Abstract
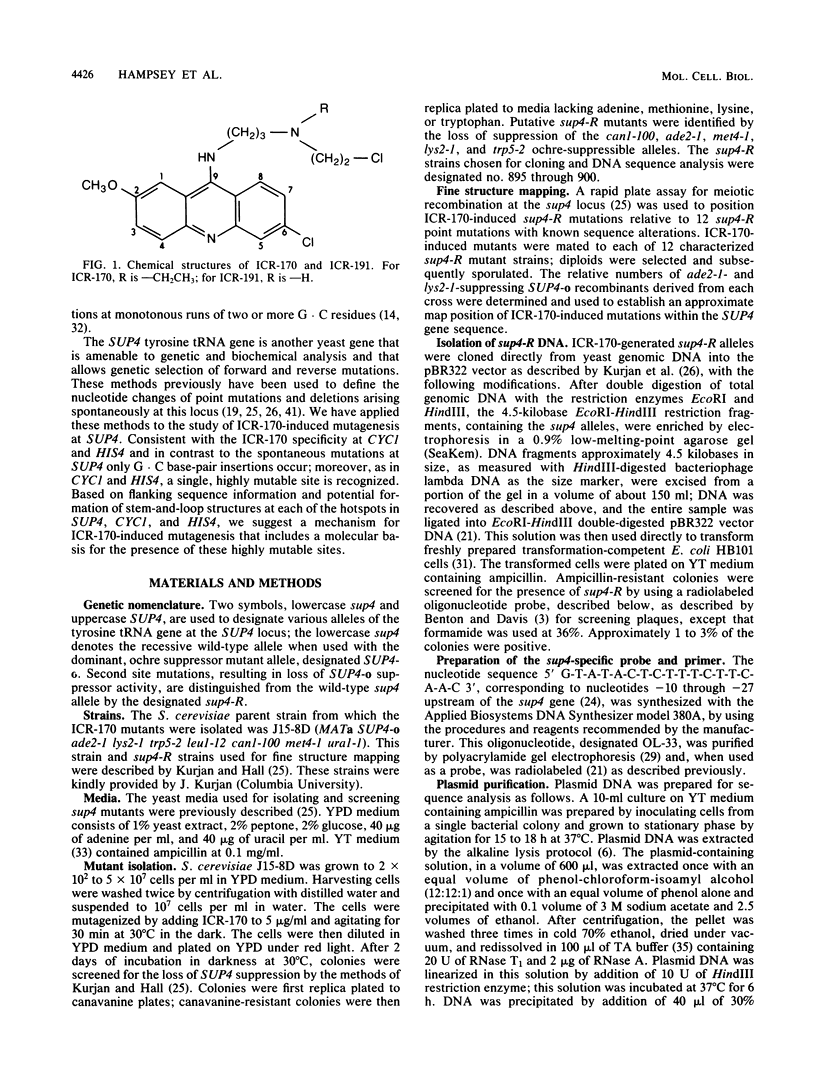
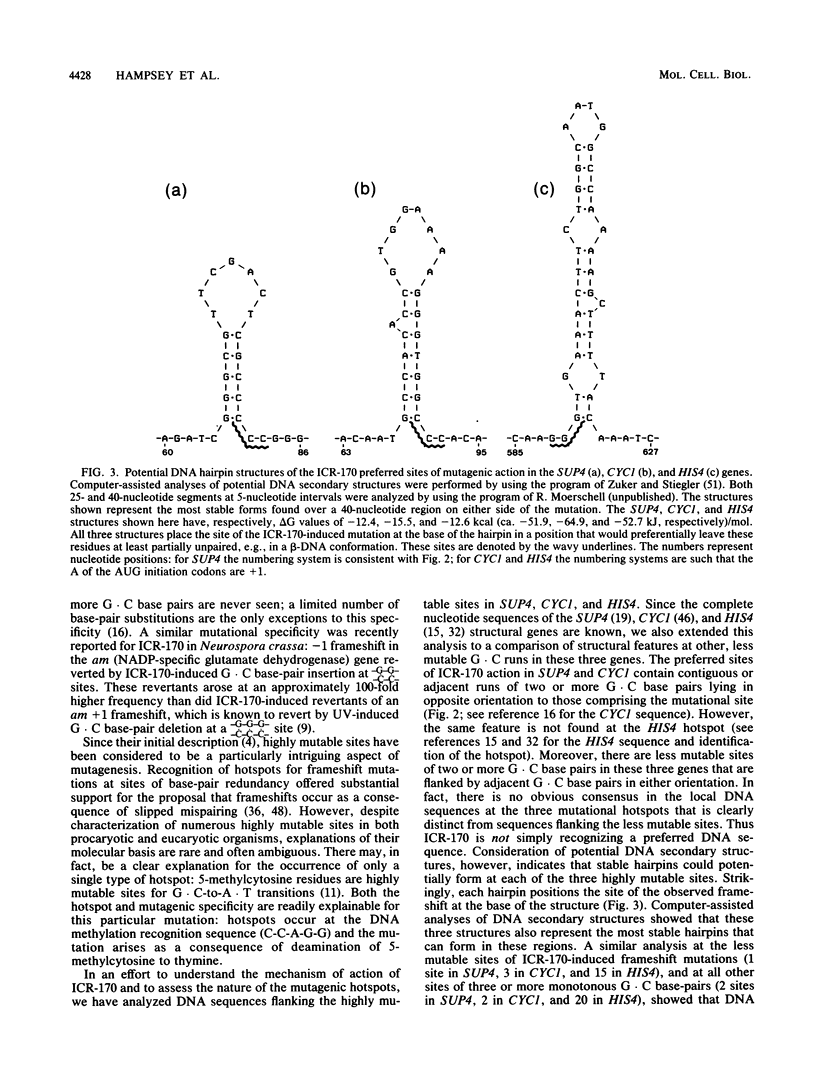
The majority of the mutations induced by ICR-170 in both the CYC1 gene (J. F. Ernst et al. Genetics 111:233-241, 1985) and the HIS4 gene (L. Mathison and M. R. Culbertson, Mol. Cell. Biol. 5:2247-2256, 1985) of the yeast Saccharomyces cerevisiae were recently shown to be single G . C base-pair insertions at monotonous runs of two or more G . C base pairs. However, not all sites were equally mutable; in both the CYC1 and HIS4 genes there is a single highly mutable site where a G . C base pair is preferentially inserted at a [sequence in text]. Here we report the ICR-170 mutagen specificity at the SUP4-o tyrosine tRNA gene of yeast. Genetic fine structure analysis and representative DNA sequence determination of ICR-170-induced mutations revealed that there is also a single highly mutable site in SUP4-o and that the mutation is a G . C base-pair insertion at a monotonous run of G . C base pairs. Analysis of DNA sequences encompassing the regions of highly mutable sites for all three genes indicated that the mutable sites are at the bases of potential hairpin structures; this type of structure could not be found at any of the other, less mutable G . C runs in SUP4, CYC1, and HIS4. Based on these results and recent information regarding novel DNA structural conformations, we present a mechanism for ICR-170-induced mutagenesis. (i) ICR-170 preferentially binds to DNA in the beta conformation; factors that increase the temporal stability of this structure, such as adjacent stem-and-loop formation, increase the frequency of ICR-170 binding; (ii) the observed mutagen specificity reflects formation of a preferred ICR-170 intercalative geometry at [sequence in text] sites; (iii) during replication or repair, ICR-170 remains associated with the single-stranded template; (iv) stuttering or strand slippage by the polymerization complex as it encounters the mutagen results in nucleotide duplication; (v) subsequent replication or mismatch repair fixes the insertion into the genome. This mechanism accounts for both the IRC-170 mutagenic specificity and the molecular basis of the highly mutable sites in S. cerevisiae.
