deluge even worse. Genome-wide small interfering RNA (siRNA) screens and proteomic scans regularly yield results for all 20 000 human genes. The Solexa/Illumina HiSeq (San Diego, California, USA) instrument can produce more than 100 billion bases of sequence information in a single instrument run, and runs are accumulating addressing questions in HIV research. Data from genome-wide association studies, HIV resistance testing and epidemiological tracking all add to the flood. Given this gigantic volume of data, the development of digestible summaries becomes as important as generating the data itself. Here we briefly list some of the main sources of data and efforts to develop new tools to work with them, particularly focusing on a new tool for analysing genome-wide screens for human genes affecting HIV replication. Links toweb sites mentioned in the article are collected in Table 1.
Table 1.
Examples of resources for HIV Systems Biology on the Web.
| Web resource | Link(s) | Description |
|---|---|---|
| NCBI | http://www.ncbi.nlm.nih.gov/ | The national repository of biomedial data and literature |
| Los Alamos HIV Databases | www.hiv.lanl.gov/ | Excellent repository of HIV sequence and epitope information |
| UNAIDS | http://www.unaids.org/en/ | Compiled information and policy materials from the UN |
| Stanford University HIV Drug Resistance Database | http://hivdb.stanford.edu/index.html | Database of mutations in HIV conferring resistance to antiviral agents |
| GPS-Prot | http://www.gpsprot.org | A site for exploring interactions between, HIV and cellular proteins, with rich tools for follow-up |
| This week in virology | http://www.virology.ws/ | Vince Racaniello’s weekly webcast on virology |
| HIV Replication Cycle | http://www.hivsystemsbiology.org/wiki/index.php/Introduction | A web-based account of the HIV replication cycle |
| Gene Overlapper | http://www.hivsystemsbiology.org/GeneListOverlapper/ | Data from genome-wide surveys together with tools for exploring overlaps |
NCBI, National Center for Biotechnology Information; UNAIDS, Joint United Nations Programme on HIV/AIDS.
Several groups have started to develop data repositories and computational tools for HIV research, providing important starting points. The National Center for Biotechnology Information (NCBI) hosts the largest effort, centralizing data on the scientific literature, DNA sequences, gene structure and many other topics [1]. NCBI serves an important archiving function, but retrieving high-throughput data is often challenging, and few analytical tools are available. The Los Alamos HIV Databases houses HIV sequences and data on mapped epitopes, together with useful alignments and some tools for working with the data [2]. The Stanford University HIV Drug Resistance Database provides a key resource for information on HIV mutations conferring resistance to antiviral agents [3]. GPS-Prot provides an innovative way for starting with an HIV protein, calling up well vetted binding proteins and exploring multiple types of annotation from there [4]. Vince Racaniello’s web show on virology is another favourite.
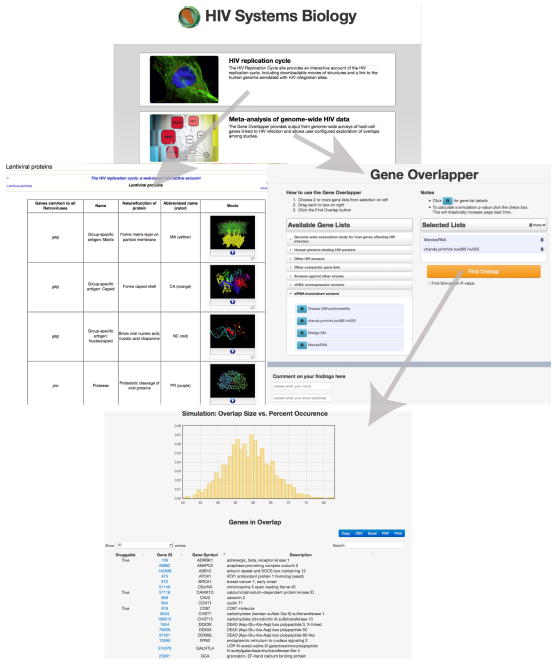
Here we introduce a new web site – HIVsystemsbiology.org – that collects Big Data on HIV and starts to provide tools to distil them (Fig. 1). One tool is the Gene Overlapper, which collects data from genome-wide screens of human gene products affecting HIVand enables analysis of overlap among sets. This is paired with a second resource, the HIV Replication Cycle site, which is accessible through HIVsystemsbiology.org and provides context for the genome-wide data. Unfortunately, large data sets often come with very limited background, greatly reducing their usefulness. Without detailed information on a sample’s origins, it is difficult to follow up with much confidence. Data need to be paired with summaries that are web based, rich in context and as inviting to users as possible, obvious from experience but surprisingly hard to implement.
Fig. 1. Progression through the HIV Systems Biology Site.
Arrows indicate links among web pages.
The HIV Replication Cycle site presents a review of HIV replication in cells, but linked to extensive web-based resources. Accounts of the HIV proteins are enhanced with movies of HIV protein structures to allow visualization in three dimensions. Numerous web links lead from the site to other resources. One link allows readers to navigate out on to the human genome and surf around, viewing positions of HIV integration sites. All images, movies and other materials are available for free download for use by AIDS researchers and educators. Importantly in this context, simple explanations of different parts of the HIV replication cycle are linked to large data sets available for study in the Gene Overlapper site.