Full text
PDF






Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allison D. S., Goh S. H., Hall B. D. The promoter sequence of a yeast tRNAtyr gene. Cell. 1983 Sep;34(2):655–664. doi: 10.1016/0092-8674(83)90398-7. [DOI] [PubMed] [Google Scholar]
- Banerjee A., Sobell H. M. Presence of nonlinear excitations in DNA structure and their relationship to DNA premelting and to drug intercalation. J Biomol Struct Dyn. 1983 Oct;1(1):253–262. doi: 10.1080/07391102.1983.10507438. [DOI] [PubMed] [Google Scholar]
- Benton W. D., Davis R. W. Screening lambdagt recombinant clones by hybridization to single plaques in situ. Science. 1977 Apr 8;196(4286):180–182. doi: 10.1126/science.322279. [DOI] [PubMed] [Google Scholar]
- Benzer S. ON THE TOPOGRAPHY OF THE GENETIC FINE STRUCTURE. Proc Natl Acad Sci U S A. 1961 Mar;47(3):403–415. doi: 10.1073/pnas.47.3.403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biggin M. D., Gibson T. J., Hong G. F. Buffer gradient gels and 35S label as an aid to rapid DNA sequence determination. Proc Natl Acad Sci U S A. 1983 Jul;80(13):3963–3965. doi: 10.1073/pnas.80.13.3963. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brusick D. J. The mutagenic acitivity of ICR-170 in Saccharomyces cerevisiae. Mutat Res. 1970 Jul;10(1):11–19. doi: 10.1016/0027-5107(70)90141-7. [DOI] [PubMed] [Google Scholar]
- Burns P. A., Kinnaird J. H., Kilbey B. J., Fincham J. R. Sequencing studies of ICR-170 mutagenic specificity in the am (NADP-specific glutamate dehydrogenase) gene of Neurospora crassa. Genetics. 1986 May;113(1):45–51. doi: 10.1093/genetics/113.1.45. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Genetic and sequence analysis of frameshift mutations induced by ICR-191. J Mol Biol. 1981 Nov 25;153(1):39–64. doi: 10.1016/0022-2836(81)90525-8. [DOI] [PubMed] [Google Scholar]
- Coulondre C., Miller J. H., Farabaugh P. J., Gilbert W. Molecular basis of base substitution hotspots in Escherichia coli. Nature. 1978 Aug 24;274(5673):775–780. doi: 10.1038/274775a0. [DOI] [PubMed] [Google Scholar]
- Culbertson M. R., Charnas L., Johnson M. T., Fink G. R. Frameshifts and frameshift suppressors in Saccharomyces cerevisiae. Genetics. 1977 Aug;86(4):745–764. doi: 10.1093/genetics/86.4.745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donahue T. F., Farabaugh P. J., Fink G. R. Suppressible four-base glycine and proline codons in yeast. Science. 1981 Apr 24;212(4493):455–457. doi: 10.1126/science.7010605. [DOI] [PubMed] [Google Scholar]
- Donahue T. F., Farabaugh P. J., Fink G. R. The nucleotide sequence of the HIS4 region of yeast. Gene. 1982 Apr;18(1):47–59. doi: 10.1016/0378-1119(82)90055-5. [DOI] [PubMed] [Google Scholar]
- Ernst J. F., Hampsey D. M., Sherman F. DNA sequences of frameshift and other mutations induced by ICR-170 in yeast. Genetics. 1985 Oct;111(2):233–241. doi: 10.1093/genetics/111.2.233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farabaugh P. J. Sequence of the lacI gene. Nature. 1978 Aug 24;274(5673):765–769. doi: 10.1038/274765a0. [DOI] [PubMed] [Google Scholar]
- Golding G. B., Glickman B. W. Sequence-directed mutagenesis: evidence from a phylogenetic history of human alpha-interferon genes. Proc Natl Acad Sci U S A. 1985 Dec;82(24):8577–8581. doi: 10.1073/pnas.82.24.8577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodman H. M., Olson M. V., Hall B. D. Nucleotide sequence of a mutant eukaryotic gene: the yeast tyrosine-inserting ochre suppressor SUP4-o. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5453–5457. doi: 10.1073/pnas.74.12.5453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guo L. H., Wu R. Exonuclease III: use for DNA sequence analysis and in specific deletions of nucleotides. Methods Enzymol. 1983;100:60–96. doi: 10.1016/0076-6879(83)00046-4. [DOI] [PubMed] [Google Scholar]
- Hampsey D. M., Das G., Sherman F. Amino acid replacements in yeast iso-1-cytochrome c. Comparison with the phylogenetic series and the tertiary structure of related cytochromes c. J Biol Chem. 1986 Mar 5;261(7):3259–3271. [PubMed] [Google Scholar]
- Hsu M. T. Electron microscopic evidence for the cruciform structure in intracellular SV40 DNA. Virology. 1985 Jun;143(2):617–621. doi: 10.1016/0042-6822(85)90400-3. [DOI] [PubMed] [Google Scholar]
- Jain S. C., Bhandary K. K., Sobell H. M. Visualization of drug--nucleic acid interactions at atomic resolution. VI. Structure of two drug--dinucleoside monophosphate crystalline complexes, ellipticine--5-iodocytidylyy (3'-5') guanosine and 3,5,6,8-tetramethyl-N-methyl phenanthrolinium--5-iodocytidylyl (3'-5') guanosine. J Mol Biol. 1979 Dec 25;135(4):813–840. [PubMed] [Google Scholar]
- Koski R. A., Allison D. S., Worthington M., Hall B. D. An in vitro RNA polymerase III system from S. cerevisiae: effects of deletions and point mutations upon SUP4 gene transcription. Nucleic Acids Res. 1982 Dec 20;10(24):8127–8143. doi: 10.1093/nar/10.24.8127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurjan J., Hall B. D., Gillam S., Smith M. Mutations at the yeast SUP4 tRNATyr locus: DNA sequence changes in mutants lacking suppressor activity. Cell. 1980 Jul;20(3):701–709. doi: 10.1016/0092-8674(80)90316-5. [DOI] [PubMed] [Google Scholar]
- Kurjan J., Hall B. D. Mutations at the Saccharomyces cerevisiae SUP4 tRNA(Tyr) locus: isolation, genetic fine-structure mapping, and correlation with physical structure. Mol Cell Biol. 1982 Dec;2(12):1501–1513. doi: 10.1128/mcb.2.12.1501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LERMAN L. S. Structural considerations in the interaction of DNA and acridines. J Mol Biol. 1961 Feb;3:18–30. doi: 10.1016/s0022-2836(61)80004-1. [DOI] [PubMed] [Google Scholar]
- LERMAN L. S. The structure of the DNA-acridine complex. Proc Natl Acad Sci U S A. 1963 Jan 15;49:94–102. doi: 10.1073/pnas.49.1.94. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo K. M., Jones S. S., Hackett N. R., Khorana H. G. Specific amino acid substitutions in bacterioopsin: Replacement of a restriction fragment in the structural gene by synthetic DNA fragments containing altered codons. Proc Natl Acad Sci U S A. 1984 Apr;81(8):2285–2289. doi: 10.1073/pnas.81.8.2285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malling H. V. The mutagenicity of the acridine mustard (ICR-170) and the structurally related compounds in Neurospora. Mutat Res. 1967 May-Jun;4(3):265–274. doi: 10.1016/0027-5107(67)90021-8. [DOI] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Mathison L., Culbertson M. R. Suppressible and nonsuppressible +1 G-C base pair insertions induced by ICR-170 at the his4 locus in Saccharomyces cerevisiae. Mol Cell Biol. 1985 Sep;5(9):2247–2256. doi: 10.1128/mcb.5.9.2247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Farrell P. H., Kutter E., Nakanishi M. A restriction map of the bacteriophage T4 genome. Mol Gen Genet. 