The Overlapper site houses 39 lists of genes called in different genome-wide screens for links to HIV, and the number is growing steadily. Included are results from three genome-side siRNA screens [5–7] and a cDNA overexpression screen [8], allowing intersections among these gene sets to be explored. Other types of data may be of interest in further comparisons, for example data from screens for human proteins binding to HIV proteins. Nineteen lists from such screens are included [9]. Additional gene lists describe computational scans for gene products important in HIV replication [10], results of genome-wide association studies [11] and siRNA screens against other viruses (e.g. [12]). Each of these types of data have significant noise in addition to the signal, but filtering over multiple such screens can help clarify the best supported genes (e.g. [6]). Once genes are identified in lists, clicking on the gene name takes the user to detailed annotation at NCBI.
For each list, data are housed together with simple descriptions of what is in the list, and what background the genes in the list were drawn from. For published studies, the reference is included as a link to the PubMed entry, allowing easy access to detailed background information.
Once genes of interest are identified, lists can be overlapped against additional types of lists for more detailed interpretation. One goal, for example, might be identifying cellular targets for potential antiviral agents. For this, one might want to identify human genes that encode proteins from ‘druggable’ gene families, or genes that are dispensable when knocked out in mice and so might not be toxic when inhibited in humans. Each of these lists are available under ‘Other comparator gene lists’.
An important goal of this initiative is to ‘crowd-source’ discovery of human genes important in HIV biology. The Overlapper site currently contains 39 gene lists, which can be formed into more than 500 billion subsets. Obviously no single individual is going to look for overlaps among all the possible subsets by hand. Thus a blog is embedded in the Overlapper site, allowing users to list data sets tested and any interesting results they would like to share. The blog also collects feedback from users on new lists to add or suggestions for improving the site.
Last year crowd-sourcing resulted in the solution of the molecular structure of a retroviral protease [13]. X-ray analysis had been performed on crystals of the Mason-Pfizer monkey virus protease, and a low-resolution nuclear magnetic resonance (NMR) structure was available, but the quality of the NMR structure was insufficient to solve the phases in the X-ray data by molecular replacement. Gamers playing Foldit improved the fold inferred from NMR data, allowing researchers to obtain starting phases and ultimately solve the X-ray structure at a high resolution. For genome-wide screens, who knows? Maybe the tools could be made inviting enough that gamers as well as scientists could be lured into playing around with HIV data and identifying some important new host factors.
Acknowledgments
This work was supported by National Institutes of Health grants AI52845 and AI 090935.
Footnotes
Conflicts of interest
There are no conflicts of interest.
References
- 1.Sayers EW, Barrett T, Benson DA, Bolton E, Bryant SH, Canese K, et al. Database resources of the National Center for Biotechnology Information. Nucl Acids Res. 2012;40:D13–25. doi: 10.1093/nar/gkr1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kuiken C, Korber B, Shafer RW. HIV sequence databases. AIDS Rev. 2003;5:52–61. [PMC free article] [PubMed] [Google Scholar]
- 3.Tang MW, Liu TF, Shafer RW. The HIVdb system for HIV-1 genotypic resistance interpretation. Intervirology. 2012;55:98–101. doi: 10.1159/000331998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fahey ME, Bennett MJ, Mahon C, Jager S, Pache L, Kumar D, et al. GPS-Prot: a web-based visualization platform for integrating host-pathogen interaction data. BMC Bioinform. 2011;12:298. doi: 10.1186/1471-2105-12-298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Konig R, Zhou Y, Elleder D, Diamond TL, Bonamy GM, Irelan JT, et al. Global analysis of host-pathogen interactions that regulate early-stage HIV-1 replication. Cell. 2008;135:49–60. doi: 10.1016/j.cell.2008.07.032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bushman FD, Malani N, Fernandes J, D’Orso I, Cagney G, Diamond TL, et al. Host cell factors in HIV replication: meta-analysis of genome-wide studies. PLoS Path. 2009;5:e1000437. doi: 10.1371/journal.ppat.1000437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Zhou H, Xu M, Huang Q, Gates AT, Zhang XD, Castle JC, et al. Genome-scale RNAi screen for host factors required for HIV replication. Cell Host Microbe. 2008;4:495–504. doi: 10.1016/j.chom.2008.10.004. [DOI] [PubMed] [Google Scholar]
- 8.Nguyen DG, Yin H, Zhou Y, Wolff KC, Kuhen KL, Caldwell JS. Identification of novel therapeutic targets for HIV infection through functional genomic cDNA screening. Virology. 2007;362:16–25. doi: 10.1016/j.virol.2006.11.036. [DOI] [PubMed] [Google Scholar]
- 9.Jager S, Cimermancic P, Gulbahce N, Johnson JR, McGovern KE, Clarke SC, et al. Global landscape of HIV-human protein complexes. Nature. 2011;481:365–370. doi: 10.1038/nature10719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Evans P, Dampier W, Ungar L, Tozeren A. Prediction of HIV-1 virus-host protein interactions using virus and host sequence motifs. BMC Med Genomics. 2009;2:27. doi: 10.1186/1755-8794-2-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fellay J, Shianna KV, Ge D, Colombo S, Ledergerber B, Weale M, et al. A whole-genome association study of major determinants for host control of HIV-1. Science (New York, NY) 2007;317:944–947. doi: 10.1126/science.1143767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hao L, Sakurai A, Watanabe T, Sorensen E, Nidom CA, Newton MA, et al. Drosophila RNAi screen identifies host genes important for influenza virus replication. Nature. 2008;454:890–893. doi: 10.1038/nature07151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Khatib F, DiMaio F, Cooper S, Kazmierczyk M, Gilski M, Krzywda S, et al. Crystal structure of a monomeric retroviral protease solved by protein folding game players. Nat Struct Mol Biol. 2011;18:1175–1177. doi: 10.1038/nsmb.2119. [DOI] [PMC free article] [PubMed] [Google Scholar]