1980;179(2):421–435. doi: 10.1007/BF00425473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy B. S., Seshadri T. P., Sakore T. D., Sobell H. M. Visualization of drug--nucleic acid interactions at atomic resolution. V. Structure of two aminoacridine--dinucleoside monophosphate crystalline complexes, proflavine--5-iodocytidylyl (3'-5') guanosine and acridine orange--5-iodocytidylyl (3'-5') guanosine. J Mol Biol. 1979 Dec 25;135(4):787–812. doi: 10.1016/0022-2836(79)90513-8. [DOI] [PubMed] [Google Scholar]
- Ripley L. S., Glickman B. W., Shoemaker N. B. Mutator versus antimutator activity of a T4 DNA polymerase mutant distinguishes two different frameshifting mechanisms. Mol Gen Genet. 1983;189(1):113–117. doi: 10.1007/BF00326062. [DOI] [PubMed] [Google Scholar]
- Ripley L. S., Glickman B. W. Unique self-complementarity of palindromic sequences provides DNA structural intermediates for mutation. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):851–861. doi: 10.1101/sqb.1983.047.01.097. [DOI] [PubMed] [Google Scholar]
- Roth J. R. Frameshift mutations. Annu Rev Genet. 1974;8:319–346. doi: 10.1146/annurev.ge.08.120174.001535. [DOI] [PubMed] [Google Scholar]
- Rothstein R. Deletions of a tyrosine tRNA gene in S. cerevisiae. Cell. 1979 May;17(1):185–190. doi: 10.1016/0092-8674(79)90306-4. [DOI] [PubMed] [Google Scholar]
- Sakore T. D., Reddy B. S., Sobell H. M. Visualization of drug-nucleic acid interactions at atomic resolution. IV. Structure of an aminoacridine--dinucleoside monophosphate crystalline complex, 9-aminoacridine--5-iodocytidylyl (3'--5') guanosine. J Mol Biol. 1979 Dec 25;135(4):763–785. doi: 10.1016/0022-2836(79)90512-6. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skopek T. R., Hutchinson F. Frameshift mutagenesis of lambda prophage by 9-aminoacridine, proflavin and ICR-191. Mol Gen Genet. 1984;195(3):418–423. doi: 10.1007/BF00341442. [DOI] [PubMed] [Google Scholar]
- Smith A. J. DNA sequence analysis by primed synthesis. Methods Enzymol. 1980;65(1):560–580. doi: 10.1016/s0076-6879(80)65060-5. [DOI] [PubMed] [Google Scholar]
- Smith M., Leung D. W., Gillam S., Astell C. R., Montgomery D. L., Hall B. D. Sequence of the gene for iso-1-cytochrome c in Saccharomyces cerevisiae. Cell. 1979 Apr;16(4):753–761. doi: 10.1016/0092-8674(79)90091-6. [DOI] [PubMed] [Google Scholar]
- Sobell H. M. Actinomycin and DNA transcription. Proc Natl Acad Sci U S A. 1985 Aug;82(16):5328–5331. doi: 10.1073/pnas.82.16.5328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Streisinger G., Okada Y., Emrich J., Newton J., Tsugita A., Terzaghi E., Inouye M. Frameshift mutations and the genetic code. This paper is dedicated to Professor Theodosius Dobzhansky on the occasion of his 66th birthday. Cold Spring Harb Symp Quant Biol. 1966;31:77–84. doi: 10.1101/sqb.1966.031.01.014. [DOI] [PubMed] [Google Scholar]
- Streisinger G., Owen J. Mechanisms of spontaneous and induced frameshift mutation in bacteriophage T4. Genetics. 1985 Apr;109(4):633–659. doi: 10.1093/genetics/109.4.633. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Todd P. A., Glickman B. W. Mutational specificity of UV light in Escherichia coli: indications for a role of DNA secondary structure. Proc Natl Acad Sci U S A. 1982 Jul;79(13):4123–4127. doi: 10.1073/pnas.79.13.4123. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]


